Xwang:Research
PROJECT 1. Regulation of early endosperm development in maize
We have constructed a local UCSC genome browser at UA to display the maize endosperm RNA-Seq transcriptome.
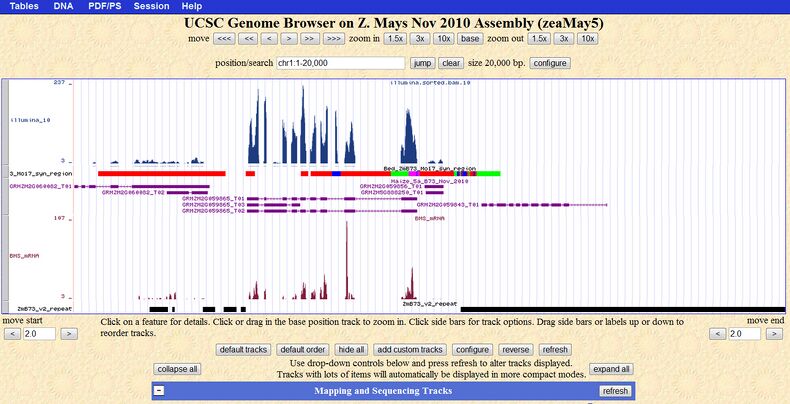
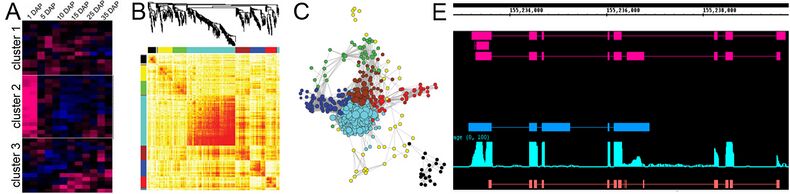
We first conducted a in silico search of maize annotated genes in its 5.0a version in plant transcription factor database, and identify hundreds potential TF maize genes. Then we utilized publish gene microarray expression data and our self-produced RNA-seq data to further filter the gene sets. Our efforts identified ~200 candidate genes that specifically expressed in early stages of endosperm development (A). We are currently performing experimental validation of selected candidate TFs. In the next step, to construct the regulatory network of those identified TFs, we identified the genes whose expression highly correlated with those TFs and calculated their topological structures (B). The regulatory network is finally illustrated in (C). Additionally, we are developing hidden markov model (HMM) to detect the alternative splicing genes during the development (D).
PROJECT 2. Chromatin and epigenomic landscape of the developing maize endosperm
Heterosis, or hybrid vigor, is the increased function of any biological quality in a hybrid offspring. It is the occurrence of a genetically superior offspring from mixing the genes of its parents. Nearly all field corn (maize) grown in most developed nations exhibits heterosis. Modern corn hybrids substantially outyield conventional cultivars and respond better to fertilizer.

Above figure was modified from two papers by (Springer and Stupar 2007 and He et al, 2010)
We are sequencing two activating histone marks H3K9ac and H3K36me3, which are usually associated with transcription initiation and elongation, respectively. The former peaks at TSSs of actively transcribed genes, while the latter spreads along the entire gene body regions (A). Therefore, the two modifications have been considered hallmarks for active promoters or actively transcribed genes (B). The current version maize genome annotation contains certain computationally predicted genes with wrongly defined gene borders (multiple genes were predicted as one gene). Because H3K9ac marks the gene start and H3K36me3 marks the gene body region, we can use these information to correct the wrongly annotated genes. In our Lab, we are developing supervised Hidden Markov Model (HMM) to integrate the epigenetic information with RNA-Seq signal information to finally improve the gene models in maize.

Genomic imprinting refers to the preferential expression of maternal or paternal alleles of genes in progeny. The imprinted genes are usually marked by silencing epigenetic modifications. In flowering plants, the endosperm is the primary tissue where gene imprinting is associated with a developmental control mechanism. However, only a handful of imprinted genes have been discovered.
PROJECT 3. Assembly and annotation of Thellungiella halophila genome
We are also collaborating with Dr. Karen Schumaker Lab on genome sequencing project of Thellungiella halophila, which is a halophytic relative of both the genetic model Arabidopsis thaliana (Arabidopsis). The genome sequence of T. halophila will be a critical resource for the fields of stress biology, evolutionary biology, and comparative genomics. We are currently working on using Arabidopsis as reference genome to assemble the hellungiella halophila. We are also analyzing the mRNA transcriptome sequenced by 454 under several stress libraries such cold, salinity and drought to identify the genes involved in stress resistance.