User talk:Richard A. Acevedo/Notebook/Biology 210 at AU
Richard Acevedo February 3, 2016 Biology 210 Lab Keanan McDaniel’s
Lab 3- Microbiology and Identifying Bacteria
Purpose: “To characterize the most populous domain on earth- bacteria. You will identify the species of bacteria based on motility, gram stain, colony morphology and sequencing of the 16s ribosomal subunit gene. There are also objectives that will help with learning how to identify bacteria.”(Biology Lab Manual: Lab 3,pg41)
Materials and methods:
The first procedure was observing the bacterial growth on the agar plates from the Hay Infusion Culture that was taken from your transect. You must find the number of colonies and then convert finding the colonies per ml for each of the dilutions that have and don’t have the antibiotic Tetracycline.
The second procedure was comparing and contrasting the agar plates with and without the antibiotic. You had to see why some agar plates did or not grow and if not why they didn’t.
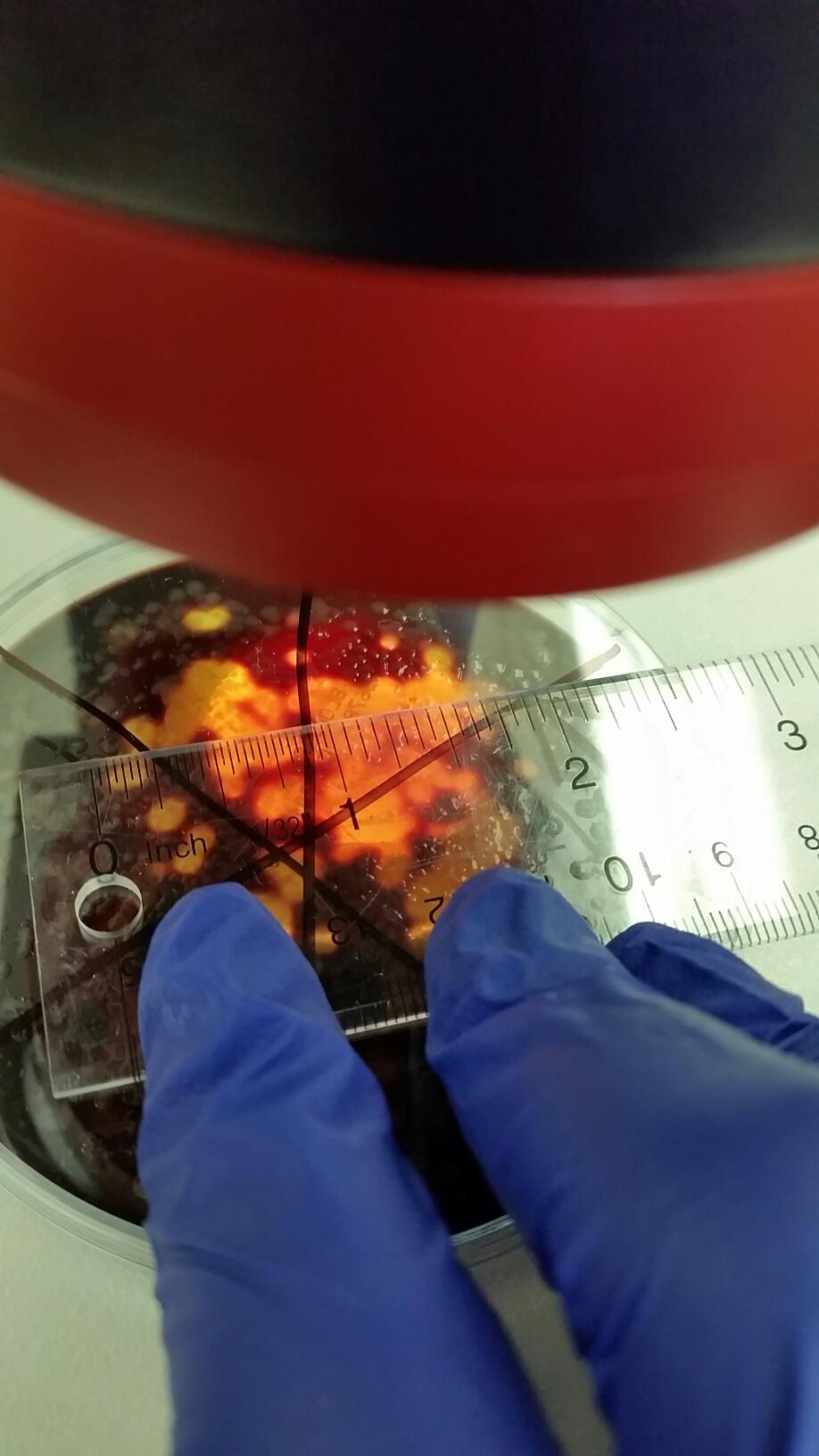
For procedure three, there were two parts. The first was observing prepared slides of bacteria, observing both native wet mount preparation and a gram stain of 2 colonies from the nutrient agar plate and 2 from the tetracycline. Once that was completed, label the colonies and describe them. Then observing more prokaryotes was required at different magnifications. The second part was observing 4 samples of microorganisms that include two from the nutrient agar plates and two from the nutrient agar plus tetracycline plates. Choosing two colonies need to be done that were isolated and then creating a wet mount and a gram stain had to be done.
When making a gram stain, sterilize a loop over a flame and scrape up a tiny amount of growth from the surface of the agar. Mix it by adding a drop of water to the slide that the growth is on. Once that is done, label each slide and draw a circle knowing where your growth piece is. After, heat up the slide to remove any water from the slide. Start by passing it at a good height where the water evaporates but does not kill the bacteria on the slide. The slide water will evaporate then add some crystal violet dye for a minute then remove it. Repeat this step but using Gram’s iodine for a minute and then remove it with water. You will then decolorize the slid by using 95% alcohol for 10-20 seconds, to determine if decolorization has occurred, check if the solvent on the slide flows colorlessly. Start by then coloring the slide with safranin stain for 20 seconds to half a minute, rinse the slide to remove after the time has expired. When finished air-dry the slide and take off any excess blots of water so it can be dry to examine it under the microscope. Examine the gram stained sample at a low magnification but then increase to 40x and 100x under oil immersion objectives.
For the wet mount, the first three steps of creating a gram stain were repeated but then a cover slip was added to cover the sample and observed at a 10x objective and 40x objective. Do not put over fire to dry water.
This now brings us to procedure IV which included using a PCR. “This was the setting up of the PCR for the 16s amplification.” Using the four plates you picked, two out of the four plates were picked that had the most complete characterization. The two agar plates had to be labeled and each group used primers and PCR to amplify the 16s rRNA gene. When amplifying the rRNA gene, first it started by labeling the 2 PCR tubes with the transect number, colony identifier and your group number. The labeling system was then written down in the lab notebook. 20 microliters of primer/water mixture was then added to a PCR tube that has been labeled and mixed together to dissolve the bead inside. A sterile toothpick was then used to pickup s small amount of a bacterial colony that you chose. The toothpick was then submerged in the primer/water mix PCR tube and twisted in it for 5 seconds, then discarded. The PCR tube was then sealed and placed in the PCR machine. This step was done when you used a different colony.
Data and Observations:
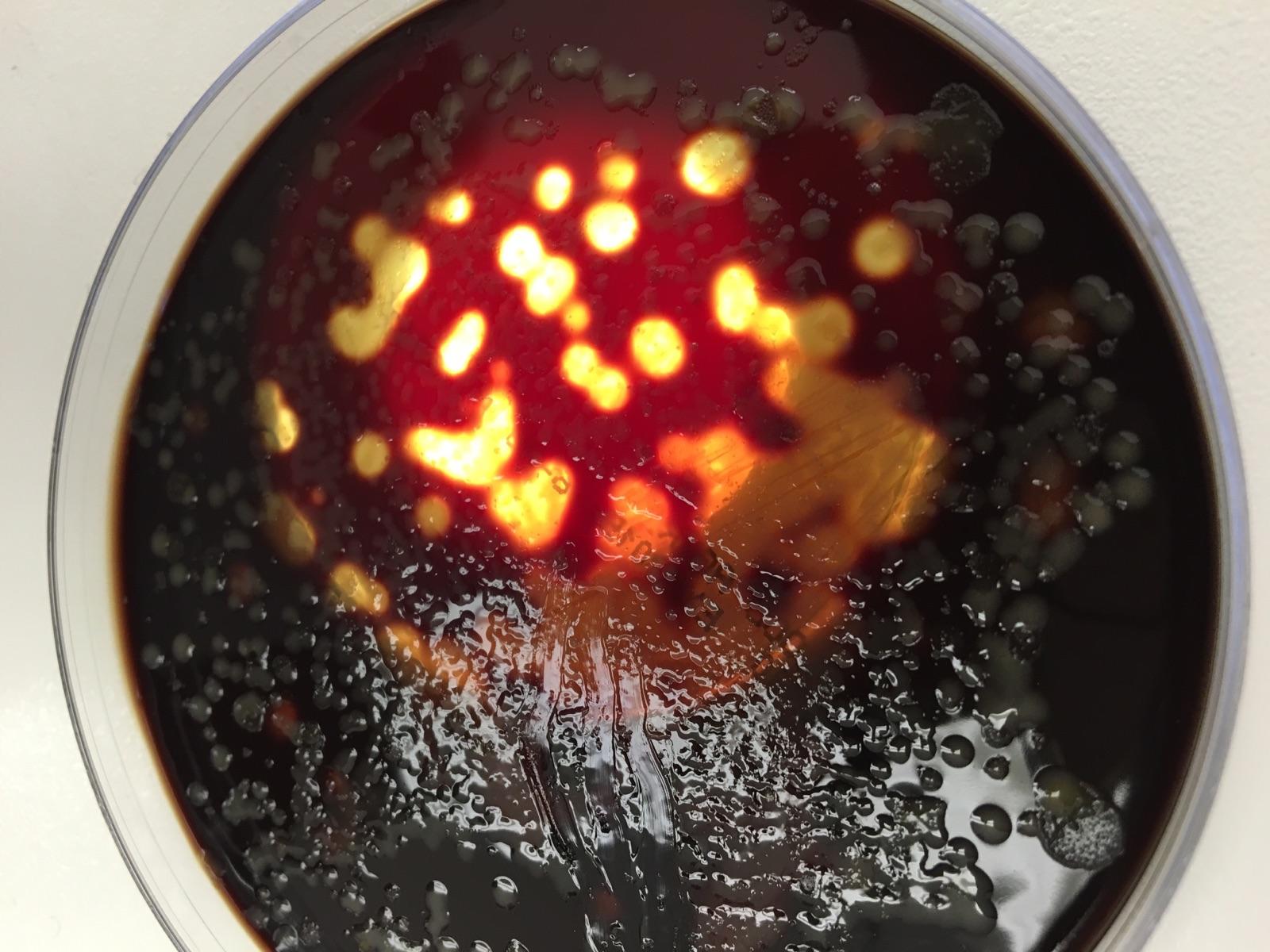
When observing our hay infusion culture, we noticed that some of the agar plates did not have any colonized bacteria when others had a good amount to thousands. They smelled really bad like the broth that was used in the lab before. To add, as the school year continues the appearance and smell might change due to the non-lethal toxin they bacteria will release and how they are growing.
The next observations is a table graph which shows the 100-fold serial dilution results which were seen and obtained by counting the number of colonies on the agar plates that had bacteria growth on them. From see the results, the number of colonies ranged from tens to thousands of colonies/ml. The ones that had no appearance of bacterial growth meant that they were not antibacterial resistance while the others showed they were. To add the effect of the Tetracycline would affect the growth of bacteria in the agar plates because we were seeing what was resistant to the antibiotic and what wasn’t. In the results it was an equal number of unaffected and affected species of bacteria.



Conclusions:
From doing this experiment, some bacteria are resistant to antibiotics and some are not. The results of the growth in the agar plates really showed that. This is important because of how truly important antibiotics are in our world due to how much they help us recover from sickness when we are at our worst. If more bacteria because resistant to more antibiotics that are very crucially to our survival if we ever get sick then that could possibly in the future kill mankind.