OpenSourceMalaria:GSK Arylpyrrole Series: Difference between revisions
| (32 intermediate revisions by 3 users not shown) | |||
| Line 3: | Line 3: | ||
== The Arylpyrrole Series == | == The Arylpyrrole Series == | ||
This page contains a summary of all the data associated with this open source project. A more [[OSDDMalaria:GSK_Arylpyrrole_Series:Story_so_far|'''human-readable summary''']] is available. | |||
[[ | |||
These compounds are currently under study by the Todd lab at the University of Sydney and the Medicines for Malaria Venture, Geneva. As an open source project, anyone may participate:<br> | These compounds are currently under study by the Todd lab at the University of Sydney and the Medicines for Malaria Venture, Geneva. As an open source project, anyone may participate:<br> | ||
[http://www.thesynapticleap.org/node/342 Coordination/Discussion site]<br> | [http://www.thesynapticleap.org/node/342 Coordination/Discussion site]<br> | ||
[http://malaria.ourexperiment.org/ | [http://malaria.ourexperiment.org/ Electronic Lab Notebooks]<br> | ||
[[Todd:Synthesis_of_Analogs_of_Arylpyrrole_Antimalarial_Drug_Leads|Undergraduate Project Report]] | |||
The initial two subseries of leads have the general structure: | |||
* TCMDC Aryl pyrrole core InChi Key: LNROIXNEIZSESG-UHFFFAOYSA-N | |||
* Near-Neighbour core InChi Key: JZQOEGKHCXONAV-ZROIWOOFSA-N | |||
[[Image:Arylpyrrole Generic.png|thumb|right|120px|GSK Tres Cantos Arylpyrrole Generic Structure]] | |||
[[Image:Nearneighbour generic.png|thumb|right|120px|Near Neighbour Generic Structure]] | |||
A number of these compounds and intermediates are [[OSDDMalaria:GSK_Arylpyrrole_Series:available_compounds|available]] for biological testing from Todd group by request. If you would like to contribute new analogues, please check the list of [[OSDDMalaria:GSK_Arylpyrrole_Series:desired_compounds|desired]] compounds. | |||
=== The two known TCMDC series starting points === | |||
Compounds are [http://www.thesynapticleap.org/node/345 commercially-available]<br> | Compounds are [http://www.thesynapticleap.org/node/345 commercially-available]<br> | ||
Series was released in a [http://www.nature.com/nature/journal/v465/n7296/full/nature09107.html large dataset] by GSK and later [http://pubs.acs.org/doi/abs/10.1021/ml200135p listed as one of the most promising leads] from TCAMS set (though lacking aryl F)<br> | |||
==== TCMDC 123812 ==== | ==== TCMDC 123812 ==== | ||
| Line 42: | Line 51: | ||
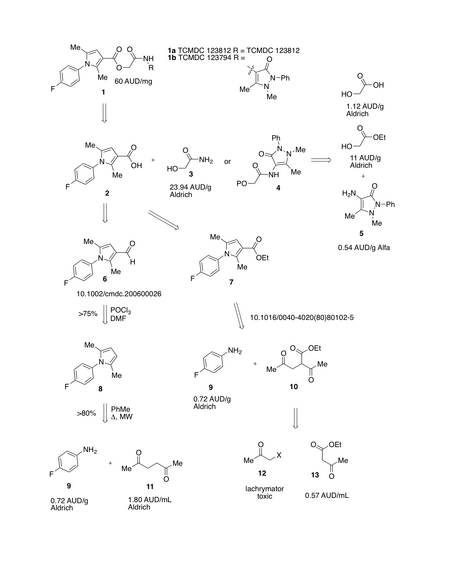
[[Image:arylpyrrolesynth.png|thumb|450px|center|Proposed Synthesis Strategy]] | [[Image:arylpyrrolesynth.png|thumb|450px|center|Proposed Synthesis Strategy]] | ||
=== Known "[[OSDDMalaria:GSK_Arylpyrrole_Series:Near Neighbours|Near Neighbours]]" contained in the Tres Cantos set === | === The Near-Neighbour Sub-series === | ||
==== Known "[[OSDDMalaria:GSK_Arylpyrrole_Series:Near Neighbours|Near Neighbours]]" contained in the Tres Cantos set ==== | |||
The original idea for investigation of the near-neighbour set came from Paul Willis and was posted on [http://www.thesynapticleap.org/node/349 the synaptic leap]. | |||
[[Image:Near neighbours.png|thumb|450px|center|Known GSK Arylpyrrole Near Neighbours]]<br> | [[Image:Near neighbours.png|thumb|450px|center|Known GSK Arylpyrrole Near Neighbours]]<br> | ||
| Line 66: | Line 79: | ||
CCn1c(cc(c1C)C=C2C(=O)NC(=Nc3ccccc3)S2)C | CCn1c(cc(c1C)C=C2C(=O)NC(=Nc3ccccc3)S2)C | ||
==== | ==== Initial synthesis strategy toward near-neighbours ==== | ||
[[Image:Nearneighbour.png|thumb|center|450px| | [[Image:Nearneighbour.png|thumb|center|450px|Initial synthesis of near neighbours]] | ||
Experimental information available from [http://malaria.ourexperiment.org/uri/32 PMY 13-1], [http://malaria.ourexperiment.org/uri/33 PMY 14-1] and [http://malaria.ourexperiment.org/uri/39 PMY 16-1], [http://malaria.ourexperiment.org/uri/41 PMY 14-3]. | Experimental information available from [http://malaria.ourexperiment.org/uri/32 PMY 13-1], [http://malaria.ourexperiment.org/uri/33 PMY 14-1] and [http://malaria.ourexperiment.org/uri/39 PMY 16-1], [http://malaria.ourexperiment.org/uri/41 PMY 14-3]. | ||
==== Current synthesis strategy toward near-neighbours ==== | |||
[[Image:Nearneighbour2.png|thumb|center|450px|Current synthesis of near neighbours]] | |||
Synthesis method taken from [http://dx.doi.org/10.1016/j.bmcl.2011.09.049 this paper]. | |||
=== Alternative side-chains === | |||
Some alternative heterocyclic side-chain ideas have been proposed [http://www.thesynapticleap.org/node/367 here] at the Synaptic Leap. | |||
==== Oxazoles ==== | |||
The [http://en.wikipedia.org/wiki/Oxazole|oxazole] side chain would target ester isosteres (see TCMDC-123812) but with greater biological stability. | |||
*Synthesis: [http://dx.doi.org/10.1021/ol101145t oxazole synthesis] | |||
*Pyrrole core acid to amide: [http://dx.doi.org/10.1021/jm100504d Amide Synthesis] | |||
=== Other known incidences of these molecules/this series === | === Other known incidences of these molecules/this series === | ||
Related compounds are known to inhibit the proteasome, according to [http://www.nature.com/nature/journal/v467/n7312/abs/nature09299.html this Nature paper]. | Related compounds are known to inhibit the proteasome, according to [http://www.nature.com/nature/journal/v467/n7312/abs/nature09299.html this Nature paper]. | ||
According to [http://dx.doi.org/10.1016/j.bmcl.2011.09.049 this paper], similar compounds are agonists of the sphingosine-1-phosphate receptor subtypes 1-5, which have a role in immune system function. | |||
A new class of antimalarial was recently published by Novartis, [http://dx.doi.org/10.1021/jm2003359 Imidazolopiperazines]. | |||
Misc papers to digest/assimilate regarding antimycobacterial/tuberculosis activity: [http://dx.doi.org10.1021/jm701560p Antimycobacterial 1,5-diphenyl pyrroles], [http://dx.doi.org/10.1016/j.ejmech.2009.06.005 10.1016/j.ejmech.2009.06.005], [http://dx.doi.org/10.1016/j.bmc.2010.09.006 10.1016/j.bmc.2010.09.006], [http://dx.doi.org/10.1002/cmdc.201000526 10.1002/cmdc.201000526]<br> | |||
''' ChEMBL-NTD ''' | |||
Searches on [https://www.ebi.ac.uk/chemblntd/compound/structure_home ChEMBL-NTD] reveal: | |||
* 19 iminothiazolidinones | |||
* 55 2,5-dialkylpyrroles | |||
* 58 rhodanines. These do not contain any biological data, presumably omitted due to promiscuity/PAINS issues. | |||
'''Scifinder Search:''' | '''Scifinder Search:''' | ||
| Line 91: | Line 134: | ||
[[Image:Proteasome.png|thumb|none|Proteasome agonist]] | [[Image:Proteasome.png|thumb|none|Proteasome agonist]] | ||
[http://www.wipo.int/patentscope/search/en/WO2006076202 WO2006076202] Steroid nuclear receptor ligands | |||
[http://worldwide.espacenet.com/publicationDetails/description?CC=WO&NR=2011075684A1&KC=A1&FT=D&date=20110623&DB=EPODOC&locale=en_EP WO 2011075684 (A1)] Inhibitors of Plasma [http://en.wikipedia.org/wiki/Kallikrein Kallikrein]/Protease | |||
[http://worldwide.espacenet.com/publicationDetails/biblio?FT=D&date=20091112&DB=EPODOC&locale=en_EP&CC=WO&NR=2009137133A2&KC=A2&ND=5 WO 2009137133 (A2)] 5-Substituted-2-Imino-Thiazolidinone Compounds And Their Use As Inhibitors Of Bacterial Infection | |||
[[Image:WO2009137133A2.png|thumb|none|Bacterial inhibitor]] | |||
== Biology == | |||
=== Malaria Assays === | |||
==== '''''In vitro''''' ==== | |||
Current tested compounds and their biological activities are summarised on a Google [http://bit.ly/OSDDcompounds spreadsheet] | |||
The results are collected on [http://malaria.ourexperiment.org/biological_data malaria.ourexperiment.org]. | |||
[http://www. | ==== '''''In vivo''''' ==== | ||
TCMDC-123812, TCMDC-123794 and near neighbour ZYH 3-1 have been [http://www.thesynapticleap.org/node/395 submitted] for study oral ''in vivo'' mouse model. Unfortunately it was found that the compounds possess zero oral efficacy (Results available [http://malaria.ourexperiment.org/uri/ed here]) in this initial trial. | |||
=== Mode of action === | |||
==== ''In silico'' prediction ==== | |||
''In silico'' prediction of targets has been carried out by [https://plus.google.com/u/0/b/114702323662314783325/115238763121035691959/posts Iain Wallace] and summarised by [[User:Matthew_Todd|Mat]] on [http://www.thesynapticleap.org/node/387 the synaptic leap]. Detail on the procedure is given on the [http://malaria.ourexperiment.org/in_silico_prediction/2675 ELN]. The following targets were identified from data derived from the ChEMBL database: | |||
* Carboxy-terminal domain RNA polymerase II polypeptide A small phosphatase 1 | |||
* Dihydroorotate dehydrogenase (DHODH)[in progress at GSK Tres Cantos] | |||
* SUMO-activating enzyme subunit 2 | |||
* SUMO-activating enzyme subunit 1 | |||
* Cyclin-dependent kinase 1 | |||
Collaborators are invited to investigate these targets to confirm if these are targeted by our compounds. | |||
==== ''In vitro'' ==== | |||
GSK Tres Cantos have offered to assay our compounds against dihydroorotate dehydrogenase (DHODH). | |||
Revision as of 04:29, 6 June 2012
The Arylpyrrole Series
This page contains a summary of all the data associated with this open source project. A more human-readable summary is available.
These compounds are currently under study by the Todd lab at the University of Sydney and the Medicines for Malaria Venture, Geneva. As an open source project, anyone may participate:
Coordination/Discussion site
Electronic Lab Notebooks
Undergraduate Project Report
The initial two subseries of leads have the general structure:
- TCMDC Aryl pyrrole core InChi Key: LNROIXNEIZSESG-UHFFFAOYSA-N
- Near-Neighbour core InChi Key: JZQOEGKHCXONAV-ZROIWOOFSA-N

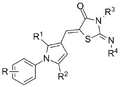
A number of these compounds and intermediates are available for biological testing from Todd group by request. If you would like to contribute new analogues, please check the list of desired compounds.
The two known TCMDC series starting points
Compounds are commercially-available
Series was released in a large dataset by GSK and later listed as one of the most promising leads from TCAMS set (though lacking aryl F)
TCMDC 123812
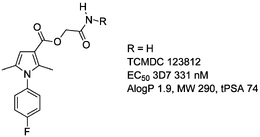
InChI=1/C15H15FN2O3/c1-9-7-13(15(20)21-8-14(17)19)10(2)18(9)12-5-3-11(16)4-6-12/h3-7H,8H2,1-2H3,(H2,17,19)
SMILES: O=C(N)COC(=O)c2c(n(c1ccc(F)cc1)c(c2)C)C
CAS Registry Number: 733026-12-5
Chemspider page
ChEMBL Page
Resynthesis:
TCMDC 123794
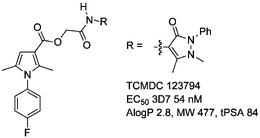
InChI=1/C26H25FN4O4/c1-16-14-22(17(2)30(16)20-12-10-19(27)11-13-20)26(34)35-15-23(32)28-24-18(3)29(4)31(25(24)33)21-8-6-5-7-9-21/h5-14H,15H2,1-4H3,(H,28,32)
SMILES: Fc1ccc(cc1)n2c(cc(c2C)C(=O)OCC(=O)NC=4C(=O)N(c3ccccc3)N(C=4C)C)C
Chemspider page
ChEMBL Page
Resynthesis:
The proposed resynthesis strategy for these two compounds:
The Near-Neighbour Sub-series
Known "Near Neighbours" contained in the Tres Cantos set
The original idea for investigation of the near-neighbour set came from Paul Willis and was posted on the synaptic leap.
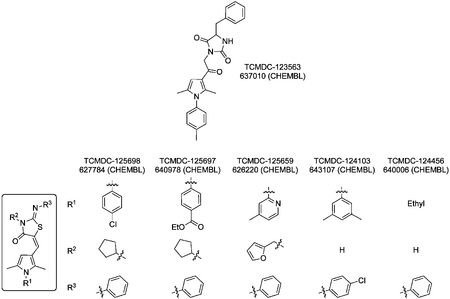
Data/links for these compounds:
TCMDC-123563, CHEMBL546966, CHEMBL page: 637010 Cc1ccc(cc1)n2c(cc(c2C)C(=O)CN3C(=O)C(NC3=O)Cc4ccccc4)C
TCMDC-125698, CHEMBL587989, CHEMBL: 627784 Cc1cc(c(n1c2ccc(cc2)Cl)C)C=C3C(=O)N(C(=Nc4ccccc4)S3)C5CCCC5
TCMDC-125697, CHEMBL581336, CHEMBL: 640978 CCOC(=O)c1ccc(cc1)n2c(cc(c2C)C=C3C(=O)N(C(=Nc4ccccc4)S3)C5CCCC5)C
TCMDC-125659, CHEMBL528140, CHEMBL: 626220 Cc1ccnc(c1)n2c(cc(c2C)C=C3C(=O)N(C(=Nc4ccccc4)S3)Cc5ccco5)C
TCMDC-124103, CHEMBL588465, CHEMBL: 643107 Cc1cc(cc(c1)n2c(cc(c2C)C=C3C(=O)NC(=Nc4ccc(cc4)Cl)S3)C)C
TCMDC-124456, CHEMBL548395, CHEMBL: 640006 CCn1c(cc(c1C)C=C2C(=O)NC(=Nc3ccccc3)S2)C
Initial synthesis strategy toward near-neighbours
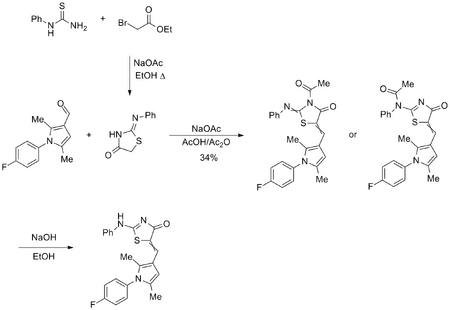
Experimental information available from PMY 13-1, PMY 14-1 and PMY 16-1, PMY 14-3.
Current synthesis strategy toward near-neighbours
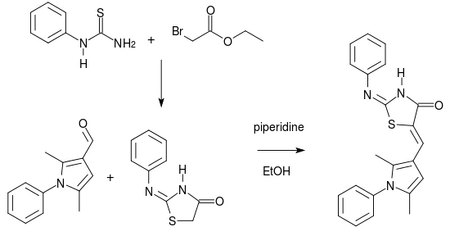
Synthesis method taken from this paper.
Alternative side-chains
Some alternative heterocyclic side-chain ideas have been proposed here at the Synaptic Leap.
Oxazoles
The [1] side chain would target ester isosteres (see TCMDC-123812) but with greater biological stability.
- Synthesis: oxazole synthesis
- Pyrrole core acid to amide: Amide Synthesis
Other known incidences of these molecules/this series
Related compounds are known to inhibit the proteasome, according to this Nature paper.
According to this paper, similar compounds are agonists of the sphingosine-1-phosphate receptor subtypes 1-5, which have a role in immune system function.
A new class of antimalarial was recently published by Novartis, Imidazolopiperazines.
Misc papers to digest/assimilate regarding antimycobacterial/tuberculosis activity: Antimycobacterial 1,5-diphenyl pyrroles, 10.1016/j.ejmech.2009.06.005, 10.1016/j.bmc.2010.09.006, 10.1002/cmdc.201000526
ChEMBL-NTD
Searches on ChEMBL-NTD reveal:
- 19 iminothiazolidinones
- 55 2,5-dialkylpyrroles
- 58 rhodanines. These do not contain any biological data, presumably omitted due to promiscuity/PAINS issues.
Scifinder Search: 2,4,5-alkyl-1-aryl-pyrrole: 77552 hits, 905 biological studies:
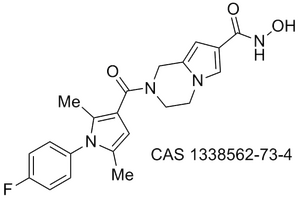
Obesity/diabetes GIP receptor inhibitor JP 2011184298 (A)
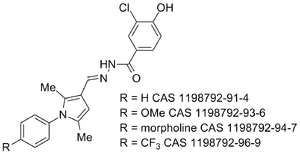
Proteasome WO 2011094545 (A2)
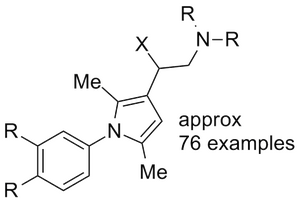
WO2006076202 Steroid nuclear receptor ligands
WO 2011075684 (A1) Inhibitors of Plasma Kallikrein/Protease
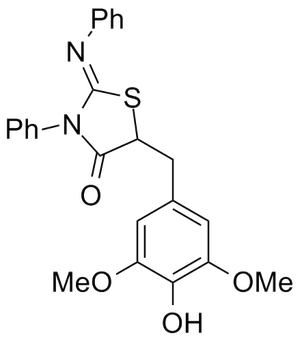
WO 2009137133 (A2) 5-Substituted-2-Imino-Thiazolidinone Compounds And Their Use As Inhibitors Of Bacterial Infection
Biology
Malaria Assays
In vitro
Current tested compounds and their biological activities are summarised on a Google spreadsheet
The results are collected on malaria.ourexperiment.org.
In vivo
TCMDC-123812, TCMDC-123794 and near neighbour ZYH 3-1 have been submitted for study oral in vivo mouse model. Unfortunately it was found that the compounds possess zero oral efficacy (Results available here) in this initial trial.
Mode of action
In silico prediction
In silico prediction of targets has been carried out by Iain Wallace and summarised by Mat on the synaptic leap. Detail on the procedure is given on the ELN. The following targets were identified from data derived from the ChEMBL database:
- Carboxy-terminal domain RNA polymerase II polypeptide A small phosphatase 1
- Dihydroorotate dehydrogenase (DHODH)[in progress at GSK Tres Cantos]
- SUMO-activating enzyme subunit 2
- SUMO-activating enzyme subunit 1
- Cyclin-dependent kinase 1
Collaborators are invited to investigate these targets to confirm if these are targeted by our compounds.
In vitro
GSK Tres Cantos have offered to assay our compounds against dihydroorotate dehydrogenase (DHODH).
