20.109(S15):Cell preparation for DNA repair assays (Day4)
Introduction
Today's the big day! The first of four pretty big days, actually: transfecting your cells, measuring their fluorescence, and analyzing relative DNA repair rates for different cell populations.
DNA repair inhibitors -- let's find one that works in CHO cells!
In Spring 2014 your 20.109 peers utilized C401, a chemical compound that inhibits the kinase domain of DNA-PKcs (cs = catalytic subunit, aka kinase domain) to block the NHEJ repair pathway. The goal of this was two-fold; 1) provide a 'back up' method to determine how much of the BFP repair that you measure is due to NHEJ (versus homologous recombination -- the other DSB repair pathway). Recall that we will utilize the xrs6 cell line as a control because it cannot repair DNA via the NHEJ pathway. But, as was discussed in lecture and lab, the Ku80 silencing mechanism in xrs6 cells is methylation dependent so it doesn't hurt to have a back-up method to perturb the NHEJ pathway. And 2) chemical inhibitors are used widely in biological engineering, so it is useful to think about their strengths and limitations.
Well...we found a major limitation of C401 when used in with our CHO cells -- it didn't work! Agi, a former 20.109 instructor, screened through a few more DNA-PKcs-specific inhibitors that had been used in human cells with success, but they did not work in our CHO cell system either. Therefore, we are calling on you -- the Spring 2015 20.109 superstars -- to help us "screen" four more inhibitors to determine if we can efficiently inhibit NHEJ in hamsters. There are four more NHEJ inhibitors that we will test in triplicate (+ 1 because there are 13 teams). The inhibitors were chosen based on searching the literature for inhibitors that may have been employed with CHO cells in the past + a paper by Goglia et al. that was recently published (and you'll read for M2D6) outlining a high throughput screening method for identifying NHEJ inhibitors.
After perusing these inhibitors, please sign up on the M2D4 talk page for what drug you'd like to try. You will use the drug for two experiments, the flow cytometry-based DNA repair assay, and an irradiation-induced cell-death experiment. The later, a 'kill curve', will utilize increasing doses of the drug and 4 Gy of irradiation to induce DNA damage. If the drug is effective, the DNA damage will not be repaired efficiently and the cells will die.
| Drug | Mechanism of action | Vendor website | Literature reference | Fun fact |
|---|---|---|---|---|
| Mibefradil dyhydrochloride | Unknown (for NHEJ) | Tocris | Goglia et al. | Used clinically to treat angina |
| Loperamide hydrochloride | Unknown (for NHEJ) | Santa Cruz | Goglia et al. | You many know this as Imodium |
| NU-7441 | DNA-PKcs inhibitor | Tocris | Zhao et al. | 45 PubMed hits for NHEJ inhibition |
| DMNB | DNA-PKcs inhibitor | Santa Cruz | Durant et al. | Chemical derivative of vanilla |
Calculating NHEJ repair values
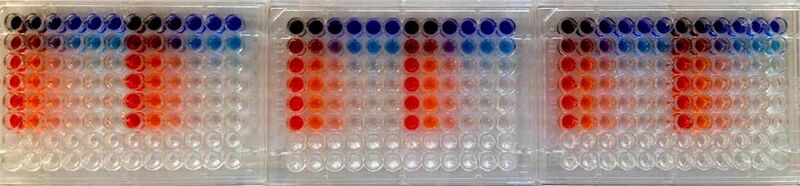
Now is the time to clearly understand the nature of the flow cytometry controls that you will perform. (We will save until next time an explanation of how flow cytometry works and a related discussion of additional controls that will be performed by the teaching faculty.) For each cell population, you will prepare two DNA mixtures: intact pMax-GFP plus damaged pMax-BFP-MCS, and intact pMax-GFP plus intact pMax-BFP. The re-circularization of pMax-BFP-MCS (less nonsense insert) will be our most direct readout of how much repair occurred. In this simplest view, broken DNA = no fluorescence and repaired DNA = blue fluorescent signal. However, what if one cell population simply took up more plasmid than another? In more technical terms, what if the transfection efficiency is higher for one cell type than for the other, and therefore the repair rate artificially appears higher? To control for this factor, we co-transfect with intact pMax-GFP – a transfection control – and normalize according to its uptake. But what if GFP and BFP plasmids are either taken up at different frequencies, or successfully expressed at different frequencies and/or signal intensities? Here is where the dual intact control is useful. It shows us the typical ratio of BFP:GFP uptake and expression, which we can use as a secondary normalization. Note that we use pMax-BFP as a control rather than pMax-BFP-MCS, because the latter will have very low expression that is not representative of the repaired construct: the nonsense insert separates the promoter and gene by too great a distance for robust expression.
Putting all of the above information together and in mathematical form, the NHEJ repair frequency equation can be determined in three steps:
- (1) Raw XFP reporter expression = % cells positive for XFP x MFI(XFP) = "RAW"
- X can be "B" or "G" in our case
- MFI is mean fluorescence intensity or median fluorescence intensity
- (2) Normalized BFP expression [math]\displaystyle{ \qquad ={RAW_{BFP} \over RAW_{GFP}}\qquad }[/math] = "NORM"
- (3) Reporter expression percent [math]\displaystyle{ \qquad ={NORM_{BFP.damaged} \over NORM_{BFP.intact}}\qquad }[/math] = NHEJ repair value
So, how will we get the expression vectors into our cells? DNA can be put into mammalian cells in a process called transfection. Recall that if you want to, say, make a hamster cell fluoresce green, you can transfect it with DNA carrying the EGFP open reading frame, a promoter directing transcription of EGFP, and a signal sequence for polyadenylation of the mRNA. The promoter tells the cell that the EGFP sequence should be transcribed by RNA polymerase. The polyadenylation sequence assists in the export and stability of the mRNA so that it gets translated by the ribosome. The coding sequence tells the ribosome which amino acids should be joined together.
Mammalian cells can be transiently or stably transfected. For transient transfection, DNA is put into a cell and the transgene is expressed, but eventually the DNA is degraded and transgene expression is lost (transgene is used to describe any gene that is introduced into a cell). For stable transfection, the DNA is introduced in such a way that it is maintained indefinitely, often using antibiotic resistance.
There are several approaches that researchers have used to introduce DNA into a cell's nucleus. At one extreme there is ballistics. In essence, a small gun is used to shoot the DNA into the cell. This is both technically difficult and inefficient, and so we won't be using this approach! More common approaches are electroporation and lipofection. During electroporation, mammalian cells are mixed with DNA and subjected to a brief pulse of electrical current within a capacitor. The current causes the membranes (which are charged in a polar fashion) to momentarily flip around, making small holes in the cell membrane through which the DNA can pass.
The most popular chemical approach for getting DNA into cells is called lipofection. With this technique, a DNA sample is coated with a special kind of lipid that is able to fuse with mammalian cell membranes. When the coated DNA is mixed with the cells, they engulf it through endocytosis. The DNA stays in the cytoplasm of the cell until the next cell division, at which time the cell’s nuclear membrane dissolves and the DNA has a chance to enter the nucleus. Cells are therefore typically evaluated for fluorescence expression 1-3 days after transfection.
Protocols
Part 1: Transfection of cells with plasmid reporters
All manipulations are to be done with sterile technique in the TC facility.
Timing is important for this experiment, so calculate all dilutions and be sure of all manipulations before you begin.
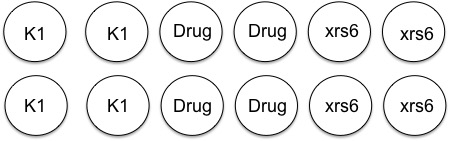
In anticipation of your lipofection experiment, the teaching faculty plated 35,000 K1 and 50,000 xrs6 cells per well in a 24-well dish at ~ 11 am yesterday. Half of the K1 cells then received NHEJ inhibitor at 10am today. A plating schematic for your experiment is shown below. Note carefully which cells are which, including those that have been pre-treated with inhibitor.
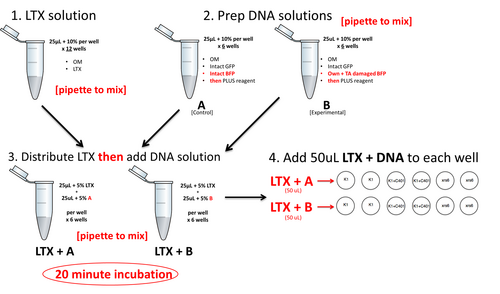
Below is a preview of the lipofection workflow. Do NOT begin these steps until you have completed the calculations further below. At each step, be sure to pipet to mix.
- Pre-label all of the eppendorfs that you will need.
- Prepare as much LTX diluted in OptiMEM (hereafter OM) as needed for all 12 transfections, plus 10%.
- The diluted LTX can sit at room temperature during the next two steps.
- Prepare the DNA mixtures – one control (A) and one experimental (B) – diluted in OM, enough for 6 transfections plus 10%, each.
- Add PLUS reagent to the DNA/OM mixtures.
- Distribute the LTX/OM to fresh eppendorf tubes, one for A and one for B. Here include 5% excess.
- For example, if 50 μL were required, you would add 52.5 instead.
- Then add the DNA/OM/PLUS (also 5% excess) to the LTX/OM.
- Incubate for 20 min at room temperature.
- Add 50 μL of the appropriate LTX/DNA/PLUS/OM mixture to each well. Read through the tips below before proceeding.
- Add the LTX/DNA/PLUS/OM drop-by-drop while making a circle with your pipet in the well, and then immediately swirl the plate (circularly) two times.
- Change tips between every addition.
- After distributing to all 12 wells, do a final mixing step for the whole plate, squeaky style: a few each of horizontal and vertical pushes.
The single-well basis reaction for each of the two DNA mixtures is shown below. Scale each one up to enough for 6 wells worth plus excess, carefully following the workflow above – for example, note that not everything is mixed together at once.
| Component | [Stock] ng/μL | Amount per control well (A) | Amount per experimental (B) |
|---|---|---|---|
| LTX (μL) | N/A | 0.5 | 0.5 |
| OM for LTX (μL) | N/A | to total 25 μL per well | to total 25 μL per well |
| intact pMax-GFP (ng) | 100 | 100 | 100 |
| intact pMax-BFP (ng) | 250 | 250 | 0 |
| damaged pMax-BFP-MCS (ng) | unique | 0 | 250 |
| extra cut DNA from Nova (ng) | 50 ng/μL | 0 | to supplement 250 above |
| PLUS (μL) | N/A | 0.5 | 0.5 |
| OM for DNA/PLUS (μL) | N/A | to total 25 μL per well | to total 25 μL per well |
Part 2: Peer Review of Methods Homework
Now that you've had two rounds of feedback and three rounds of drafting practice, you will put your knowledge to work by reviewing and correcting the Methods sections of your peers. First, read the text below as a guide to what your instructors look for while grading Methods sections. Next, read completely through the text of Methods draft that you will offer comments on. Finally, using whatever style is easiest for you, offer specific comments to your fellow 109er about what they have done well and what needs further work. Complete this activity using the "golden rule" -- offer the type of feedback that you hope to also receive. One of the best ways to improve your own skills is to teach something, so take advantage of this opportunity.
Once you have completed your review, please turn in your comments and the Methods section that you reviewed to the front bench. Shannon and Leslie will provide further comments on the Methods section that you reviewed, as well as comments to you as the reviewer to help you with these activities in the future.
(A) Holistic view of Methods section
When the instructors first read your Methods section, we begin by taking a holistic view of its structure. There are three main elements that all good methods sections must contain:
- An appropriate number of sub-sections
- Has the author divided the Methods into logical sub-sections based upon an entire experiment and not based upon completion day in the 20.109 lab?
- Descriptive sub-section titles
- Do the sub-section titles give enough information for the audience to easily pinpoint where the information they need will be found?
- Are the titles specific (i.e. Culture of CHO-K1 and xrs6 cells) or could they belong in any paper that involves cell culture (i.e. Cell culture)?
- Introductory sentences
- Does each sub-section start with an introductory sentence that states the goal of the experiment that was done? (For example, A Western blot was completed to determine Ku80 expression in CHO-K1 and xrs6 cells.)
(B) Specific experimental details
Next, we examine each Methods section to make sure that all of the details that are required to replicate your study are included. For this assignment, the important details are as follows (here we've included them within the sub-sections that would be appropriate for M2D1-D3):
- Cell culture sub-section
- cell types and origin (where did you get them?)
- cell culture media formulation
- culture conditions (temperature, CO2%, and relative humidity)
- Note: later you may choose to include the inhibitor name, concentration, and treatment times in this section.
- Western blot sub-section
- cell seeding density and incubation time
- lysis buffer formulation and lysis conditions (PBS rinse, temperature)
- protein quantification reagent, total protein separated on gel
- SDS-PAGE gel %, TE buffer formulation, electrophoresis conditions
- transfer buffer formulation, transfer conditions, membrane type, blocking buffer
- names of antibodies and dilutions, incubation times
- washing conditions and blot development
- DNA damage sub-section
- name of plasmid, mass digested
- name of enzymes, concentration of enzyme, reaction buffer
- reaction conditions, evaluation by electrophoresis
(C) Quality of writing
Finally, we provide comments on the writing style employed in each Methods section, concentrating on the following:
- Is this a protocol or a formal Methods section?
- Are volumes, vortexing steps, incubation times, etc. that are extraneous (or flexible) included? If so, this is a protocol.
- Is it concise?
- Are "filler words" avoided? Are sentences combined where possible to eliminate extraneous phrases and/or information?
- Is it logical?
- Do the 'steps' follow a logical progression? Again, is the section constructed to explain an experiment as a whole and not simply listed in the order in which it was performed in class?
- Is it written in complete sentences?
- Is the section easy to read or are there words missing/misspelled/misused?
For this activity, please comment on each of the three major evaluation criteria (A-C) listed above. You may do this by following the format that Noreen, Leslie, and Shannon employ -- numbering the paper and then writing an accompanying document. Or, you may simply write comments in a Word doc and provide a copy of that to your Methods section buddy.
Homework
- Your Module 1 microbiota summary revision is due by 5pm on Saturday Note: now Sunday, submitted to Stellar. Please remember to indicate the changes that you made (as instructed by the email sent to everyone) in order to qualify for the maximum possible grade increase.
Homework due on M2D6
- The major assessment for this module will be a research article describing your DNA replair study. For this assignment, you will start outlining and drafting the introduction to your report. Recall that the introduction provides a framework for the story you are about to tell, with respect to both background and motivation. You are encouraged to revisit the scientific writing guidelines before you begin.
- What might the big picture section consist of? Will you focus on cancer in general, the use fluorescent sensor technologies, mechanisms of DNA repair, or ... what?
- What background would the reader like to have as you zoom in? What biology topic(s) and what engineering topic(s) are relevant? Be sure to cite previous work to establish credibility.
- When you define your specific study, be sure to share your hypothesis about the affect of the damage topology and the NHEJ inhibitor you chose. You may also choose to speculate about how your system conditions will perform in relation to the others used by the class.
- At a minimum, you should hand in some high-level ideas and supporting notes for the first two sections, and write a draft of the third section in complete sentences.
- Revise your Methods section based on the peer comments that you were given in class. In addition to the complete draft of M1D1-M1D3, also include a sub-section title and outline for M1D4.
- Finally, on M2D6 we will discuss a paper about high throughput screening technologies applied to drug discovery. Goglia et al. identified two of the inhibitors that you could choose for today. Start scanning this paper now.
Reagent list
- Media as on Day 1
- Lipofectamine® LTX Reagent with PLUS™ Reagent (Life Technologies)
- Opti-MEM reduced serum medium (Life Technologies)
Navigation Links
Next Day: DNA repair assays Previous Day: Complete Western and prepare damaged DNA