Amplified insert assembly
Overview
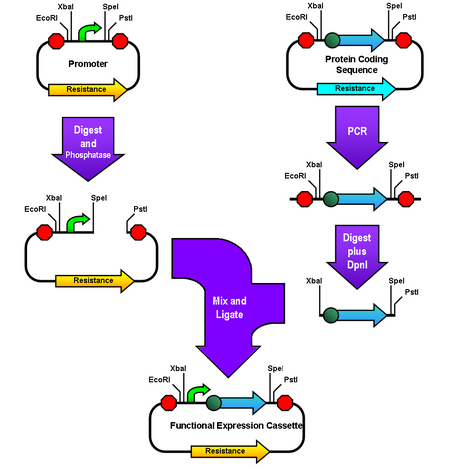
Amplified Insert Assembly is a method of "BioBricking™" two biological parts (i.e. pieces of DNA) together. For more information on BioBricking see this link. A You-Tube video summary of the Amplified Insert Assembly method provides a quick visual introduction.
This method combines the ease and flexibility of 3A assembly with the reliability of standard assembly. In comparison to 3A assembly, this method can take up to two hours longer; however, the additional time spent at the bench is minimal. Major benefits of this assembly method over other BioBrick™ assembly methods include:
- No need for gel electrophoresis or gel extraction.
- The ability to insert small (i.e. invisible on a gel) parts.
- No need to use multiple antibiotic resistances.
- No having to make construction vectors.
- No need to order custom oligos for each assembly.
- Really low background (99% of colonies are correct)
- This means less sequencing
- Easy transformation (use homemade competent cells)
- Less culturing
- This is because one plasmid prep can supply many PCR inserts. So minipreps of common parts (i.e. promoters and RBSs) can be used over and over again.
This protocol is typically used to do BioBrick™ assembly with restriction sites in the RFC 10 BioBrick Standard :
-----EcoRI--XbaI--Part--SpeI--PstI-----
But it can also be applied to any other format in which the two inner sites form an inactive "scar" and the two outer sites can be heat inactivated.
The two parts you want to assemble will be labeled "insert" and "vector" and will be initially contained on separate plasmids. The eventual goal of assembly is to get these parts on the same plasmid next to one another.
This is a consensus protocol. For more information see the pages below
Materials
General
- Pipettors
- Microcentrifuge
- Water baths
- Thermocycler
- Electroporator
- Selective media plates
Enzymes
- EcoRI Restriction Endonuclease
- XbaI Restriction Endonuclease
- SpeI Restriction Endonuclease
- PstI Restriction Endonuclease
- DpnI Restriction Endonuclease
- Vent DNA Polymerase (or another equivalent high fidelity polymerase)
- Antarctic Phosphatase
- T4 Ligase
Procedure
1. Miniprep both "insert" and "vector" from their respective cultures using a kit or this protocol (30 mins).
2. PCR the "insert" plasmid.
- Use a high-fidelity polymerase (e.g. pfu Turbo or Vent).
- Use the same primers you use for Colony PCR.
- These should flank your restriction sites by 100-150bp thus allowing even very small parts (e.g. RBS) to be purified using a column.
- They should probably have a Tm of 55-60°C.
- For most BioBrick applications these can be primers VF2 and VR.
- Only run 25-30 cycles as this will help ensure high fidelity.
- This will take 1-2 hrs, but start the vector digest right away while the PCR is cycling.
- Use a high-fidelity polymerase (e.g. pfu Turbo or Vent).
3. Digest the "vector" with the appropriate restriction endonucleases for 2 hours.(do this while while PCR is running)
4. Purify the PCR product using a kit or this protocol.
5. Digest the purified insert for 1 hour with enzymes complementary to your vector digest.
- Include DpnI along with the other restriction endonucleases.
- Include DpnI along with the other restriction endonucleases.
6. Add 6μL Antarctic Phosophatase Buffer and 1μL Antarctic Phosphatase to the "vector" digest and incubate until the "insert" digest is done.
7. Kill all reactions by incubating for 20 mins at 80°C.
8. Ligate at a molar ratio of 4:1 (insert:vector).
9. Transform your cells.
10. Plate the transformed cells on plates with the same antibiotic as the "vector" resistance.
11. Celebrate.
- If you already have PCR insert ready to go (i.e. you ran the PCR the night before from old miniprep) then it only takes about 4 hours.
Notes
- The DpnI eliminates any background from the insert PCR.
- The phosphatase eliminates any background vector.
- The "vector" will be digested for a total of thee hours (including nearly one hour with Antarctic Phosphatase)
- The "insert" will only be digested for one hour. This is okay as there is a lot of it.
- Detractors of this method may say that it's risky to PCR the inserts because of mutations. We say:
- This hasn't been a problem for us.
- This is why we use a high-fidelity polymerase
- We're sequencing the constructs anyway so we'd spot any mutations.
References
- Speer, M.A. and T.L. Richard. 2011. Amplified insert assembly: an optimized approach to standard assembly of BioBrick™ genetic circuits Journal of Biological Engineering 5:17
Contact
- This protocol was developed by Mike Speer and Tom Richard
- You can also discuss this protocol here.