Zhang Lab @ UMB
Welcome to the Network Informatics LabUniversity of Maryland Marlene and Stewart Greenebaum Cancer Center, Department of Epidemiology and Public Health, University of Maryland School of Medicine |

Recent News. 06/05/2009, New BMC Bioinformatics Paper Yuanxin's paper titled 'BSMAP: whole genome Bisulfite Sequence MAPping program' has been accepted by BMC Bioinformatics. Congratulations Yuanxin!
Kaifu Chen has accepted our postdoc offer. Kaifu has a PhD in Genomics and Bioinformatics from Beijing Institute of Genomics, Chinese Academy of Sciences. Welcome Kaifu!
Peng Yu has accepted our postdoc offer. Peng will soon finish his PhD in computer engineering at the University of Texas at Austin, and join our lab this summer. Welcome Peng!
A paper titled 'Reprogrammed Androgen Receptor Function in Androgen-Independent Prostate Cancer' has finally been accepted by Cell.
Please join us on May 2nd (Saturday) 2009 at MD Anderson Cancer Center. This meeting is organized by Steffi Oesterreich, Wei Li and Jean-Pierre Issa.
Julia Kristine Blackmore, a PhD student in BCM MCB dept., will be doing a two-month rotation in our lab.
. 01/21/2009, New Paper A paper titled 'Integrative analysis of HIF binding and transactivation reveals its role in maintaining histone methylation homeostasis' has been accepted by PNAS.
| |
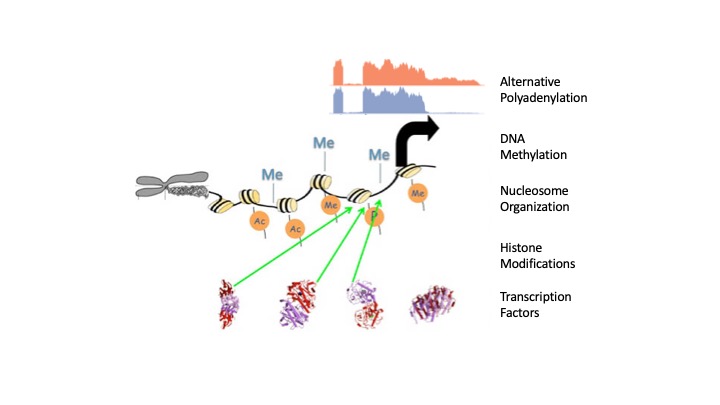
A Genomic View of Epigenetic and Transcriptional RegulationOur lab is focused on the design and application of statistical and computational algorithms to elucidate global epigenetic and transcriptional regulatory mechanism, by interpreting and integrating data from ChIP-chip/seq, DNA methylation, Nucleosome positioning, Alternative splicing and Motif finding. An elaborate system of epigenetic and transcription regulation is responsible for the morphological and behavioral complexity in higher eukaryotes. This regulatory system consists of diverse trans-acting protein factors, cis-acting regulatory DNA sequences and the underlying epigenomic background, such as histone modifications, DNA methylation and Nucleosome localizations. Recently, Chromatin ImmunoPrecipitation coupled with whole genome tiled microarray (ChIP-chip) and/or next-generation sequencing (Solexa, SOLiD and 454) has evolved as a powerful and unbiased technique to study this genome-wide regulatory system. The application of this technology to multiple factors and/or in multiple conditions allows biologists to study how transcription is differentially regulated in a combinatorial manner. However, it also poses great challenges for the development of effective algorithms, the key link between massive raw data and biological hypotheses. We developed a series of algorithms to reliably detect and annotate ChIP-enriched regions using Next-generation sequencing (MACS; Genome Biology 2008) and Affymetrix whole-genome tiling arrays, including 1) Model-based Analysis of Tiling-arrays (MAT; PNAS 2006) and a hidden Markov model (Bioinformatics 2005) for ChIP-region detection, 2) extreme MApping of OligoNucleotide (xMAN; BMC Genomics 2008) for microarray probe mapping, 3) Cis-regulatory Element Annotation System (CEAS; NAR 2006) for ChIP-region annotation. Since the inception in early 2006, they have been adopted by hundreds of academic users and are now considered as the ChIP-chip data analysis standard in many labs. We worked with ENCODE consortium to systematically analyze the performance variability introduced in ChIP-chip protocols, array platforms, and analysis methods (Genome Res. 2008). Furthermore, we are also in close collaboration with several labs on identifying global regulation targets of several key transcription factors, including Estrogen Receptor (Cell 2005; Nature Genetics 2006); Androgen Receptor (Molecular Cell 2007; Cell 2009) and FoxA1 (Cell 2008). We are currently collaborating with many BCM laboratories to use the Next generation sequencing to study 1) Transcription factor binding (ChIP-seq); 2) DNA methylation at single nucleotide resolution (BS-seq); 3) Nucleosome positioning (Nu-seq); 4) Cancer specific alternative splicing junctions (RNA-seq). My laboratory also plays an important role in the BCM Epigenomics Data Analysis and Coordination Center for a five-year $190 million NIH Roadmap Epigenomics Program.
|