20.109(S07): Bio-material engineering/Sequence analysis
Introduction
The invention of automated sequencing machines has made sequence determination a fast and inexpensive endeavor. The method for sequencing DNA is not new but automation of the process is recent, developed in conjunction with the massive genome sequencing efforts of the 1990s. At the heart of sequencing reactions is chemistry worked out by Fred Sanger in the 1970’s which uses dideoxynucleotides.
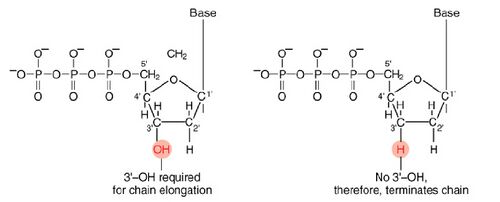
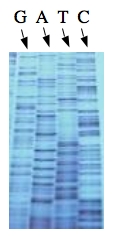
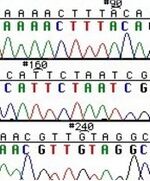
These chain-terminating bases can be added to a growing chain of DNA but cannot be further extended. Performing four reactions, each with a different chain-terminating base, generates fragments of different lengths ending at G, A, T, or C. The fragments, once separated by size, reflect the DNA’s sequence. In the “old days” (all of 10 years ago!) radioactive material was incorporated into the elongating DNA fragments so they could be visualized on X-ray film (image on left). More recently fluorescent dyes, one color linked to each dideoxy-base, have been used instead. The four colored fragments can be passed through capillaries to a computer that can read the output and trace the color intensities detected (image on right). Your sample was sequenced in this way on an ABI 3730 DNA Analyzer.
Analysis of sequence data is no small task. “Sequence gazing” can swallow hours of time with little or no results. There are also many web-based programs to decipher patterns. The nucleotide or its translated protein can be examined in this way. Thanks to the genome sequence information that is now available, a new verb, “to BLAST,” has been coined to describe the comparison of your own sequence to sequences from other organisms. BLAST is an acronym for Basic Local Alignment Search Tool, and can be accessed through the National Center for Biotechnology Information (NCBI) home page
Protocol
The data from the MIT Biopolymers Facility is available for you to examine. Retrieve the sequences from this link. Choose the "Login to dnaLIMS" link and then use "nkuldell" and "20.109" to login. At the bottom of the left panel should be a link to download your sequencing results. Select the appropriate order # (you'll be told which one is correct) and then "submit." From the list find your samples. The quickest way to start working with your data is to follow the "view" link. From this link you'll see the sequencing traces themselves and can add the sequence to the workbox by clicking on "sequence text."
A good place to start your sequence analysis is to copy the sequence text into the translation program that is freely available from EMBL. Be sure to translate in all 3 reading frames since the first nucleotide that's listed on the sequence file may not be the first nucleotide of a codon. The table below should help orient you to the salient parts of your data. The translated sequence is presented in single letter code, where X indicated ambiguity in the sequence data. The four library sequences do not necessarily bind gold.
| pCT-CON | YALQA SGGGG SGGGG SGGGG SASCG GGGTS KISHF LKMES LNFIR AHTPY INIYN CEPAN PSEKN SPSTQ YCYSI QSSQV DCGGG SEQKL ISEED L**LEI **QQ |
| pAu1 | YALQA SGGGG SGGGG SGGGG SASQV QLQQS GPGLV KPSQT LSLTC AISGD SVSGN TAAWN WIRQS PSRGL EWLGR TYYRS KWHYD MRHL* KVE* |
| Library seq1 | YALQA SGGGG SGGGG SGGGG SASQG GGGSG PPRRR SNVWA PV*LA RPVAW GRIRT KAYF* |
| seq2 | YXXXA SGGGG SGGGG SGGGG SASQG GGGSG VYGLS GTARS RG*LA RPVAW GRIRT KAYF* |
| seq3 | YXXXA SXXGG SGGGG SGGGG SASQG GGGSG KRGCS RALWW IA*LA RPVAW GRIRT KAYF* |
| seq4 | YXLQA SGGGG SGGGG SGGGG SASQG GGGSG WKMFI GGTWL GC*LA RPVAW GRIRT KAYF* |
As you consider your data, you should also explore what is known about amino acid interaction with metals, using search engines such as PubMed, MIT’s homepage or even Google, and also consider the data from your classmates. Collaborating in this way may support any developing theory you have. Before you leave, please post your data (sequence, relative strength of gold binding, and so on) to the discussion page associated with this lab and write a few comments about the results.
REALLY DONE!
