BME100 f2013:W1200 Group9 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
OUR COMPANY
LAB 6 WRITE-UPComputer-Aided DesignTinkerCAD
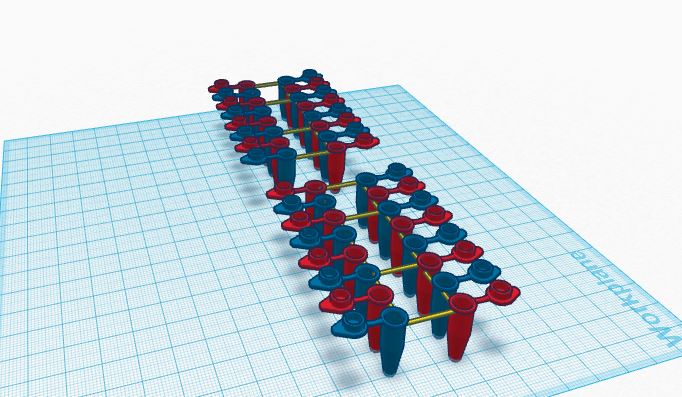
Our group decided to take the already created PCR tubes and color coded them to that they we would be able to identify whether they were positive or negative. This way we can save time without the extra labeling and as we discovered while performing this lab, if you accidentally make a mistake, the writing easily becomes illegible. Labeling would be a huge problem when working with PCR because interpretation of the wrong data could drastically affect the results. We also decide to create thirty-two PCR tubes that are attached by cheap plastic that is breakable or able to be separated with scissors. This way all your tubes are in one place and you can easily break off the desired amount from the rest of the tubes while staying organized and well kept in one place.
Feature 1: Cancer SNP-Specific PrimersBackground on the cancer-associated mutation Rs17879961, studied in our lab, is a mutation described in the Database of short genetic variations (dbSNP) that is cancer-related. It is categorized as a Single Nucleotide Polymorphism (SNP)which means the mutation is an occurrence of a different variation of the gene among a population, affecting the structural unit of nucleic acid in RNA and DNA. This polymorphism is a pathogenic SNP affecting the twenty-second chromosome and is only found in Homo Sapiens, who usually have twenty-three pairs chromosomes. This gene was affected by Checkpoint Kinase 2 (CHEK2) where the mutation of this gene was found to cause breast cancer, brain tumors, and predisposition of sarcomas.
How the primers work: For a primer to begin, a specific target sequence is required. In this lab, our target sequence was the cancer specific SNP, rs17879961. This cancer specific reverse primer will only bind to a complementary match of the sequence only if it matches the primer completely. If it is not a perfect match, the strand will not duplicate because it is unable to bind together. With this specific cancer gene, only cancer primers can be replicated and non-cancer primers will not.
Feature 2: Consumables KitThe packaging of our consumables will be very similar to the way they are currently packaged. The plastic tubes will come in rows of sixteen, eight tubes on each side (as shown in the image above). These tubes will come in cardboard packages and will hold 240 tubes per package. each package will include a rack for storing the tubes. Two racks holding 120 tubes each will be included with each package of 240 tubes. The micropipeter will be packaged separately and will include 200 plastic tips with purchase. These tips will be racked in a plastic rack and extra tips will also be available for purchase in quantities of 50 and 100. The PCR mix and the primers will also be packaged together. All of the items in the consumables kit are available as one package for a set price and are also available for separate sale. Our group decided to do this so that consumers could buy all the necessary items they needed at one reasonable price or they could buy separate items if they only needed a few individual things.
One of the problems that our group mentioned was the lack of labeling on the plastic tubes. The tubes that we designed on TinkerCad are color coated in order to help differentiate the different primers, and DNA samples. The packages of tubes will also come with small labeling strips so that consumers will be able to label their specific tubes and have some way to keep their samples organized.
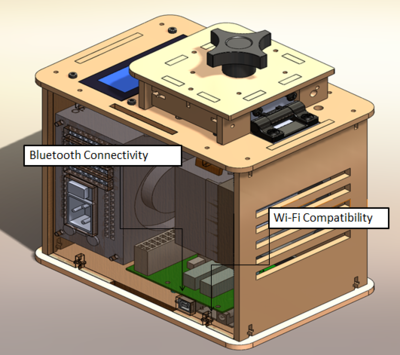
Feature 3: PCR Machine Hardware
Feature 4: Fluorimeter HardwareThe flourimeter hardware was ineffecient in sustaining the cellular device which would take the picture of the DNA. This would cause pictures to be of poor quality or of nothing that was of use for the lab. Therefore, with a an adjustable fluorimeter hardware any phones of any size can be sustained adequately with the complete disregard of whether or not the phone will fall at any given moment. This improvement will help overall as the pictures will not be of bad quality.
Bonus Opportunity: What Bayesian Stats Imply About The BME100 Diagnostic Approach[Instructions: This section is OPTIONAL, and will get bonus points if answered thoroughly and correctly. Here is a chance to flex some intellectual muscle. In your own words, discuss what the results for calculations 3 and 4 imply about the reliability of CHEK2 PCR for predicting cancer. Please do NOT type the actual numerical values here. Just refer to them as being "less than one" or "very small." The instructors will ask you to submit your actual calculations via e-mail. We are doing so for the sake of academic integrity and to curb any temptation to cheat.] |
||||||