BME100 f2013:W1200 Group9 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||
|
OUR TEAM
LAB 1 WRITE-UPInitial Machine TestingThe Original Design
After unplugging part 3 from part 6 of the PCR machine, the LED screen shut off. Once we reconnected the wire, the light came on again. From this information we can conclude that parts 3 and 6 are connected to the bulb that lights up the screen. When we unplugged the white wire that connects (part 6) to (part 2), the temperature on the screen dropped.
The date that we first worked with Open PCR was October 22, 2013 at 9:00 am. The number of the machine our group was assigned was 10.
ProtocolsThermal Cycler Program
PCR Reaction Mix
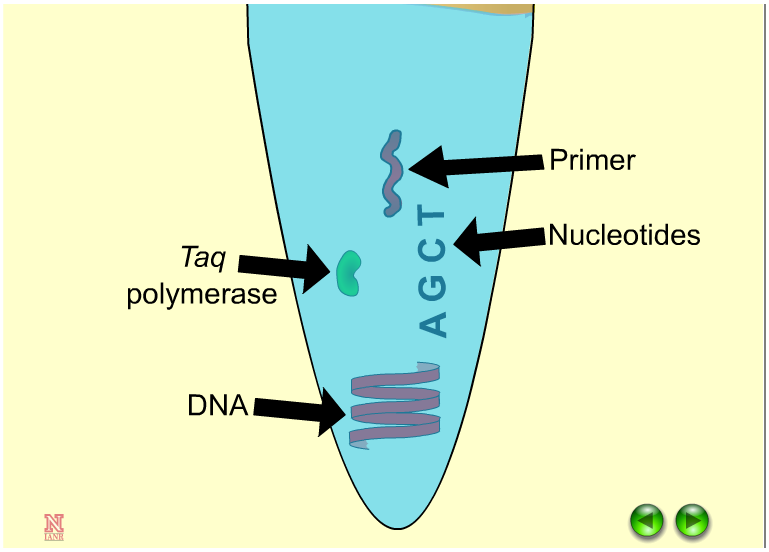
Research and DevelopmentPCR - The Underlying Technology PCR has various components. The multiple aspects have an important role in multiplying and duplicating DNA. The PCR also has an important role in a sensitive test for tissue typing and vital for organ transplants but the main cause we will be studying is the study of patterns of gene expression. At the start of the process, a template DNA labels the DNA selected to copy and the RNA protein which will be created. Primers are the attaching component to the PCR reaction. Simultaneously, the primers copy the DNA and stop the polymerase from attaching to the template DNA. A Taq polymerase are high resisting heat molecules that attach to the primers to match the nucleotides and copy the DNA. Magnesium Chloride is the catalyst within this reaction which helps to tag the polymerase. The last component of a PCR reaction are deoxyribonucleotides abbreviated as (dNTP's) which pairs up with the complementary bases to replicate DNA strands.
During thermal cycling the samples are heated up at ninety-five degrees Celsius for three minutes. Then at the same temperature for thirty seconds the DNA is denatured, where the helix separates creating two separate strands. The temperature then drops to fifty-seven degrees Celsius for thirty seconds where the primers anneal to their complementary strands, Adenine pairs with Thymine and Cytosine pairs with Guanine. The temperature is again increased where the tag polymerase attaches at the priming site and extends to form a brand new DNA strand. The last step stays at the same temperature where the polymerase continues to the end of the strand until it falls off. The final hold is dropped from seventy-two to four degrees Celsius to prevent potential nonspecific binding of primers and amplification.
| |||||||||||||||