User:Yeem/BE.180 notes/Semester review
Programming in BE
Abstract concepts
- Abstraction hierarchy: high to low
- Decouling: breaking a complex system down
Comparison of Python and Biology
Python
- Functions
- Classes
- Inheritance
- Instances
- Passing data
Biology
- DNA
- TAATA, etc.
- Parts
- Basic biological function encoded in DNA
- Devices
- One or more parts that encode a human-defined function
- Systems
- One or more devices that perform a human-defined function
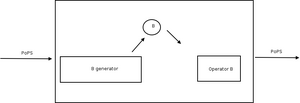
Recall that for an example of a genetic device, we spent a bunch of time making genetically encoded inverters, where one protein controls the operator of another protein. (see figure)
Protein kinases
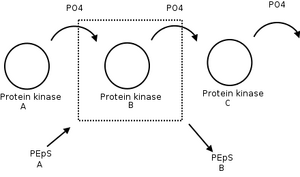
(figure) Protein kinase B phosphorylates protein kinase C to prevent it from phosphorylating something else. Is this a good device?
- No. Signal is specific. PEpS not interchangeable. Need common signal carrier.
How can you fix it?
- ???
Goal: can you describe a biological function that you can use without worrying about the details?
Redundancy
DNA to RNA to Protein
- ATCG to AUCG (one to one mapping for DNA to RNA)
- Mapping via a triplet code (RNA to protein)
- Looking at a piece of DNA, since there's 20 amino acids and 64 possible combinations of 3 nucleotides, there's redundancy
- Redundancy means flexibility in the specific DNA sequence
If you had the coding region of a particular open reading frame, and you found that there was a restriction site in the middle of it, you could remove it by specifying the same amino acid with a different nucleotide sequence.
Programming in time & space
- See other notes for more info
Explored two examples of early languages
- Growing point language ("GPL") developed by Coore
- "Crop circle language" language
- With "structured computer programming", you can
- Reuse functions
- Call function more than once
- Nest functions
- Reuse functions
- Even with differences, we've been able to map to biology
- We've made genetic devices that can respond to intermediate levels of an input signal
Sender/receiver
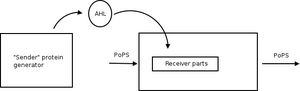
Band detectors: detect intermediate concentration
- i.e., AHL low, PoPS low
- AHL high, PoPs low
- AHL juuuuuust right, PoPS high