BE.109:DNA engineering/FACS analysis
Introduction
FACS stands for "Fluorescence Activated Cell Sorting." The FACS machine has revolutionized biology by allowing researchers to isolate cells based on their spectral qualities. For example, if you have a fluorescently tagged antibody that binds to a certain cell type, you can isolate a pure sample of this cell type from a complex mixture by using a FACS machine. In addition to purification, the FACS machine can count the number of cells that have a certain spectral quality. If a FACS machine is used just for counting and not for separating subpopulations of cells, then the procedure is called "flow cytometry," and this is what you will be doing today.
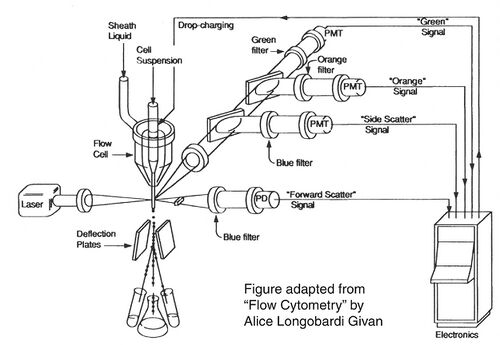
Before there were FACS machines, there were Coulter counters. Coulter counters are automated cell counting machines developed in the 1950s that count cells as they flow in a liquid stream. In an ingenious conceptual leap, Mack Fulwyler combined the technology of ink jet printers with that of Coulter counters to develop the first FACS machine. The ink jet printer head works by vibrating a nozzle so that a spray of discrete droplets is formed. Similarly, in a FACS machine, a liquid suspension of cells is forced at high pressure through a vibrating nozzle to create tiny charged droplets, each containing a single cell. The stream of droplets pass in front of a laser beam, and the scattered light is analyzed by a series of filters and photomultiplier tubes that convert the light signal into electrical impulses. Thus, each cell is "interrogated."
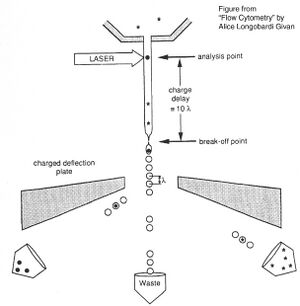
For FACS, the spectral qualities of the cell are analyzed nearly instantaneously and the machine "decides" whether or not the cell matches your desired spectral qualities. For example, if you have a mixture of green fluorescent cells and non-fluorescent cells, you can ask the machine to isolate the green cells. If a cell registers as green, an electrical charge deflects the cell to make it fall into a collection chamber.
FACS is technically challenging and most FACS machines are only run by experts. In contrast, biologists are often trained to perform flow cytometry in order to analyze the proportion of their sample that has particular spectral qualities. You will be using flow cytometry to measure the percentage of cells that are fluorescent. You have lipofected cells with two non-functional EGFP genes. Recombination between these two genes can restore the full length EGFP coding sequence so that cells express EGFP. By measuring the percentage of cells that fluoresce green, you will have some measure of the frequency of homologous recombination within mammalian cells.
Protocol
Preparing your cells for FACS analysis The following protocol should be performed in the sterile hoods unless otherwise indicated.
- Aspirate the media from your cells and wash them with 1 ml PBS, aliquotted with a 10 ml pipet.
- Add 200 ul trypsin to each well, aliquotted with a 2 ml pipet. After the last addition, start a 1’ timer. During this time, rock the plate in each direction to distribute the trypsin over the cells.
- Aspirate the trypsin and incubate the cells at 37°C for 10 minutes, using your timer to precisely time this incubation.
- Resuspend the cells in 100 ul OptiMEM, using your P200 to make an even suspension. Move each sample to a labeled FACS tube
- Keep your tubes on ice as you walk to the FACS facility.
For next time
Write a lab report describing the experiment you have performed. You should include information about building the Δ5 plasmid, an essential component created for your ultimate investigation, but you should focus your report on the recombination experiment you performed. As you sit down to write, it might be wise to reread the information about lab reports that is included as supplementary material with the lab manual and here: BE.109:Guidelines for writing a lab report.
