BME100 f2013:W1200 Group17 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |
|
OUR TEAMLAB 1 WRITE-UPInitial Machine Testing
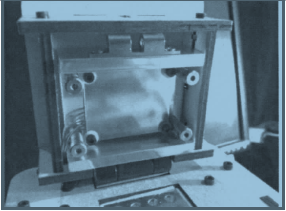
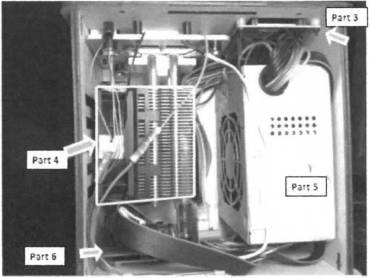
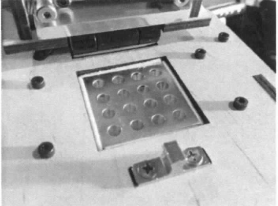
Experimenting With the Connections When we unplugged the LCD display (part 3) from the circuit board (part 6), the screen shut off. When we unplugged the white wire that connects the circuit board (part 6) to the heating plate (part 2), the recorded temperature was removed from the LCD display.
Open PCR was first tested October 23, 2013. The machine was fairly cheaply made and the DNA replications were slow. The DNA replications did not happen at the rate expected.The PCR was slow and did not calibrate correctly when it was connected to the computer. The total time that was expected to run the heating and cooling protocol was 2 hours or less. Within 15 minutes during the operation of the machine, the test run was failing. The displayed results on the screen was slower than the expected compared to the computer monitor. As of result, the machine failed to pass the test run.
ProtocolsThermal Cycler Program
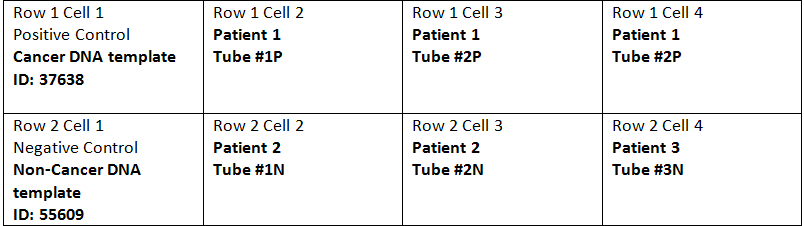
DNA Set-up Procedure Step 1) Label tubes according to the chart above. Of the 6 tubes given, 3 tubes must be for the Positive Control (Cancer DNA template) and 3 other tubes must be for the Negative Control (Non-Cancer DNA template). The ones that are for the Positive Control must be labeled #1P, #2P, #3P respectively. The tubes for the Negative Control must be labeled #1N, #2N, and #3N respectively. They can be labeled with a dry erase marker or with sharpie on a small piece of tape that is attached to each tube. Step 2) Move DNA/Primer mix into correct PCR tubes with a pipette tip (50 microliter in each). Only use each pipette tip once. Dispose of pipette tip after each transfer. Step 3) Put PCR reaction mix (50 microliter) into each PCR tube with pipette tip (reaction mix includes DNA polymerase, magnesium chloride, and dNTP's). Only use each pipette tip once. Dispose of pipette tip after each transfer. Step 4) Place PCR tube filled with all reaction components into a DNA Thermal Cycler (PCR machine) Step 5) Set the Open PCR Software to run a specific program in order to heat/cool the DNA appropriately. Make sure the HEATED LID portion is set at 100 degrees Celsius and the INITIAL STEP section is set at 95 degrees Celsius for 3 minutes. Make sure the DNA Thermal Cycler runs for 35 cycles denature at 95 degrees Celsius for 30 seconds, anneal at 57 degrees Celsius for 30 seconds, extend at 72 degrees Celsius for 30 seconds. Next it should be set at 72 degrees Celsius for 3 minutes, and the final hold should be at 4 degrees Celsius. Step 6) Close lid of DNA cycler and press start to collect data. PCR Reaction Mix The PCR reaction mix includes Taq DNA polymerase, MgCl2(Magnesium Cholride), and dNTP's(deoxynucleotide triphosphates ). We used 8 tubes and each had 5 microliters of the mix in each.
DNA/ primer mix Each DNA/Primer mix contains a different template DNA. All tubes have the same forward primer and reverse primer.
Research and DevelopmentPCR - The Underlying Technology What is PCR, and Why Do We Care? Polymerase Chain Reaction, commonly called 'PCR', is a relatively simple technology that provides an invaluable tool to the research biologist. PCR uses naturally found enzymes to quickly make large numbers of identical copies of the original "template DNA" that one desires to copy. PCR gives effectively gives scientists and endless supply of any DNA they are able to extract and isolate, giving them the freedom to perform large numbers of experiments and diagnostics that would otherwise be impossible with the tiny amount of DNA that is typically extracted from sources. What is Needed to Perform PCR, and Exactly What is Going on in Those Tubes? Perhaps the most important ingredient necessary for PCR is the Template DNA. This is the DNA of interest, the master copy that the scientist intends to reproduce thousands or millions of times. In PCR, this template (which is double stranded like most DNA) is denatured and split into two complimentary single strands. This allows for binding of the primers. Primers are small, single-stranded pieces of DNA that have been engineered to bind to the template DNA at a specific place and serve as the origin of replication. Once the primer is bound, this allows for the attachment of the DNA Polymerase, which begins the process of DNA replication. The polymerase used in PCR is uniquely thermostable. Most DNA polymerases denature and are destroyed in the high temperatures necessary to denature the template DNA, preventing the automation of multiple cycles. PCR uses Taq Polymerase, a DNA polymerase extracted from bacteria found in hot springs, that is capable of withstanding the extreme temperatures needed for PCR and allows for its repeated use over many cycles. Once the Taq Polymerase is bound, it reads the template strand and adds the required complimentary building blocks, known as deoxyribonucelotide triphosphates (dNTPs), to copy the DNA. dNTPs are the individual nucleic acids that join together to make DNA. There are 4 deoxyribonucleotides: Adenine (A) Thymine (T) Guanine (G) and Cytosine (C). A and T are compliments, as are G and C. Taq Polymerase continues down the template DNA strand, adding the complimentary dNTP's until the replication cycle has completed. Taq Polymerase needs a particular concentration of Magnesium Chloride in its presence to effectively function; too little Magnesium Chloride will cause the polymerase to work slowly and inefficiently, while too much Magnesium Chloride will cause the polymerase to work too quickly and make errors in replication. This process of PCR is performed in many consecutive cycles, with the number of DNA copies doubling with each consecutive cycle. How Does a PCR Thermocycler Work? What Steps are Needed for Proper PCR? 1. Initial Step - Here, the PCR machine is heated at 95°C for 3 minutes. This is only done once, in the beginning of the process. The purpose is to bring the machine up to a constant temperature before beginning the PCR process and to thaw out any necessary ingredients that may have been frozen 2. Denaturing - This step signifies the beginning of each PCR cycle. Here, the DNA is heated to 95°C for 30 seconds. This heat causes the template DNA strands to denature and split from double stranded to single stranded 3. Annealing - This next step, in which the temperature is held at 57°C for 30 seconds, is when the small primers will find the sequences on the template DNA they were designed to bind and serve as origins of replication 4. Extending - This is the last step of the PCR cycle, where the temperature is held at 72°C for 30 seconds. In this step the Taq Polymerase binds to the primers on the template DNA and replicates it by reading the template strand and adding the appropriate dNTP's as it progresses. 5. Final Step - At the end of the desired number of PCR cycles, the temperature is held at 72°C for 3 minutes. This step allows the polymerase to finish its last cycle and signifies the end of the PCR process 6. Final Hold - In the event that the newly created DNA will be stored for later use, the DNA is chilled to 4°C and held there until it is removed from the thermocycler. This prevents DNA degradation and unwanted water condensation.
| |