Wikiomics:RNA phylogenetics
From OpenWetWare
Jump to navigationJump to search
Many phylogenetic analysis tools assume that each site in an alignment is independant of all other sites. This is not generally true, particularly in the case of structural RNAs where base-paired sites co-vary with respect to each other. Some of the early work on site-dependant models was by Arndt von Haeseler and co-workers [1]. More recently Paul Higgs and co-workers have compared a number of different evolutionary models [2]
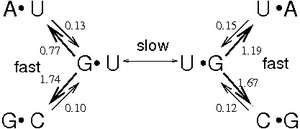
References
- von Haeseler A and Schöniger M. Evolution of DNA or amino acid sequences with dependent sites. J Comput Biol. 1998 Spring;5(1):149-63. DOI:10.1089/cmb.1998.5.149 |
- Savill NJ, Hoyle DC, and Higgs PG. RNA sequence evolution with secondary structure constraints: comparison of substitution rate models using maximum-likelihood methods. Genetics. 2001 Jan;157(1):399-411. DOI:10.1093/genetics/157.1.399 |
- Higgs PG. RNA secondary structure: physical and computational aspects. Q Rev Biophys. 2000 Aug;33(3):199-253. DOI:10.1017/s0033583500003620 |
RNA phylogenetic analysis tools
- CBCanalyzer Inferring phylogenies based on compensatory base changes.
- jRNA Exploring insect (and other less interesting) phylogenies using RNA secondary structure. Provides tools structural RNA alignment and analysis.
- MrBayes Bayesian Inference of Phylogeny. Allows doublet (secondary structure) models.
- PHASE Designed specifically for use with RNA sequences that have a conserved secondary structure, e.g., rRNA and tRNA. Substitution models of sequence evolution that consider pairs of sites rather than single sites are implemented in this package along with standard nucleotides substitution models used nowadays. When a RNA molecule with a secondary structure is used in conjunction with a RNA substitution model, PHASE requires a structure-based alignment of the sequences with the consensus secondary structure indicated in bracket and dot notation at the top of the alignment.
- RAxML Implements most of the RNA models described in PHASE (S6A,S6B,S6C,S6D,S6E,S7A,S7B,S7C,S7D,S7E,S7F,S16,S16A,S16B).
- RNAsalsa is suitable for routine applications in molecular phylogenetics based on structured RNAs, in particular rRNAs.
- RRNADIST, RRNAML and RNAML Empirical substitution models for Ribosomal RNA. Estimate phylogeny using maximum likelihood structural RNA models of sequence evolution.
- SISSI Simulating Sequence Evolution with Site-Specific Interactions: a software tool to generate data of related sequences along a given phylogeny, taking into account user defined system of neighbourhoods and instantaneous rate matrices.