IGEM:IMPERIAL/2006/project/Oscillator/project browser/Test Killing Predator Construct/Design
| Super Parts | Predator Construct | |||
|---|---|---|---|---|
| Actual Part | AiiA Testing Construct <bbpart>J37022</bbpart>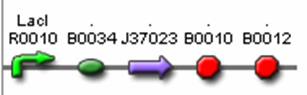
| |||
| Sub Parts | <bbpart>R0010</bbpart> | <bbpart>B0034</bbpart> | <bbpart>J37023</bbpart> | <bbpart>B0015</bbpart> |
Design Introduction
Part J37022 is comprised of an AiiA protein coding region with a FLAG immunotag on the 5’ end and a LVA degradation tag on the 3’ end of the sequence. This AiiA protein coding region is ligated to both an RBS and a terminator. The transcription of the entire sequence is controlled by IPTG inducible LacI promoter.
The actual construct that will be placed into the predator cell will be the test construct without the LacI promoter.
Ideally, we would like to be able to characterise AiiA before we implement it into our predator cell and actually build the biological oscillator construct. We strive to measure the half-life of our AiiA as well as Vmax and Km, as defined by Michaelis-Menten enzyme kinetics.
| INPUTS | biological component(s) | comments |
|---|---|---|
| IPTG | Inducing Sugar | Isopropyl thiogalactopyranoside (Galactose analogue) sensed by the LuxR promoter (see also description below) |
| OUTPUTS | biological component(s) | comments |
| AiiA | AHL-lactonase | Degrades AHL, produced by <bbpart>J37023</bbpart>, LVA & FLAG tagged, see also description below |
| FUNCTIONS | biological component(s) | comments |
| Inducer | pLacI promoter <bbpart>R0010></bbpart> | senses IPTG and initiates transcription, see also description below |
| Prey Killing | enzymatic reaction | enzymatic reaction from aiiA on AHL . |
| Predator Death | growth dilution | Degradation of AiiA through LVA tag and natural metabolic pathways; cell dilution and growth also affect concentration of AiiA |
The LacI Promoter
The LacI promoter is derived from the Lac operon sequence normally found in E. coli. In bacterial cells, lactose is generally not mebolized if glucose is present. This is due to lactose requiring more energy to cleave the 1-4 B glycosidic bond and processing of glactose (one of the breakdown products of lactose). When lactose is not present, CAP (Catabolite Activated Protein) binds tightly to the DNA not allowing the RNA polymerase to bind, and thus not allowing transcription to occur. The arrival of lactose induces a second messenger system inducing cAMP (cyclic adenosine monophosphate) to bind to CAP, causing a conformation change in the protein, thus releasing it from the DNA. Transcription is then allowed to proceed.
Lactose analogues such as IPTG (Isopropyl thiogalactoside) were found to activate this second messenger cascade system, thus inducing transciption of the lac operon. We can use this as an alternative way of activating the transcription of the downstream genes allowing us a degree of control of expression. IPTG is favoured over lactose in experimental biology since it will not interfere with metabolism, as it is not broken down by the enzyme B-galactosidase, which cleaves lactose into glucose and galactose.
The LacI promoter part (R0010) is derived from a segment of DNA coding for the repressor protein, a CAP binding site, and the promoter site. The DNA coding for the repressor protein is inactivated by splicing in the middle, as CAP and cAMP are produced automatically by the E. coli cells.
We can then use the LacI promoter to observe various amounts of protein (in our case, AiiA) being produced for a given concentration of IPTG.
J37023
The part <bbpart>J37023</bbpart> contains the AiiA with an LVA tag and an immunoassay tag. In solution, the AiiA protein is stable and has a relatively long degradation period. [1] However, our final construct depends on having a fluctuating level of AiiA producing the prey oscillations controlling the predator population. The addition of an LVA tag increases degradation rate by "flagging" the protein as a metabolite. Since we do not know a quantitative value for the degradation rate with the LVA tag, we must perform testing of the construct before combining it to form the final predator construct.
The immunotag is not used in the final construct once we combine the two cell populations; however, it is used as a method of measuring the amount of AiiA in our solution. This will be helpful when characterising the enzyme activity through using Michaelis-Menten kinetics as discussed further.
B0034 & B0015
Part <bbpart>B0034</bbpart> codes for a ribosome binding site (RBS) on the mRNA to enable translation of our gene. Part <bbpart>B0015</bbpart> is a combination of two terminator regions which signals the RNA polymerase to stop transcribing.