20.109(S07): Lipofection
Introduction
DNA can be put into mammalian cells in a process called transfection. Mammalian cells can be transiently or stably transfected. For transient transfection, DNA is put into a cell and the transgene is expressed, but eventually the DNA is degraded and transgene expression is lost ("transgene" is used to describe any gene that is introduced into a cell). For stable transfection, the DNA is introduced in such a way that it is maintained indefinitely. Today you will be transiently transfecting your cultures of mouse embryonic stem cells.
There are several approaches that researchers have used to introduce DNA into a cell's nucleus. At one extreme there is ballistics. In essence, a small gun is used to shoot the DNA into the cell. This is both technically difficult and inefficient, and so we won't be using this approach! More common approaches are electroporation and lipofection. During electroporation, mammalian cells are mixed with DNA and subjected to a brief pulse of electrical current within a capacitor. The current causes the membranes (which are charged in a polar fashion) to momentarily flip around, making small holes in the cell membrane that the DNA can pass through.
The most popular chemical approach for getting DNA into cells is "lipofection." With this technique, a DNA sample is coated with a special kind of lipid that is able to fuse with mammalian cell membranes. When the coated DNA is mixed with the cells, they engulf it through endocytosis. The DNA stays in the cytoplasm of the cell until the next cell division at which time the cell’s nuclear membrane dissolves and the DNA has a chance to enter the nucleus.
Today you will lipofect the genetically-encoded calcium sensor into your mouse embryonic stem cells. This will cause the cells to fluoresce green. Next time we will measure fluorescence to assess the success of the transfection and to detect differences in intercellular calcium in response to agents that affect calcium flux and free calcium concentration.
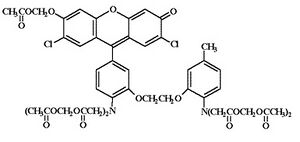
We'll also compare the genetically-encoded calcuim sensor to a commonly used calcium-sensitive indicator dye, Fluo-3, AM. We used the Fluo-3 salt last time to measure the concentration of Ca2+ in vitro. Today we will use an acetoxymethyl ("AM") ester derivative of the Fluo-3 salt which is cell-permeant. Once inside the cells, the Fluo-3, AM is cleaved by resident esterases to yield the active indicator.
Protocols
Half the class will begin by working in the cell culture facility while the other half discusses the journal article that was assigned for today, namely Nagai et al PNAS (2004) 101:10554. Midway through the lab time, you'll switch tasks.
Part 1: Transfection
All manipulations are to be done with sterile technique in the TC facility.
Timing is important for this experiment, so calculate all dilutions and be sure of all manipulations before you begin.
Before you begin, decide with your partner who will prepare the "slide #1" and who will prepare "slide #2"
Each transfection will require 2.5 ul of Lipofectamine 2000 in 50 ul OptiMEM. You and your partner should work together to dilute enough "carrier" for 10 lipofections. Let the dilution sit in the hood undisturbed for at least 5 minutes but not more than 30.
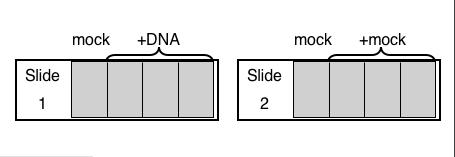
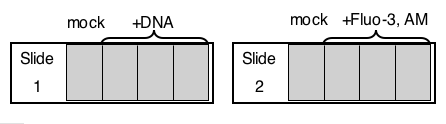
SLIDE #1 : inverse-pericam
- Slide #1 will have one "mock transfection" well, i.e. no DNA in the transfection mix. Prepare an eppendorf tube with 50 ul OptiMEM, enough for one mock transfection.
- Slide #1 will have three wells transfected with DNA. For each well you'll need 0.1 ug of "inverse-pericam" in 50 ul OptiMEM. Prepare a single eppendorf tube with enough DNA and OptiMEM to transfect all 3 wells. By making one transfection cocktail that is later divide between replicates, you can be confident that each replicate was treated identically.
- You should now have three eppendorf tubes, two with OptiMEM +/- DNA and one with diluted Lipofectamine 2000. Add an equal volume of diluted Lipofectamine to the eppendorf tubes with OptiMEM +/- DNA (i.e., 50 ul if the tube has 50 ul). Pipet up and down to mix.
| Tube | [DNA] stock | DNA/lipofection | Volume DNA for 3 transfections | Volume OptiMEM for 3 transfections |
|---|---|---|---|---|
| inverse pericam DNA | 0.1 ug/ul |
SLIDE #2: Fluo-3, AM
- Slide #2 will have four "mock transfection" wells, i.e. no DNA in the transfection mix. Prepare an eppendorf tube with 200 ul OptiMEM, enough for four mock transfection.
- Add an equal volume of diluted Lipofectamine to the eppendorf tubes with OptiMEM (i.e., 50 ul if the tube has 50 ul). Pipet up and down to mix.
For both Slide #1 and Slide #2
- Incubate the OptiMEM +/- DNA and lipofectamine cocktails at room temperature for 20 minutes. To allow the DNA/carrier complexes to form, it is important that you do not disturb the tubes during this incubation. During this time, aspirate the media from the cells in your 4-well slide, wash the wells with 0.5 ml PBS, then put 0.5 ml of fresh pre-transformation media on the cells. The PBS and media can be aliquoted with a 2 ml pipet.
- After the 20 minute incubation is over, use your P200 to add 95 ul of the appropriate DNA:lipofectamine complexes to each well. Since the carrier is quite toxic to the cells it’s a good idea to gently rock the slide back and forth after each addition.
- Return the slide to the petri dish and the dish to the 37°C incubator.
- Tomorrow, one of the teaching faculty will remove the lipofection media from your cells and will replace it with 1 ml of fresh media. The following day, one of the teaching faculty will add 1 ml of 2 uM Fluo-3, AM to the last three wells of "slide #2" which wasn't transfected with DNA. The cells require only 45 minutes at 37° to uptake the dye, then only 30 minutes at 37° to cleave it into an active form inside the cell.
- Approximately 48 hours after performing the lipofection, you and your partner will examine the cells and analyze their fluorescence after different treatments that might affect calcium signaling. Before you leave today, see one of the teaching faculty to schedule your microscope time.
Part 2: Journal article discussion
Guided by the questions in the "for next time assignment" from last lab and also by the figures and supplementary material that are included in Nagai et al PNAS (2004) 101:10554, we will discuss the relevant aspects of this article. You will spearhead a particular aspect of the discussion, but you should be willing and able to participate in all aspects of the conversation.
DONE!
For next time
- Review the descriptions of the lab's microscopes that we covered during our lab orientation.
- Plan your experiment for next time. The plasmid you transfected today was originally described in Nagai et al PNAS (2001) 98:3197.
Look at Figure 5 in this article and then choose some calcium-mobilizing agents you might like to try in lab next time. You can also look at the Miyawaki et al Nature (1997) 388:882 article, Figure 3 in particular, for additional ideas.
Be sure to learn a little about the agent(s) you've chosen, thinking about the concentration(s) you'll work with and how you expect the chemicals to affect the fluorescence.
A schematic of the slides you and your partner set up today is provided to help you plan your experiment. A table is provided to help as well. Please fill these out as needed, in consultation with your lab partner, and bring them with you next time you come to lab.
| agent | [stock] | working concentration (if [final] is <1:100 of [stock]) | final concentration |
|---|---|---|---|
| histamine | 100 mM | ||
| cyproheptadine hydrochloride | 0.1 M | ||
| A23187 | 2 mM | ||
| BAPTA-AM | 30 mM | ||
| CaCl2*2H20 | 3 M | ||
| MgCl2*6H20 | 3 M | ||
| EGTA | 0.5 M |
Reagents list
- OptiMEM
- Lipofectamine 2K
- Plasmid DNA for "inverse-pericam" described in Nagai et al PNAS (2001) 98:3197