BISC110/F11: Lab 7 Taster SNP2
Lab 7 -Taster SNP Week 2
Today you will finish determining your genotype at the PTC gene locus by testing for one of the SNPs in the PTC gene. Remember that last week you began the process by isolating your own DNA and setting up a PCR reaction to amplify a region of the PTC gene which contains the SNP of interest. Today you will digest your PCR product with the restriction enzyme Fnu4H1 and analyze the products by agarose gel electrophoresis. We should also have time for a data analysis discussion and writing workshop in preparation for the paper you will write on your findings.
PART I: Digestion of your PCR products with the restriction enzyme Fnu4H1
Now that you have plenty of copies of the region of DNA we’re interested in, you will set up a restriction enzyme digest. How will this differentiate the SNP you have? Restriction enzymes are quite specific about the DNA sequences that they will cut. If the sequence is different, even by one base, they will not cleave it. We have made use of this specificity in choosing a restriction enzyme, Fnu4H1, which cuts one variant of the PTC gene SNP, but not the other. Because the restriction digest will take an hour, you will set it up at the beginning of the lab period.
- Locate your PCR tubes from last week, and set them out at your bench to thaw.
- Each student will set up a restriction enzyme digest of her own PCR product and one student of each pair will also digest the pcr product of the heterozygous(PAV/AVI) control.
- Get a clean microfuge tube and label it with your id number (from the course spreadsheet), and D to indicate that this will be the digested DNA. If you are doing the control, label another microfuge tube with a C and a D.
- Use your P20 set to 10 μl and carefully pipette 10 μL of the appropriate PCR product into the D labeled microfuge tube(s). Be sure to save the remaining PCR product! You will also run this undigested DNA from on your gel. Add the label U, for undigested, to the PCR tubes that contain the rest of PCR product (after you have removed the 10 μL aliquot for digestion).
- Add 10 μL of restriction enzyme “cocktail” to the D tube(s) only. The "cocktail" contains Fnu4H1 and an appropriate buffer. (In each 20 μL reaction volume there is 2 µl New England Biological Buffer #4 (http://www.neb.com), 2.5 units (in 0.5µl) of FNU4H1 (New England Biological product RO171), 7.5 µl purified water and the remainer is template DNA). Mix well by pipetting up and down.
- Incubate your digests in the 37C heat block for 60 minutes.
DO NOT THROW AWAY ANY OF THE PCR TUBES!
PART II: Agarose gel electrophoresis.
View the animation of how to pour and run an agarose gel.
You can do several of these steps while you are waiting on your restriction digests.
We will be using a Biorad gel system. Each pair of students will use one of these systems.
Setting up the gel apparatus
- Place the gel-casting tray in the gel box on the raised platform, positioning the comb slots nearest to the negative electrode (Black).
- Place the metal wedges into the gel box in the slots on either side of the gel-casting tray.
- Place the comb into the slots closest to the edge of the gel-casting tray.
- Ask your instructor to check your set up.
Dilute the buffer and make an agarose solution:
Each pair of students will make a 30 ml solution of 1.5% (w/v) agarose in 1X TBE (89mM Tris Base (TRIZMA), 89mM boric acid, 2mM EDTA (pH 8.0).
In order to do this:
- You will be provided with a 10X TBE stock solution. You will need a total of 300ml of buffer for preparing your agarose solution and for running your gel. Calculate the amount of 10X TBE you will need in order to prepare 300ml of 1X TBE.
- Measure out _____ ml of 10X TBE
- Add_____ml of water.
- Use a stir bar to mix this 1X TBE solution (briefly) in a 500ml flask.
- Calculate the amount of agarose you should weigh out to make your gel.
- For 30 ml 1.5% agarose weigh out _____ g of agarose and put that in ______ ml of the 1x TBE buffer that you diluted.
- Check your calculations with your lab instructor before proceeding.
- Using one of the balances in lab, carefully weigh out the appropriate amount of agarose using one of the top loading balances and a piece of weighing paper. Transfer the agarose to the small, empty 125 ml Erlenmeyer Flask at your bench and add the proper amount of 1X TBE buffer that you previously diluted. Swirl to mix. The agarose will NOT go into solution without high heat (the next step).
- Cover the open top of your 125ml flask with the little beaker you found on it originally and microwave the agarose solution for ~15-20 seconds on high power. Use the “hot hands” to handle the flask when removing it from the microwave, since the glass will get very hot. The agarose should be completely melted and the solution should be uniformly clear when you swirl it around. If it isn't, microwave it again for a few more seconds (~10).
- Immediately take your flask to the instructor's bench and ask your lab instructor to add 3 μl of SYBR Safe stain to your flask. PUT ON GLOVES. Any time you work with your agarose or gel for the remainder of the lab, you will need to be wearing gloves.
- Wearing gloves, swirl the flask gently to mix, taking care to avoid the formation of bubbles. If you sit the flask on the lab bench and make a slow figure eight, this form of mixing works pretty well. Then immediately proceed to the next step since everything is highly time sensitive at this point.
Cast a gel:
If you pour the agarose while it is too hot, you can crack or warp the casting tray, which may cause leaking, but if the solution is too cool it will be a lumpy mess when you pour it and you will have to start all over again.
- When the lip of the glass flask is not too hot to hold with a bare hand (approximately one minute after you add the Sybr green), you may begin to pour the agarose into the get-casting tray (if your instructor has previously checked it for correct assembly.
- Start out very SLOWLY pouring agarose into the CENTER of gel-casting tray, trying to avoid the formation of air bubbles. If you start in the center, by the time the agarose gets to the edges, it will be just cool enough to seal them without leaking. If you start out pouring VERY slowly, the casting tray is less likely to crack from the heat because the temperature can equilibrate incrementally. If bubbles form, remove them quickly before the gel starts to solidify by having your partner pop them with a Pasteur pipette.
- When you have poured the whole flask of molten agarose, allow it to set completely. When it has solidified, it will appear relatively opaque. Ask your instructor to check before you go on the next step.
- Carefully remove the metal wedges from the gel box.
- Fill both sides of the gel box with 1xTBE running buffer, until the buffer is about 3mm above the gel surface.
- To remove the comb from the gel, pull it slowly and straight up so that the wells are not damaged.
While you are waiting for your restriction digests to finish:
- Prepare your undigested samples: Each student will run their own undigested PCR product as well as their own digested product. You only have to run the digested control (heterozygous DNA).
- Label a microfuge tube with your code number and indicate that it contains undigested DNA (U).
- Pipette 10 μL of the appropriate PCR product from the PCR tube into the labeled microfuge tube.
- If you do not have 10 μL left, add as much of your PCR product as you have.
- Add 10 μL of water to each labeled microfuge tube. Mix thoroughly by pipetting up and down.
- Add 2.2 μL of loading dye to the microfuge tube. Mix thoroughly by pipetting up and down. Loading dye contains the following components:0.4 % Ficoll 400, 1.8 mM EDTA, 0.55 mM Tris-HCl, 0.00117 % SDS, 0.025 % Bromophenol Blue pH 8.0 @ 25°C)
- Don’t forget to use a clean, new tip for each tube.
- Your lab group should acquire a sample of marker lane DNA (ready to load) from your lab instructor. This marker lane DNA contains DNA fragments of the following sizes: 766 bp, 500 bp, 300 bp, 150 bp and 50 bp. Using what you know about the SNP region we amplified with PCR and the location of the Fnu4H1 II cutting site, you should be able to predict where you might see DNA fragments (looking like glowing bands) in relation to the position of the marker lane bands.
- Practice gel loading. Loading an agarose gel can be tricky. Your lab instructor will demonstrate how to use the practice dishes to learn to load a gel properly. Practice until you are comfortable and proficient.
Bromophenol blue: is a negatively charged dye that moves towards the positive pole.
Ficoll: is added to increase the density of the sample. This causes the sample to sink into the well, since it is heavier than the liquid in the well.
EDTA: helps keep the nucleic acid in the sample stable by inhibiting nucleases that might be present. It does this by binding divalent cations, such as Mg2+, that are necessary for nuclease activity.
Do not throw the stock tube of loading dye away when you clean up after this lab!
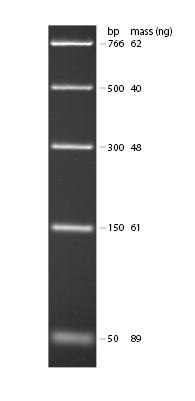
DNA Ladders: 0.3µg of “PCR Ladder” and visualized by ethidium bromide staining on 1.8% TBE agarose gel. Ladder from New England Biological (NEB3234).
Prepare your restriction digest for loading:
- After your restriction enzyme digest is complete, microfuge your tubes briefly (5-10 seconds) to get any condensation from the lid into the bottom of the tube.
- Add 2.2 μL of loading dye to each tube of digested DNA (one tube for each member of your lab group, plus the heterozygous digested control).
- Each pair of students should now have 6 tubes which contain DNA and loading dye: one digested tube for each group member, one undigested tube for each group member, one digested control, and one tube of marker lane DNA.
Loading and Running the Gel
- Fill out the gel template provided so you know which lane each sample will be loaded. Copy this template into your lab notebook. You will load 20 microliters from each tube (including the pcr marker) into separate gel lanes.
- Check your tubes before loading: microfuge briefly (around 15 seconds) if necessary. Be careful not to let any bubbles into the pipette tip when you are drawing the sample into the tip. Give the template to your lab instructor.
- When all of the samples have been loaded, check the orientation of the gel with respect to the electrical leads.Remember “RED AHEAD”; DNA, having a net negative charge, will migrate toward the positive (red) electrode.
- Placing the lid on the box connects the leads to the gel box. Connect the other ends of the leads to the power supply. Check that the power supply is set at 100 volts, and then turn on the power. Observe bubbles rising from the electrodes at each end of the tank (more from the negative electrode than the positive electrode). After 1–2 min, observe the dye leaving the wells. Run the gels for approximately 30-35 minutes or until the tracking dye has moved about half way down the gel. Ask your instructor to check that your dye front has migrated sufficiently before you stop the electrophoresis by turning off the current.
Gel photography
- When the gel has separated your DNA fractions, disconnect the leads from the power supply and remove the lid from the gel rig.
- Wearing gloves, carefully remove your casting tray (still containing the gel) from the gel rig and transfer it to a labeled plastic sandwich box at your bench.
- After the electrophoresis is complete, your instructor will take you to the instrument room next door, so that you may take pictures of your gels. Follow the directions in Appendix G to use one of the imagers to visualize your DNA fragments and to take a photo.
- SYBR SafeTM stain is a fluorescent dye that binds to nucleic acids. It has excitation maxima at 280nm and 502nm, and an emission maximum at 530nm. You will place your gel on a UV transilluminator, a device that shines ultraviolet light through the gel. Where nucleic acids have been stained with the SYBR SafeTM stain, a fluorescent green glow will be detected by the digital camera connected to the transilluminator.
- Once you have captured a satisfactory image of your gel, email it to yourself, and upload it to your lab Sakai conference.
Analyzing the Gel
- Ascertain whether your enzymes cut your DNA. How can you tell? How many fragments do you see for each person in your lab group?
- Use the DNA ladder to estimate the size of the fragments you see.
Recording Your Data
- Determine what genotype you are: PAV/PAV, PAV/AVI, or AVI/AVI. Be sure to check your interpretation of the gel with your lab instructor.
- Once you are sure of your genotype, record your data in the class excel spread sheet where you have already recorded your PTC sensitivity phenotype. DO NOT LEAVE LAB UNTIL YOU HAVE RECORDED YOUR GENOTYPE ON THE COURSE SPREADSHEET AT THE INSTRUCTOR COMPUTER!
Class results will be compiled and presented to protect anonymity. You are not required to share your personal data with anyone else in lab (though you are free to do so if you wish).
Part III. Data Analysis & Science Writing Discussion
While your restriction enzyme digestion and electrophoresis are running, we will have time to talk about your expected results and their significance. We will also address how you might write about these data in the form of a primary research report due at the beginning of the next lab. This will be a full paper (except that you may omit the Materials and Methods section) so we will concentrate on the sections you have yet to write (Abstract and Introduction) using the Wooding article as a possible model. You should have brought a hard copy of this article with you to lab: Wooding et al., Natural Selection and Molecular Evolution in PTC, a Bitter-Taste Receptor Gene. Am. J. Hum. Genet. 74:637–646, 2004.
Assignment
Your instructor will post the preliminary course data to a Data Folder in Resources in your lab Sakai site. It is time for you to start analyzing the data to discover what it says about your main experimental question, "What is the distribution of the Taster PTC phenotype and genotype at Wellesley College?" What are you going to do with that table of phenotype and partial genotype information? At the moment it is just a list of code names and raw data showing who could taste, partially taste or not taste the PTC paper and whether or not those subjects' DNA showed a cytosine or a thymine nucleotide at SNP #785 on each of their two copies of human gene TAS2R38. The first thing to do is to get that raw data table into sortable form in Excel. If your instructor hasn't already done this for you, remove the lab section identifier information so that you have long uninterrupted columns of numbers. For how many subjects do you have complete data sets (n=?). What should you do with the subjects for whom you have only partial data? Does it matter if you lack phenotype or genotype information for some subjects?
There are many possible ways to sort and use these data to answer different parts of our basic question. Before you come to lab next week, use this preliminary COURSE data (not just your lab section) to create a graph that shows the PTC Taster phenotype distribution of Wellesley College students. Make another graph that shows the genotype distribution. Create a different graph of Allele frequencies: PAV vs. AVI. Make an additional graph illustrating how well you were able to predict phenotype from genotype in this population. Turn your graphs into figures with legends. Remember that your legend must allow the reader to understand how the data shown were collected and processed into the form shown in the graphs and it must clarify ambiguous terms. Remember that your reader is NOT likely to be familiar with acronyms like PAV, AVI, SNP, PTC, etc. or understand how you were able to determine this partial genotype for the PTC gene. The legend is independent of the results narrative; this means that your reader must be able to understand the data and the take home message without reading the narrative. Conversely, the narrative must do more than point to the data shown in the figure. The figure title should be the main take-away message OR summarize what's in the graph if a conclusion isn't appropriate.
Remember that although your basic experimental question is, "what is the distribution of the Taster PTC phenotype and genotype at Wellesley College?", you also want to apply the answer to that question to a broader concept in the discussion section of your paper. Consider the following broader topics: Is Wellesley College unusually diverse and Do your findings support Fisher's Balancing Selection Theory for the PTC taster gene? Which of the graphs you have created help you answer those questions? Review the paper by Wooding et al., Natural Selection and Molecular Evolution in PTC, a Bitter-Taste Receptor Gene. Am. J. Hum. Genet. 74:637–646, 2004. | http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1181941/?tool=pubmed and Wooding et al. (2006)review and some of the the Kim and Drayna papers on their work in this field and consider what non-agreement between the prediction of phenotype from genotype might mean. Can you now think of ways to revise or expand your graphs or make new ones or combine them into composite figures of more than one graph to show a new reader how your data might be applied to those concepts? If you can, do so and bring both the required graphs and these revisions to lab next week to use in our science writing workshop part of lab.
Construct a rough draft of your results section, please start a separate paragraph describing each of your draft figures. Each paragraph should begin, "In order to find out_______". Complete that sentence with whatever part of the experimental goals are answered in the figure. The second sentence in each results paragraph should be a summary of what you did experimentally to obtain the data shown in the figure it will analyze. DO NOT include procedural details but instead, start with the beginning material (paper impregnated with PTC or cheek cells from Wellesley College students) and describe, in as few words as possible, how that material was processed into whatever is shown in your graph. If the reader is unlikely to be familiar with a term, like FNU4H1, you should either define it or use a more general descriptor instead of the term, such as replacing FNU4H1 with "digested with a restriction enzyme that cuts the DNA of the most common taster allele but not the most common non-taster allele". Once you have given the context, you will be ready to point out the most relevant information in the figure that answers that question addressed in the goal. List the most important numbers in your figure or the most important comparison. Write a few sentences for each graph that explains how or why a particular comparison or the numbers you have listed as important show something interesting about the genotype or phenotype distribution, the allele frequency, or how well you were able to predict phenotype from genotype. Do those sentences answer the goal described in the first sentence of each paragraph? If not, clarify and construct a conclusion sentence using the data you described in the previous sentences.
Use the Structure of Science Writing handout (available for download at Media:Structure_of_Science_Writing110_1.doc ) to construct a skeleton outline of your paper's introduction and discussion sections. A skeleton outline includes all the topic sentences for each paragraph in each section. Please make sure that your topic sentences address each of the expected elements for that section (in the order they are described in the handout). Include in the skeleton outline of each paragraph (after the topic sentence) the full citation (in Cell format) for outside information and a brief summary of that relevant outside information. In the outline of the discussion section, please include the figure number of the evidence from your study that will be used in that paragraph and a summary and citation for outside evidence.
Bring a hard copy of all of these draft materials to lab next time. Print them single sided, one figure per page (with legend), and skip several lines between each topic sentence in your skeleton outlines. If time permits, we will workshop some of the figures and dissect the skeleton outlines in collaborative groups. Make sure you have uploaded your drafts to your lab Sakai site before lab.