User:Sonia Chothani

About Me
<html> <body>
PhD student at Duke-NUS school of medicine, Singapore (Aug 2016 - Present)
Master of Science in Computational Biology, Carnegie Mellon University, Pittsburgh, PA (Aug 2012 - Dec 2013)
Bachelor of Technology in Biotechnology, Indian Institute of Technology Madras, India (Aug 2008 - July 2012)
LinkedIn : https://sg.linkedin.com/in/sonia-chothani-b192b515
Research Gate : https://www.researchgate.net/profile/Sonia_Chothani
Youtube link of an interview at CMU : https://youtu.be/O_ir9r-B7oc
Google Scholar : https://scholar.google.com.sg/citations?user=HdP8YrIAAAAJ&hl=en&oi=ao
</body>
</html>
Publications & Conference Proceedings
- Daniel Taliun, Sonia P. Chothani, Sebastian Schönherr, Lukas Forer, Michael Boehnke, Gonçalo R. Abecasis, Chaolong Wang; LASER server: ancestry tracing with genotypes or sequence reads. Bioinformatics (2017) btx075. doi: 10.1093/bioinformatics/btx075
- Raju Nagarajan, Sonia Pankaj Chothani, Chandrasekaran Ramakrishna, Masakazu Sekijima and M Michael Gromiha, Structure based approach for understanding organism specific recognition of protein-RNA complexes. Biology Direct (2015)
- Nilanjana Banerjee, Sonia Chothani, Lyndsay Harris, Nevenka Dimitrova. Identifying RNAseq-based coding-noncoding co-expression interactions in breast cancer. IEEE International Workshop on Genomic Signal Processing and Statistics - GENSIPS (2014)
- Nilanjana Banerjee, Sonia Chothani,et,al,. Abstract P4-04-02: RNA-seq reveals functional lncRNAs associated with estrogen-receptor status in breast cancer Cancer Research-American Association for Cancer Research (2014)
- Received acknowledgement in Chowdhury, S., et,al,. Structural and Logical Analysis of a Comprehensive Hedgehog Signaling Pathway to Identify Alternative Drug Targets for Glioma , Colon and Pancreatic Cancer, PLoS ONE (2013)
Patents
- Degenerate primer sets. Sonia CHOTHANI, Sitharthan Kamalakaran. (2016)
WO 2016193846 A2; PCT/EP2016/055195 [1]
- Methods of displaying the antimicrobial sensitivity of biological isolates. Sonia CHOTHANI Sitharthan Kamalakaran, Pramod MAYIGOWDA, Henry Lin (2016)
WO2016142890 A1; PCT/IB2016/051352 [2]
- Infection management and control. Sonia CHOTHANI Sitharthan Kamalakaran, Pramod MAYIGOWDA, Henry Lin (2016)
WO2016142493 A1; PCT/EP2016/055195 [3]
- Methods and systems to generate noncoding-coding gene co-expression networks. Yee Him Cheung Nilanjana Banerjee, Nevenka Dimitrova, Sonia CHOTHANI, Wilhelmus Franciscus Johannes Verhaegh (2016)
WO2016092444 A1; PCT/IB2015/059389 [4]
Coursework
Algorithms and Data Structures, Machine Learning, Mathematical Modeling, Biostatistics, Phylogenetics, Bioinformatics, Computational neuroscience, Principles of Imperative Computation, Molecular Biology, Structural Biology, Cell Biology, Evolution of Regulatory Genomics, Linear Algebra and Numerical Analysis, Calculus I,II, Probability
Skills
- Languages: C, C++, C0 (Safe ver. of C), BlueJ, Python, Perl, SQL, HTML
- Data Analysis: R, MATLAB
- Tools: vcftools, king, VerifyBamID, GotCloud, samtools, gatk, CellNetAnalyser, COBRA, Cytoscape
Awards
- Academic Achievement Fellowship, Carnegie Mellon University, Pittsburgh, PA (Aug ‘12 – May ‘14)
- Gold, Massachusetts Institute of Technology, Boston MA, International genetically Engineered machines World Championship (iGEM)), (Nov '11)
- Asia’s Best Safety Commendation and Asia’s Best Biobrick, iGEM Asia Regionals, Hong Kong University of Science and Technology, Hong Kong, (Oct ‘11)
- Gold, Massachusetts Institute of Technology, Boston MA, iGEM World Championship, (Nov ‘10)
- Awarded a summer research fellowship (IAS-NASI-INSA), Centre for Cellular and Molecular Biology, India (Summer ‘10)
- Secured an all India rank in top 1% in the Joint Entrance Examination’08 (IIT JEE) (amongst ~400,000 students) (June ‘08)
Research Projects
Areas of Interest: Bioinformatics, Machine Learning
- Research Officer, Genome Institute of Singapore, A*Star, Singapore [Guide : Dr. Chaolong Wang, Dr. Jianjun Liu, Computational and Systems Biology : May’16-May'17 ] Population structure, imputation and ancestry estimation in Asian populations
- Graduate Research Assistant, Dr. Schwartz Laboratory, Lane Center for Comptational Biology [Guide : Dr. Russel Schwartz, Carnegie Mellon University Duration : Dec’12-May'13 ] Marker based dimensionality reduction of aCGH/DNA copy number data prior to Principal Component Analysis and un-mixing to improve ability for phylogenetic inferences of tumor development
- Organism Specific Protein-RNA interactions [Guide : Dr. M. M. Gromiha, IIT Madras Duration : Dec’11-May'12] Understanding the recognition mechanism of Protein-RNA complexes has been a challenging task in molecular and computational biology. Computational biologists have studied non-redundant sets of these complexes and have formulated interesting facts. We would like to study whether a Protein-RNA recognition mechanism studied in only one organism can be generalized for all others or not. Expecting a varied behavior we would like to study how this change has evolved and observe a pattern over evolution.
- Reconstruction and Modelling of human specific Mitogen Activated Protein Kinase signalling pathways [Guide : Dr. Ramrup Sarkar, Mathematical Modeling and Computational Biology Laboratory, Centre for Cellular and Molecular Biology, Hyderabad, India Duration : May-July 2010] The initial phase of the work required a meticulous and patient reconstruction of a comprehensive MAPK signaling pathway (297 ‘species’ and 160 ‘reactions’) from various databases and literature. We used the Logical Steady State Analysis (LSSA) approach for modeling and analysis using the software CellNetAnalyzer (CNA). Further, a systematic perturbation study analysis to identify alternative pathways and optimal intervention points (targets) was carried out.
<html> <a href="http://openwetware.org/images/1/17/Report_Sonia_Chothani.pdf">Click here to view the report.</a>
<a href="http://openwetware.org/images/b/ba/Reconstruction_%26_Modeling_Of_MAP_Kinase_Signaling_Pathway.ppt">Click here for the presentation</a>
</html>
International Competitions/Conferences
<html>
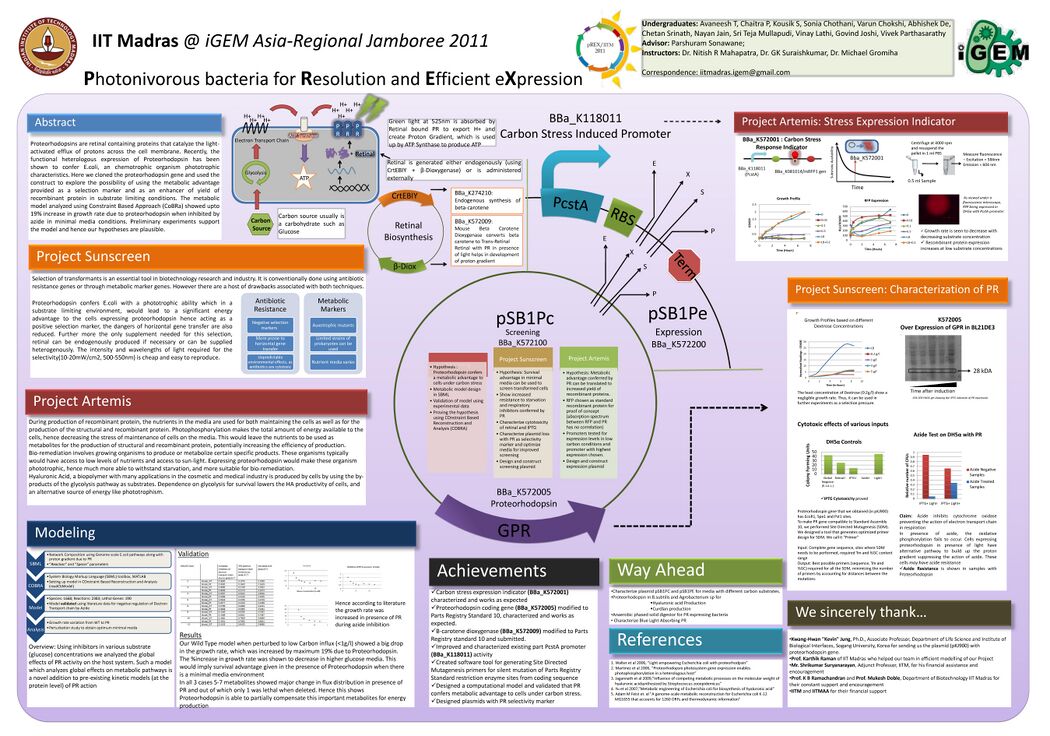
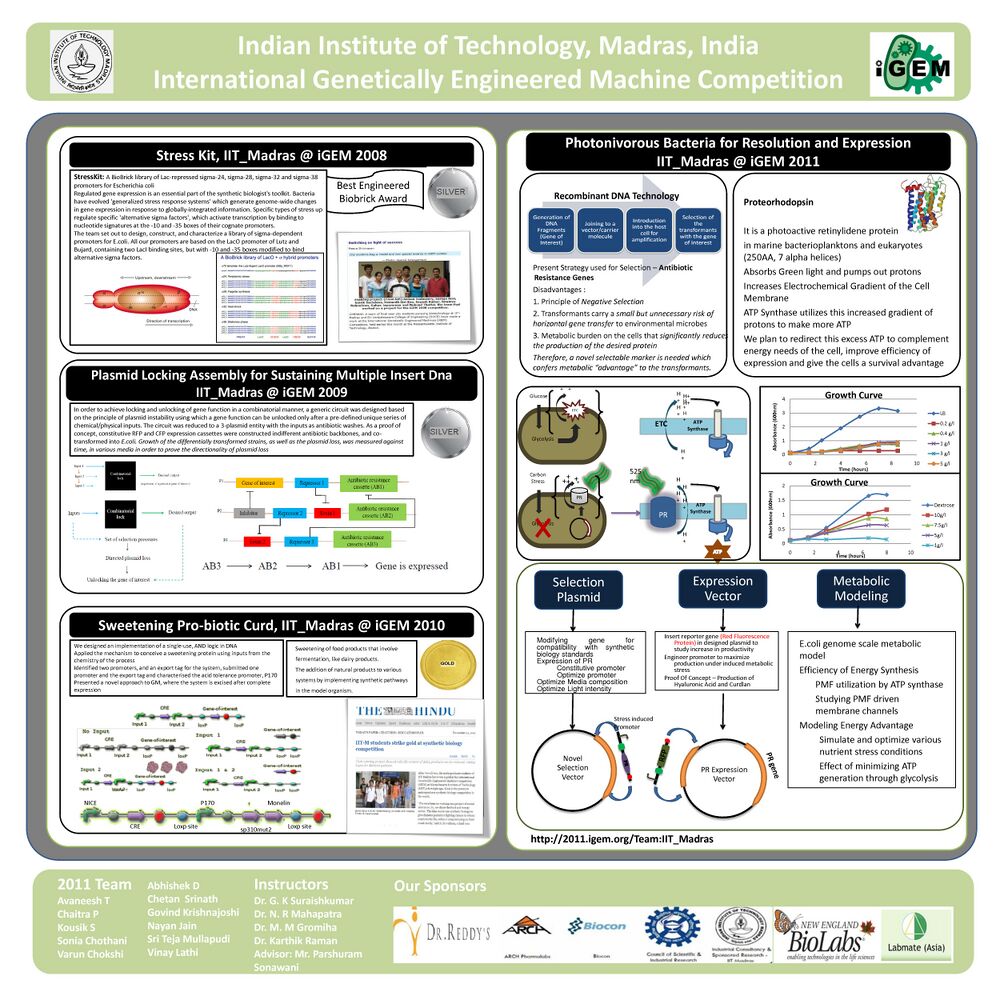
- International Genetically Engineered Machines (iGEM) 2011 Photonivorous bacteria for Resolution and Expression, IIT Madras, Jan-Nov’11
- A light based technique to screen transformants using a positive control An alternative to antibiotic screening technique for transformants. The screening plasmid here consists of a <a href="http://en.wikipedia.org/wiki/Proteorhodopsin">Proteorhodopsin(PR)</a> gene instead of the antibiotic resistance which in presence of light and <a href="http://en.wikipedia.org/wiki/Retinal">Retinal (A chromophore)</a> gives a proton gradient, this is used by ATP synthase to produce ATP. Hence giving an energy advantage allowing only the transformed cells to grow in a low carbon media.
- A light based technique to improve protein productivity Plasmid consisting of a Carbon Stress Promoter (pCSTA) and Proteorhodopsin, giving a survival advantage to the cells in low nutrient media. This would help improve the productivity of recomibnant proteins, biopolymers like Hyaluronic Acid and Curdlan.
- Mathematical Modeling and computational tools
- Developed an open-source online tool for site directed mutagenesis primer design
- An extensive Reconstruction and Flux Balance Analysis study of metabolic pathways in E.coli at the genome scale, considering 1668 metabolites and 2383 reactions and their respective stoichiometry matrices was carried out using a Constraint Based approach. Upto 30% increase in growth rate was observed due to Proteorhodopsin at high carbon stress conditions
Won Gold, Asia's Best Natural Biobrick, Asia's Best Safety Commendation
[Guide: Dr. Nitish Mahapatra, Dr. G.K Suraishkumar, Dr. Michael Gromiha]
[Asia Regionals, HKUST, Hong Kong (15th Oct’11)]
[World Championship, MIT, Boston (7th Nov’11)]
<a href="http://2011.igem.org/Team:IIT_Madras/Dry_lab/Modelling">Click here to see more details on the modelling.</a>
</html>

- <html>
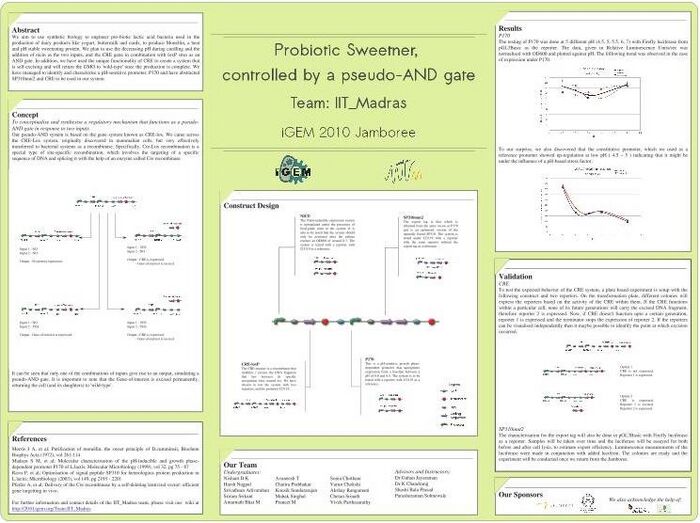
- International Genetically Engineered Machines (iGEM) 2010 Sweetening Pro-biotic Curd, Department of Biotechnology, IIT Madras, Jan-Nov’10
- Our innovations are two-fold: the idea of using a lactic acid bacteria (often used in the curdling of milk) to produce the sweetening protein(Monellin), and in using a regulatory mechanism that functions under a combination of constraints, like an AND gate
- Our regulatory mechanism combines the use of promoters that can be either induced or repressed by an external action, like providing Heat-shock, adding Niacin, or by the system falling below a certain pH <a href="http://2010.igem.org/Team:IIT_Madras">Click here to go to our media wiki</a>
- Philips Research centre North America [Briarcliff manor, NY, USA Jan'15-Jan’16]
- Monsanto Research Centre [Bangalore, India May-Jul’11]
- Molecular Cloning [Dr. Thillai Chidambaram, Dept: Molecular Cloning, Vector Technology and Protein Analytics (MVP), 7 weeks]
- Gained experience in numerous sophisticated Molecular Biology Techniques for E.coli and Agrobacterium
- Submitted 3 successful clones which were forwarded to the next vertical (Transformation, MRC, Bangalore)
- Quality check time for transformants was reduced from 1 day to 4 hrs by optimizing cell growth
- Optimized a new cloning technique which uses Overlap Extension fusion. It helps to clone more than 1 inserts at a time. This is being continued at MRC to reduce the cost and time
- Experimentally calculated optimum insert concentration for a successful ligation hence optimized time for PCR amplification of the insert. The time was reduced from 3.5 hrs to 1 hr
- Bioinformatics [Dr. Subodh Rawool, Computational Biology Function , 2 weeks]
- Learnt Perl, a scripting language-within 4 days and developed scripts to overcome the shortcomings of currently available methods for dynamic visualization of metabolic networks.
- Larsen & Toubro Infotech Ltd. [Powai, Mumbai, Jun-Jul’09]
- Trained for 8 years at Prerana Naratan Kala Kendra, Examining Body: Bharatiya Sangeet Samiti
- Master Diploma with “Distinction” in Bharatnatyam, ‘04
- Foundation Sr. with “Distinction” in Manipuri Naratan,‘97
- Awarded “NXG Hindu Chennai’s Best Dancer” at Campus Jive’10; Photograph and article featured in Hindu on 1st Sept’10
- Member of Institute dance team (‘08-’11) and a consistent winner at Solo dance, Lit-Soc: 6th in ‘08, 1st in ‘09, 2nd in ‘10
- Won 1 gold medal and 2 silver medals in a classical group dance competition at ISKCON (2002, 2003, 2004)
- Lead Actor in Sharavati hostel Short Film Making, LitSoc’11; Placed 3rd;
- Supporting Actor in “Variety”, an independent short film venture by MM Productions in 2011
- Social Service: Certificate of Appreciation for Alert India, Urban Leprosy control programme and National Association for Blind, India. Have also been involved in Neighborhood Development with National Service Scheme, India 2008-2009 Positions of Responsibility
- Choreo Coordinator, Saarang, IIT Madras, Sept’10-Aug’11
- Coordinated one of the biggest events at Saarang; Professional show, 8000 audience, 500 participants
- Conceptualized and conducted a new event, $treet$; 3000 audience, 40 participants, judged by: Int’l. hip hop crew
- Headed the Institute Dance team and coordinated the dance competitions all year round
- Co-Founder and Coordinator, SPArK, Dance Club, IIT Madras, ‘08-‘10
- Instrumental in organizing dance sessions on campus by renowned choreographers at 50% rates since freshman year
- Social Affairs Secretary, Sharavati Hostel, IIT Madras, Elected by 600 electorate, May’10-Apr’11
- Effectively handled a budget of INR 36,500 for conducting various cultural and social activities
- Led Sharavati hostel to place 3rd (out of 17) in Lit-Soc ‘10-‘11(Year round inter hostel literary and cultural competitions)
- Coordinator, Quality Management System, Shaastra (ISO 9001:2008 certified), IIT Madras, Mar’10-Feb’11
- Structured and implemented a formal procedure for improving the quality and management of Workshops at Shaastra
Won Gold
[Guide: Dr. Guhan Jayaraman]
[iGEM , MIT, Boston(7th Nov’10)]
<a href="http://openwetware.org/images/f/f5/11_02_10_Jamboree_presentation.pdf">Click here to view our presentation</a>
These were student initiated and organized project where we brainstorm towards a feasible, novel and socially significant idea. We also handled the Sponsorship, Finance and the laboratory equipment management. We generated a sponsorship of INR 8,00,000 to fund the projects and managed the laboratory requirements.

Industrial Experience
Offered a Research Consultant Position at Monsanto Research Centre, Bangalore
Understanding of Product Lifecycle Management and its Current Usage in Biotechnology Equipped the team with prospect maps and SWOT analysis of various clients
Co-Curricular & Extracurricular activities
Contact Info
Useful links
<html> <a href="http://www3.clustrmaps.com/counter/maps.php?url=http://openwetware.org/wiki/User:Sonia_Chothani" id="clustrMapsLink"><img src="http://www3.clustrmaps.com/stats/maps-no_clusters/openwetware.org-wiki-User-Sonia_Chothani-thumb.jpg"?url="http://www3.clustrmaps.com/user/a5bebfcf" style="border:0px;" alt="Locations of visitors to this page" title="Locations of visitors to this page" id="clustrMapsImg" onError="this.onError=null; this.src='http://www2.clustrmaps.com/images/clustrmaps-back-soon.jpg'; document.getElementById('clustrMapsLink').href='http://www2.clustrmaps.com'" /> </a> </html>