20.109(F10): Mod 2 Day 5 Assessing re-tuned system
Assessing Re-tuned System
Introduction
Today is a day when you'll start to collect information about the mutant candidates you've chosen. Over the next 3 lab sessions, you will look at
- the expression level of the mutant Cph8 proteins relative to the wildtype light sensor
- the DNA sequence of the region around the K+ hotspot
- the β-gal activity of the mutant photography strains grown in the dark
- the resulting bacterial photograph taken with your mutant strain
The data should be informative, though preliminary.
Cph8 Library
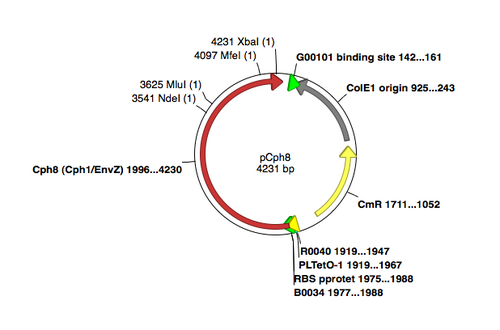
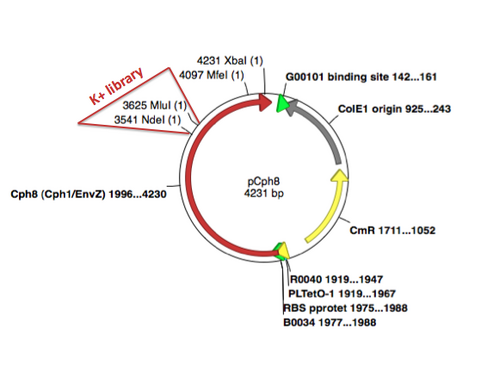
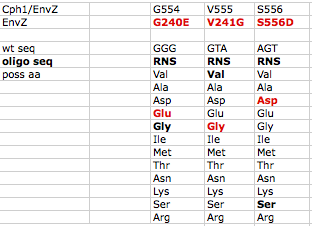
A map of the plasmid that encodes the light-sensor protein, Cph8, is shown. To make the libraries that we used last time, the plasmid was modified with a degenerate oligonucleotide spanning the sequence between the NdeI and MluI restrictions sites in Cph8. The pool was synthesized with a mixture of nucleotides at the K+ relevant positions G554 V555 S556, as shown in the reagents list from today's lab. PCR was used to extend the degenerate oligos to incorporate them into the pCph8 backbone. The PCR products are a mixture of different sequences since the initial oligo template that was PCR amplified had a variety of sequences. Twelve possible amino acids could be encoded at each of the three variable positions. Consequently, there are more than 1700 possible variants (12x12x12) in the library. The resulting pool of plasmid variants is what you transformed and screened last time.
DNA Sequencing
Though you don't know for certain which, if either, of your mutants exhibits detectably different activity in the context of the bacterial photography system, you will send plasmid DNA from both mutants for sequencing. The invention of automated sequencing machines has made sequence determination a fast and inexpensive endeavor. The method for sequencing DNA is not new but automation of the process is recent, developed in conjunction with the massive genome sequencing efforts of the 1990s. At the heart of sequencing reactions is chemistry worked out by Fred Sanger in the 1970’s which uses dideoxynucleotides.
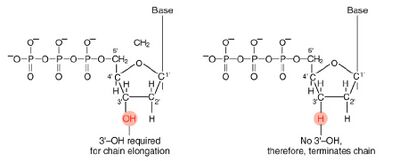
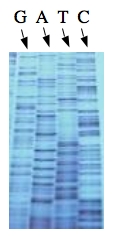
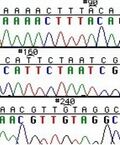
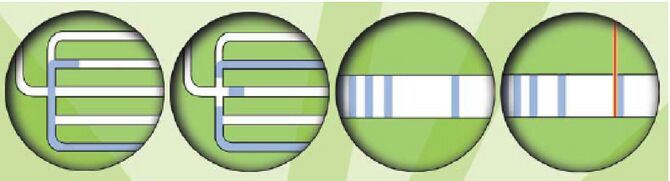
These chain-terminating bases can be added to a growing chain of DNA but cannot be further extended. Performing four reactions, each with a different chain-terminating base, generates fragments of different lengths ending at G, A, T, or C. The fragments, once separated by size, reflect the DNA’s sequence. In the “old days” (all of 15 years ago!) radioactive material was incorporated into the elongating DNA fragments so they could be visualized on X-ray film (image on left). More recently fluorescent dyes, one color linked to each dideoxy-base, have been used instead. The four colored fragments can be passed through capillaries to a computer that can read the output and trace the color intensities detected (image on right). Your sample will be sequenced in this way on an ABI 3730 DNA Analyzer.
Today you will isolate plasmid DNA from the cells to send for sequencing and to check by digest. You'll also measure the β-gal activity associated with your mutant candidates. If you choose, you can set up new cultures of your mutants in the light and the dark to measure the difference in β-gal activity next time.
Part 1: DNA analysis and sequencing
You will isolate plasmid DNA from the overnight cultures of the 2 mutant candidates and the wild type photography system, NB334. The miniprep protocol we'll follow is already familiar to you from Module 1. Each sequencing reactions requires 200-500 ng of plasmid DNA, and 3.2 pmoles of sequencing primer in a final volume of 12 μl.
- Resuspended your plasmid DNA pellets in 40 ul of sterile H2O. The miniprep'd plasmid should have ~1 ug of nucleic acid/ul but that will be a mixture of RNA and DNA, so we will guess at the amount of plasmid DNA.
- Next you should prepare three eppendorf tubes, one to sequence each of the two mutants and one to sequence the wild type . Each tube should have
- 2 ul of your plasmid DNA, mutant candidate or wild type
- 6.4 ul of sequencing primer NO289 (seq = ATTACCGCCTTTGAGTGAGC, this has already been diluted 1:100 from the stock).
- 15.6 ul sterile water
- Pipet this sequencing solution up and down to mix and then transfer 12 ul to one of the tubes in the 8-tube strip we will turn in at the biopolymer's facility in E17 for sequencing. The data will be available for you to examine < one week from today. Keep your fingers crossed.
- To check the library candidates for the presence of the library fragment, the candidate DNA samples and the wild type will be digested. Add 10 ul of each candidate DNA to eppendorf tubes, label them with your team color and a number, and give them to the teaching faculty. These aliquots of DNA will be digested with NdeI and XbaI, enzymes that will release a 700 bp fragment if the DNA from the mutant candidates has Cph8. The digests will be run on a gel by the teaching faculty and the data posted to today's "talk" page when it is available.
Part 2: β-galactosidase Assay
You will measure the β-gal activity associated with the wild type and the mutant candidates grown in the dark. You can also set up overnight cultures for your samples in the light and dark if you choose to collect that data next time. Review the protocols here and here for details and ask the teaching faculty if you need help. Calculate the b-gal activity of your samples before you leave today.
DONE!
For Next Time
- To be turned in: You should prepare an outline for the introduction to the research article that you will write to describe your work in this experimental module. Your outline should include references that you have found to describe the system and your tuning of it. At this point the references can be in any format you choose.
- NOT to be turned in: You should update the Materials and Methods section that you are writing for your research article to include the experiments you have performed today.
Reagents
- for β-gal assay
- Z-buffer
- 0.06 M Na2HPO4
- 0.04 M NaH2PO4*H2O
- 0.001 M KCl
- 0.0001 M MgSO4
- 4 mg/ml ONPG
- 1M Na2CO3
- Z-buffer
- for alkaline lysis miniprep
- Solution I
- 25 mM Tris pH8
- 10 mM EDTA pH8
- 5 mM Glucose
- Solution II
- 1% SDS
- 0.2M NaOH
- Solution III
- 3M KAc, pH 4.8
- for agarose gel analysis
- Loading Dye
- 0.25% xylene cyanol
- 30% glycerol
- RNase
- 1% agarose gel in 1X TAE
- 1X TAE
- 40 mM Tris
- 20 mM Acetic Acid
- 1 mM EDTA, pH 8.3
- Sequencing primer NO289
- 5' ATTACCGCCTTTGAGTGAGC
- Loading Dye