BME100 f2013:W1200 Group1 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
OUR COMPANY
LAB 6 WRITE-UPComputer-Aided DesignTinkerCAD Implications of Using TinkerCAD for Design
Feature 1: Cancer SNP-Specific PrimersBackground on the cancer-associated mutation The cancer-associated mutation that was studied in lab was rs17879961, which is a SNP standing for single nucleotide polymorphism. A nucleotide is an organic molecule that serves as the monomer of nucleic acids such as DNA and RNA. Polymorphism is when two or more different phenotypes exist in the same population of a species. rs17879961 is found in the the species homo sapiens. This SNP is considered pathogenic, which means that is causes a disease. It is found on chromosome number 22. Typically, humans have 23 pairs of chromosomes. The gene affected from a SNP is called CHEK2. This stands for Checkpoint Kinase 2. CHEK2 is a cell cycle checkpoint regulator and putative tumor suppressor that stabilizes the tumor suppressor protein p53.
Primer design
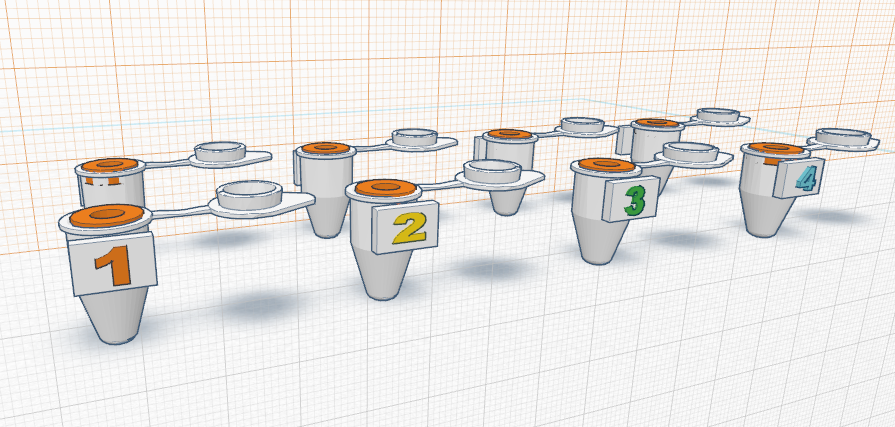
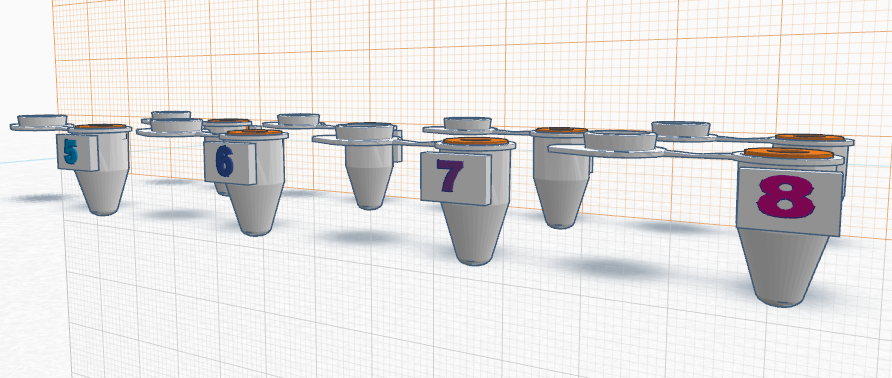
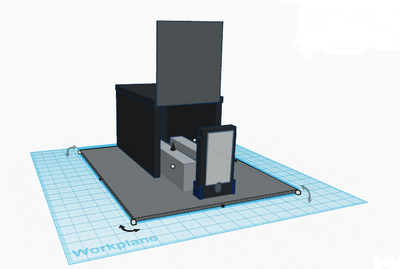
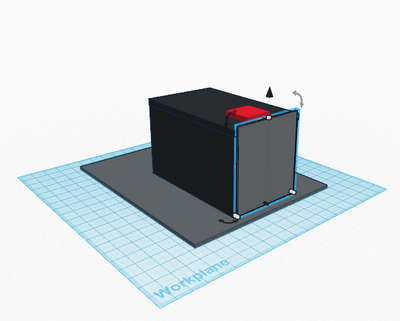
How the primers work: Primers work by first identifying a target sequence in a certain segment of DNA. In this lab, the cancer-associated SNP is rs17879961. The cancer-specific reverse primer will bind to the complementary cancer-SNP template. In order for proper binding to occur, the primer and the complementary sequence must match. The forward primer binds similarly to the reverse primer by matching to complementary sequences. If primers do not bind in a complementary fashion, the DNA will not be replicated. Once the primers are blinded complementary, DNA polymerase is directly to synthesize the complementary DNA strands. DNA polymerase only replicates segments with the attached primer. For this reason, the non-cancer alleles will not be amplified because it does not contain a primer. Feature 2: Consumables KitThe new kit will follow the same general structure of the one used in class, however it will be improved upon the security and ease of use of the consumables. The consumables will come packaged in the same plastic containers, however they will not be packaged within the same plastic wrap. In order to reduce the chance of potential contamination, all consumables will be packaged individually. This will ensure that accidental spillage or cross-contamination is not possible. Also, each consumable will have a designated spot within a styrofoam cutout to reduce the chance of breaking during transportation and shipment. Lastly, an instruction booklet will also be included in the kit, in order to teach the user how to properly use and dispose of each piece. The booklet will provide clear information of the kit including pictures for every step, and, to minimize confusion, each component will be separately labeled with a corresponding color and letter in the instructions booklet. Feature 3: PCR Machine HardwareThe PCR machine will be incorporated into the new redesigned system in the same way that it is already used. Similar to the setup in class, the PCR machine will be used solely to heat and cool the samples, allowing for the DNA contained within the Eppendorf tubes to multiply rapidly. In the new system, the PCR machine will heat and cool the samples to the same temperatures as the current one because this has been established to be most successful at multiplying DNA, however, it will be significantly more efficient. The modified PCR machine will be more efficient and will improve the previous one on its major weaknesses. One problem associated with the previous PCR machine was the fact that it was supposed to be programmed via computer. The redesigned PCR machine will be programmable from the machine itself, removing the need for time consuming and challenging computer programming. Another problem with the original machine is that a user cannot see the process in which the heating plate is in direct contact with the Eppendorf tubes. To solve this problem, the new redesigned machine would have a clear lid with a pressure sensor. This would allow the user to ensure that the samples don't spill over and that the heating plate is in direct contact with the samples. An additional improvement is that the heating plate would be larger and the Eppendorf tubes would be smaller. Rather than the current 16 well plate, a 96 well plate would be incorporated as to allow more samples to be tested at one time, eliminating the need of having to run countless PCR processes. Another drawback of the current machine is that the complete process takes hours. It was noticed that there was a fair amount of open space inside the PCR machine. This open space would be filled by making a larger and more powerful heating/cooling system, which would effectively allow the temperatures to change much more rapidly and would significantly reduce the time needed for the PCR process. Lastly, the brand name of redesigned machine "Packin' Heat" would be added to the side walls of the PCR machine. source:(http://www.wisbiomed.com/ec/index.php?cPath=12&main_page=index) Feature 4: Fluorimeter HardwareSimilar to the fluorimeter used in lab, our redesigned fluorimeter will be used to shine a beam of blue light through the drop of sample containing DNA. Then pictures of the drop will be taken in complete darkness. However, the fully upgraded and redesigned fluorimeter will consist of the fluorimeter with LED light, an automated closing lid, and a built in camera that works in sync with the lid. This setup will result in more accurate and higher quality photos. It would also ensure that all pictures are the exact same. As previously mentioned, the improved fluorimeter would improve on a few of the weaknesses of the current model. One major problem is that it is not accurate and convenient to keep the phone in place and at the same distance away from the sample drop and beam of light. The new redesigned fluorimeter improves on this weakness by including an adjustable clamp that holds the phone ins the same position. The upgraded model with built-in camera and an automated lid would ensure that ideal pictures are taken every time and that pictures are only taken when the lid is completely placed. With a simple press of a button, the lid would shut and a perfect picture would be taken when the lid is completely lowered. This eliminates the possibility of user error. Bonus Opportunity: What Bayesian Stats Imply About The BME100 Diagnostic ApproachBayesian statistics is a method to determine the reliability of detecting cancer SNPs and predicting cancer. The equation for Bayesian statistics is: P(A|B)=P(B|A)*P(A)/P(B). Calculation 3 asks what is the probability that the patient will develop cancer, given a cancer DNA sequence. Because the P(A|B) value for this calculation is very small and less than one, it is not accurate at all. Calculation 4 asks what is the probability that the patient will not develop cancer, given a non-cancer DNA sequence. Because the P(A|B) value for this calculation is also very small and less than one, it is not accurate at all either. Thus, the reliability of CHEK2 PCR for predicting cancer is not good and should not be used. The calculations may not have produced reliable results because of human error using the OpenPCR machines. Many of the trials were not completed to full term, which threw out samples and may have skewed the overall data with a smaller amount of trials than expected. In order for the CHEK2 PCR to actual be a reliable predictor for cancer, the values of P(A|B) for calculation 3 and 4 should be very close to one instead of being such small numbers.
Bonus Opportunity #2: Instruction manual for PCR machine |
||||||