BME100 f2013:W1200 Group12 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||
|
GROUP 12LAB 1 WRITE-UPInitial Machine TestingThe Original Design "Completed OpenPCR Machine." Photo.openpcr.org, n.d. 28 Oct. 2013 [1] An Open PCR is a machine that will duplicate a desired strand of DNA. It can generate 100 billion copies of a specific strand of DNA in a matter of hours. The process of making billions of copies is called amplification. It is used to diagnose disease, identify viruses, bacteria, and many other things. They way it works is it starts off by heating up the double stranded DNA to near boiling point. This causes the DNA to denature, or separate, into two strands. Then the temperature is greatly decreased causing primers, which are DNA sequences, to attach to complementary matches on the target DNA. The primers make it so the DNA can be copied. The temperature is slightly raised again causing an enzyme to attach to the primers and adds nucleotides to extend the second strand. That is the first cycle. This happens about 34 more times to make a total of 35 cycles. Experimenting With the Connections (extracted from OpenPCR Lab Worksheet 1 page 2) (extracted from OpenPCR Lab Worksheet 1 page 2) "Internal Components of OpenPCR Machine." Photo.openpcr.org, n.d. 28 Oct. 2013 [2]
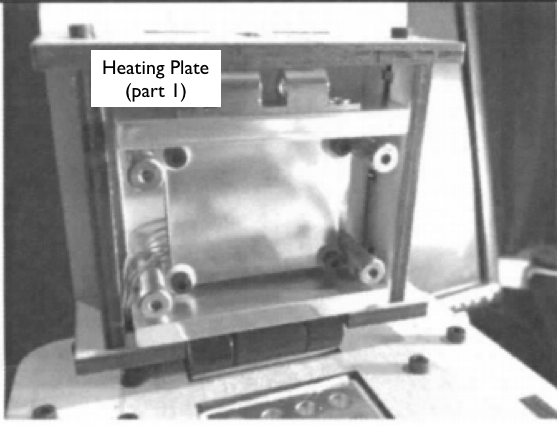
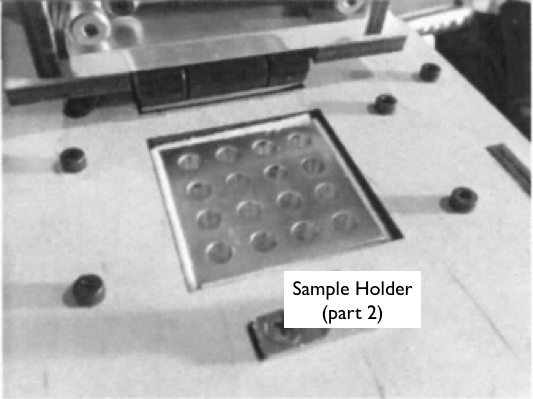
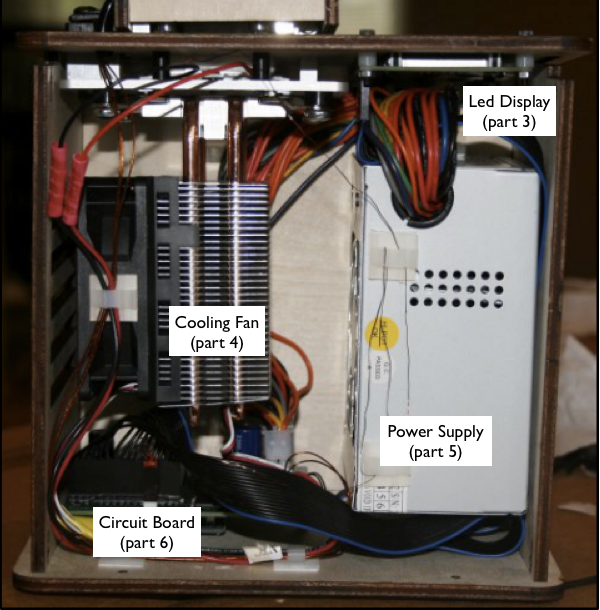
When we unplugged the white wire that connects (part 6) to (part 2), the machine's display stopped reading the temperature of the machine. That leads us to believe that part six is the machines thermometer. Part 1 in the above picture is a heating plate and lid which is used to maintain a greater temperature at the top of the test tubes which prevents the DNA samples from evaporating and rising to the top. Part number 2 is the test tube sample holder which is needed to hold the DNA samples as they are heated and cooled throughout the experiment.
The Open PCR machine was tested on October 23, 2013. We performed a simple test with the machine connected to a computer in order to see if the machine properly went through all of its cycles. The conditions the test was run under were to heat the heating lid to 100°C , with an initial step of 95°C for 3 minutes. We ran 35 cycles for the step, and denatured the faux test samples at 95°C for 30 seconds, then annealed the samples at 57°C for 30 seconds. The extend was set to 72°C for a 30 second duration, and the final step was 72°C for 3 minutes. Finally, the final hold was set to 4°C .The Open PCR functioned properly and completed its cycles without any issues. The overall experience was good, although some of the cycles took a long time, which was expected.
ProtocolsThermal Cycler Program
There are 8 test tubes that are 50 microliters each. The mix contains taq DNA polymerase, MgCL2, as well as dNTP's.
There are 8 test tubes that are 50 microliters each. Each mix contains a different template DNA and all test tubes have the same forward primer as well as the same reverse primer. PCR - The Underlying TechnologyFunction of each component of a PCR reaction Template DNA At the beginning of the reaction high temperature is applied to the original DNA molecule to separate the strands from each other. The template DNA is the sample DNA that contains the beginning sequence to which the reactions will be derived from. Taq Polymerase Taq polymerase is the most common polymerase used in synthesizing new strands of DNA. Taq DNA has a higher fidelity when the DNA is copied. Polymerases use DNA templates and primers to generate new strands of DNA that are heat resistant. Magnesium Chloride (MgCl¬2) Magnesium Chloride acts as a catalyst in the reaction because the Thermo stable polymerase requires the presence of magnesium to act as a cofactor throughout the reaction process. Because of this Magnesium chloride is the preferred method of adding magnesium to the PCR. Deoxyribonucleotides (dNTP’s) dNTP’s are the building blocks of new DNA strands. dNTP’s are the single units of the bases A,T,G, and C.
Initial step: Heat to 95°C for 3 minutes The DNA is separated into right and left sides because the water is almost boiling. The reason it is for 3 minutes is to make sure all of the DNA has been separated. Step 1: Heat to 95°C for 30 seconds This step is similar to the initial step, where the purpose is to separate DNA into two pieces, but this is the first step of the cycle. It only lasts 30 seconds because the DNA will be smaller and smaller as the target area is separated from the rest. Step 2: 57°C for 30 seconds This step is to allow the primers to attach to the DNA. One primer will connect at point A of the target strand while the other will connect at point B. Only one primer can connect to any one piece of DNA. Step 3: 72°C for 30 seconds This steps allows for the Taq Polymerase to create new DNA using the dNTPs. This step is curtail to allow for DNA to be separated further. In essence, this step has created copies of the original DNA. Final step: 72°C for 3 minutes This step is similar to step 3, but is extended to allow for all of the Taq Polymerase to react. Final hold: 4°C This step isn't necessary, however, it is to prevent any water condensation from entering the solution filled with millions of copies of the target DNA strand.
dNTPs consist of A, T, G, and C. A will connect with T T will connect with A G will connect with C C will connect with G During PCR In step 4, Taq Polymerase will connect dNTPs to the DNA. If there is an A in the DNA, it will connect a T in that spot. If there is a G in the DNA, it will connect a C to it, etc. |
|||||||||