User:Siena Eve Romano/Notebook/Biology 210 at AU
Lab 7: “16S Sequence” 3/5/2015
Purpose: The purpose of this experiment is to identify the two colonies of bacteria from the experiment titled “Lab 3: Microbiology and Identifying Bacteria with DNA Sequences 02/04/2015” which were originally taken from the transect sample. This experiment aims to examine antibiotic resistance using tetracycline but mainly, to appreciate the importance of DNA sequences in identifying species. The hypothesis is that the 16S sequence will correlate with the previous research findings regarding colony morphology, gram stain, and motility studies to help identify the bacteria from the transect.
Materials and Methods: One bacteria from the nutrient agar plate and one from the tetracycline plate was selected. Then the a single colony of bacteria was transferred to 100 ul of water in a sterile tube and incubated at 100oC for 10 minutes in a heat block with the tubes floating in the water. Following the heat block the samples were centrifuged for 5 minutes at 13,400 rpm while 20 ul of primer/water mixture were added to a labeled PCR tube. Then this was mized to dissolve the PCR bead. Five ul of supernant were transferred from the centrifuged samples to the 16S PCR reaction before being placed in the PCR machine. The following week the PCR products were run on an agarose gel and analyzed next to a DNA ladder. Then the two PCR products that that formed a thick band beside the 16S gene marker on the DNA ladder were purified for sequencing. Four weeks later, the codes for identifying the bacteria were analyzed using a Blast website. The website relayed a DNA sequence used to identify the bacteria from the agar plates.
Data and Observations:
Above is an image of the agarose gel that was used to identify the 16s gene for sequencing. The visible bright pink bands indicated the PCR products that should have DNA purified.
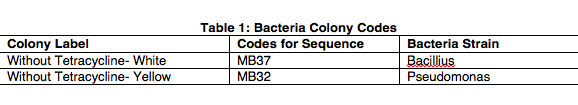
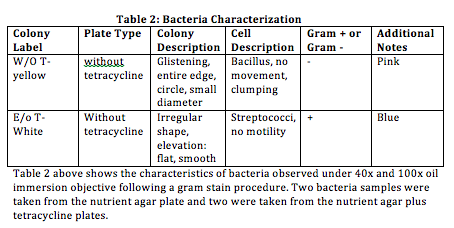
Table 1 represents the colonies that were purified for sequencing along with the resulting code from the blast website. Table 1 also shows the bacteria strain that was identified using the code and Blast website.
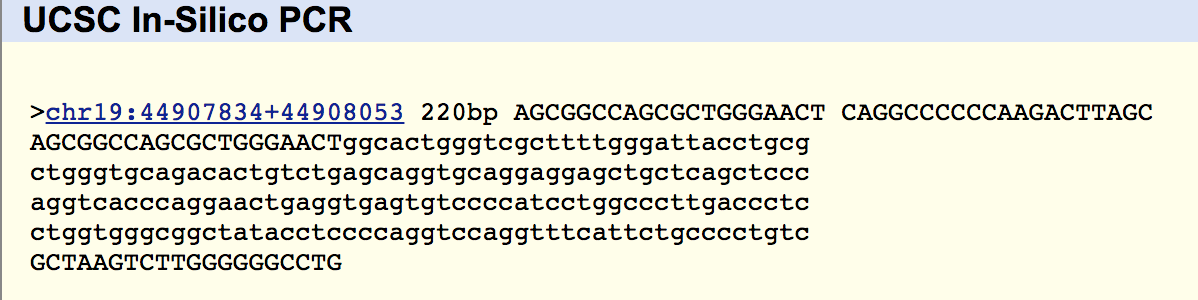
Sequence Code for ‘Without Tetracycline- White’:
NNNNNNNNNNNNNNCNANANTGCAGTCGAGCGAATGGATTAAGAGCTTGCT CTTATGAAGTTAGCGGCGGACGGGTGAGT AACACGTGGGTAACCTGCCCATAAGACTGGGATAACTCCGGGAAACCGGGGCTAATACCGGATAACATTTTGAACTGCAT GGTTCGAAATTGAAAGGCGGCTTCGGCTGTCACTTATGGATGGACCCGCGTCGCATTAGCTAGTTGGTGAGGTAACGGCT CACCAAGGCAACGATGCGTAGCCGACCTGAGAGGGTGATCGGCCACACTGGGACTGAGACACGGCCCAGACTCCTACGGG AGGCAGCAGTAGGGAATCTTCCGCAATGGACGAAAGTCTGACGGAGCAACGCCGCGTGAGTGATGAAGGCTTTCGGGTCG TAAAACTCTGTTGTTAGGGAAGAACAAGTGCTAGTTGAATAAGCTGGCACCTTGACGGTACCTAACCAGAAAGCCACGGC TAACTACGTGCCAGCAGCCGCGGTAATACGTAGGTGGCAAGCGTTATCCGGAATTATTGGGCGTAAAGCGCGCGCAGGTG GTTTCTTAAGTCTGATGTGAAAGCCCACGGCTCANCCGTGGAGGGTCATTGGAAACTGGGAGACTTGANTGCAGAAGAGG AAAGTGGAATTCCATGTGTAGCGGTGAAATGCGTAGAGATATGGAGGAACACCAGTGGCGAANGCGACTTTCTGGTCTGT AACTGACACTGAGGCGCGAAAGCGTGGGGAGCAAACAGGATTAGATACCCTGGTAGTCCACGCCGTAAACNATGAGTGCT AANTGTTAGAGGGTTTCCGCCCTTTANTGCTGAAGTTAACGCATTAAGCACTCCGCCTGGGGAGTACGGCCGCAAGGCTG AAACTCAAGGAATTGACNGGGGCCCGCACAAGCNGTGNANCATGTGGTTTAATTCNAANCAACGCGANNACCTTACCAGG TCNTGACNTCCTCTGANACCCTANAGATNNGGCTTNNNCNTCGGGANCAGAGTGACAGTGNNCATGNTGTCNTCANCTCN NGTCNTGAGNNGTTGGTNANNNCNNCAACNAGCNGCAACCCNNNATCTTNNTTGCCNNNNTNANNTNN
Sequence Code for ‘Without Tetracycline- Yellow’:
NNNNNNNNNNNNNNNNNNNNNTANNNTNCNNNCGGGGNNCGATAGNANCNTGCTNAAGATTCNNGANNGACGGGTGAGTA ATGCCAAGGAATCTACCTAGGGGTGGGGGACACCTTTTCGAACGGAAACCTCATACCGCATACATCGTACNGGANAGGGC AGGGGACCCTCCGGCCCTGCCCTAATAAATGAGCCCAGGTCGGATTAGCTGGTTGGTGAGGTAATGGCTCACCAAGGCNA CGATCCGTAGCTGGTCTGAAANGATGATCNGTCACCCTGGAACTGAGACACGGTCCNNACTCCTACGGGAGGCNGANTGG GGAATATGGGAGAATGGGTGATCGCNTNNNCCANCCCGCCCGCGTGTGTGANNAAGGTCTTCNGATCGTTTTGACNTTNA NANANGATNNAACGCGTGAAATTCNGANCGTACTGTTTTAAAATACACCNNNNAACACCGAGCCCTNNNCCNNGTNCCAC CANNNGGTNGTAACNTTNTNCGGATTTCTTTTGTTCANAGTTTCTGGACGTGGAGCNGNATANNTGNNGAAATATCTTAN TNTNANATCACCCTGCCANNNATANNGNCTGTCTTGANAAATTACAAGCGNCAGGATGAAGTAGTGTANCGGAGATCCGC NTAGATANTACTGANAACANNNATTGCGAAGGGAAGTCGCTATGTCNTAACGGGACCCCCGATAGCACAAN
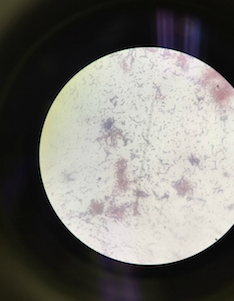
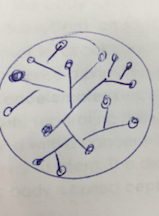
Above is an image through the microscope of the colony labeled 'Without Tetracycline- White' that was identified as Bacillius.
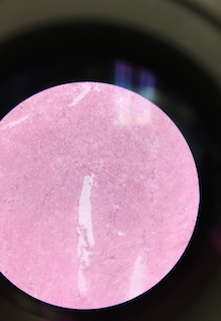
Above is an image through the microscope of the colony labeled 'Without Tetracycline- Yellow' that was identified as Pseudomonas.
- Table 2 represents our observations from "Lab 3: Microbiology and Identifying Bacteria with DNA Sequences 02/04/2015” (when studying the same bacteria identified during this lab)
Conclusion: After using 16S sequence analysis, the hypothesis proves to be true overall. However there were some discrepancies between the bacteria characteristics observed and those that were later proven to be true upon identification with the 16S sequence analysis. The data displayed in Table 1 largely correlates with the research findings from 02/04/2015 that are displayed in Table 2. Outside research indicates that Bacilius’ characteristics are in fact that it is irregular or rod shaped and gram positive (Cook & Sandeman, 2000). Yet we observed no motility when in fact Bacilius are known to display swarming movements (Cook & Sandeman, 2000). Similarly, Pseudomonas is known to be gram negative yet we observed the bacteria as circular when it is actually known to be rod shaped (Du et al., 2011). Furthermore, we observed no movement when in fact Pseudomonas is known to have flagella (Du et al., 2011). This experiment was effective in showing the importance of 16S sequencing for bacteria identification because we were clearly mistaken when making preliminary observations. In the future, it would be interesting to compare our observations under the microscope from a greater number of bacteria samples with results from 16S sequencing to see if our ability to observe particular features of bacteria improves.
Reference:
Cook, G. M., & Sandeman, R. M. (2000). Sources and characterisation of spore-forming bacteria in raw milk. Australian Journal of Dairy Technology, 55(3), 119. Retrieved from http://search.proquest.com/docview/199493576?accountid=8285
Du, L., Wang, S., Li, G., Wang, B., Jia, X., Zhao, Y., & Chen, Y. (2011). Biodegradation of malachite green by pseudomonas sp. strain DY1 under aerobic condition: Characteristics, degradation products, enzyme analysis and phytotoxicity. Ecotoxicology, 20(2), 438-46. doi:http://dx.doi.org/10.1007/s10646-011-0595-3
Lab 6: “Zebra Fish and Nicotine” 2/26/2015
Purpose: The objective of this lab was to study and compare embryonic development in an array of organisms and design an experiment to study how environmental conditions impact embryonic development. Zebrafish can be used as a model for comparison to larger vertebrae development. For this specific experiment, we used zebrafish to observe and understand the stages of embryonic development because they are opaque, accessible, and share many parallels in embryonic development with humans. In our experiment, one group of zebrafish was the control group in normal deerpark water and one group of zebrafish were treated with 5 mL of nicotine. The hypothesis was that zebrafish treated with nicotine will display defects in their development, resulting in poorer response to stimuli and mutations body length/structure.
Materials and Methods:Twenty zebrafish embryos were placed in 20 mLs deerpark water and another twenty zebrafish embryos were placed in 20 mLs of the same deerpark water with an additional 5 mL of nicotine in the water solution. The day after the initial set up (day 1) the number of dead eggs, living embryos still in egg cases, the number of hatchlings, and the number of dead hatchlings was observed. Additionally, the zebrafish embryos were observed on both day 1 and day 4 following their initial set up. Observations were recorded of how many were still alive, what specific features had developed over time, and any other features such as heartbeat. On day 4 10 mLs of water were removed while egg cases and dead embryos were removed as well. An additional 10 ML of water was added to each test dish and 2.5 mL of nicotine was added to the experimental group. Lastly, one drop of fish food was given to each test dish. The same steps from day 4 were repeated on day 7 as well as 11. In addition, on day 7 two larvae from the control and three from the test dish were removed from their dishes using a dropper. The larvae were then put in a tub with 1 drop of tricaine solution, 1 mL of water, and paraformaldehyde which was added by the TA. Another fixation following the same procedure took place on day 14 when final observations perviously listed were made again. The length of the tail, length of the entire larvae, and diameter of the eyes were measured.
Data and Observations:
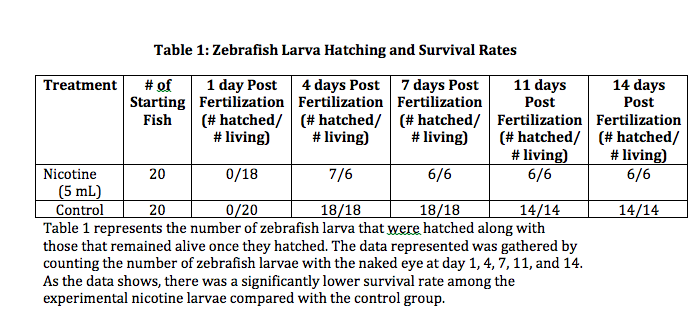
Conclusion: Although the experiment is not complete the results indicate the hypothesis may be proven true upon completion. Although both zebrafish expressed some movement when the dish was tapped (stimuli), there were noticeable differences between the control and experimental groups. The slower movement, and sometimes no movement at all, displayed by the zebrafish exposed to 5 mL of nicotine already shows the negative effects of nicotine on response to stimuli. As this was the most noticeable difference at 96 hours, it will be interesting to see how the zebra fish continue to develop in regards to their ability to respond to stimuli. In future observations, it will also be interesting to better observe the heartbeat of the fish. It will also be beneficial to compare body structures as no differences were observed at either 24 or 96 hours.
2.24.15
Very good notebook entry.
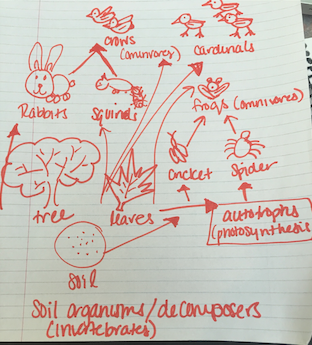
Nice food web
SK
Lab 5: "Invertebrates" 02/18/2014
Purpose: The objective of lab 5 was to better understand how simple systems were able to evolve into more complex systems by studying the importance of invertebrates. Through observation of acoelomates, pseudocoelmoates, and coelomates the experiment also serves to show the simplicity of these organisms as well as how body structure and movement correlate between invertebrates species. Furthermore, this lab involved observing soil invertebrates from our own transect to better identify, classify, and understand invertebrates. The hypothesis for this experiment is that by observing the soil invertebrates from our own transect the diversity, characteristics, multitude within ecosystems, and importance of invertebrates will be appreciated.
Materials and Methods: In order to prepare the Berlese funnel, one week ago about 500g of leaf litter was placed into one bag for which a handful of (dead) leaves along with small amount of dirt was obtained from edge of the transect with softer soil and a variety of dead leaves. Then 25 mL of the 50:50 ethanol/water solution was poured into a flask. A piece of screening material was placed into the bottom of the funnel, and the sides were taped securely so that the leaf litter could not fall into the preservative. Then, the funnel was placed into the neck of the square-sided bottle. Following that, the leaf litter sample was put into the top of the funnel, and the entire Berlese Funnel was covered in foil. A 40 watt lamp was set in place above the funnel with an incandescent bulb 2 inches from the top of the leaf litter. The lighted set up and Berlese funnel remained on the lab bench for one week.
First the Acoelomates, Pseudocoelmoates, and Coelomates were studied under a microscope during which their movement and body structure were observed. Then, the Berlese setup was broken down and 10 mLs of the preservative solution containing liquid and organisms was transferred into each of two Petri dishes in order to be examined under the dissecting microscope. Three Arthropoda invertebrates from the Berlese Funnel were then identified using dichotomous keys while their characteristics including length, number in sample, and description was recorded in Table 1. Finally, vertebrates who may inhabit or pass through the transect were considered along with the biotic and abiotic characteristics of the transect that would benefit each species. The five vertebrates were classified and a food web was created based upon the observed organisms in the transect.
Data and Observations:
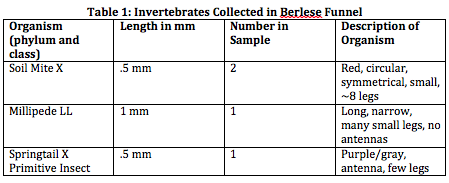
As displayed in Table 1 above, the size range of the organisms we measured was from 0.5 mm in length to 1 mm in length, a 0.5 mm difference. The largest organism we discovered was the Millipede LL, and the smallest organisms were the Springtail X Primitive Insect as well as the Soil Mite X. The organisms that were most common was the Soil Mite X.
 This image is the invertebrate identified as “Soil Mite X” as observed through the microscope.
This image is the invertebrate identified as “Soil Mite X” as observed through the microscope.
 This image is the invertebrate identified as “Millipede LL” as observed through the microscope.
This image is the invertebrate identified as “Millipede LL” as observed through the microscope.
 This image is the invertebrate identified as “Springtail X Primitive Insect” as observed through the microscope.
This image is the invertebrate identified as “Springtail X Primitive Insect” as observed through the microscope.
The images above are of the acoelomate, pseudocoelomates, and coelomates studied. The first photo is planaria, the second is a nematode, and the third is an earthworm. They all had extremely fluid and “squiggily” movements. The movement of the earthworm was similar to an accordion where the folds would allow the organism to move.
Conclusion:
The hypothesis proposed at the beginning of the experiment proved true as we were able to appreciate the diversity, characteristics, multitude within ecosystems, and importance of invertebrates through careful observation. In the future, it would be beneficial to continue this experiment by studying the invertebrates present in transects from other locations with potentially different environments.
Some vertebrates that might inhabit the transect:
1. Rabbits (Chordata, Mammalia, Lagomorpha, Leporidae, Oryctolagus, Cuniculus)
Biotic Advantage: Grass, grass seeds, or other plants for food
Abiotic Advantage: Water puddles, soil, large rocks for shelter
2. Crows (Chordata, Aves, Passeriformes, Corvidae, Corvus, Corvus brachyrhynchos)
Biotic Advantage: Invertebrates for food, such as worms
Abiotic Advantage: large flat rocks, water puddles
3. Cardinals (Chordata, Aves, Passeriformes, Passeri, Cardinlidae, Periporphyrus, Cardinalis)
Biotic Advantage: Invertebrates for food, such as worms
Abiotic Advantage: soil, water puddles
4. Common Frog (Chordata, Amphibia, Anura, Ranidae, Rana, Rana)
Biotic Advantage: Presence of bugs, worms, soil animals who help in breaking down and recycling decomposing plant/animal material
Abiotic Advantage: soil, drain, small rocks, water puddles
5. Squirrels (Chordata, Mammalia, Rodentia, Sciuridae, Sciurus, Sciurus carolinensis)
Biotic Advantage: seeds, green plants (grass) for food
Abiotic Advantage: Water puddles, soil, large flat stones
The image above is of a food web drawn using the organisms that were observed as well as those that were speculated to inhabit the transect. The organisms represent community as they are the many different species living in the transect as the same time (currently). Furthermore, there are few number of cardinals compared to invertebrates, for example, which indicates the environment can only sustain a few cardinals (carrying capacity) and many more invertebrates. Lastly, the organisms are located at various tropic levels as indicated by their position on the food web.
2.19.15 Excellent entry. Data section should also include some text. SK
Lab 4: "Plante and Fungi" 02/11/2015
Purpose: The main purpose of this lab was to observe the characteristics and diversity of plants while also being aware of the function and importance of Fungi. In order to understand the characteristics/diversity of plants, five plants were collected from the transect and studied based on specialized structures, vascularization, reproductive mechanisms, and the general size/shape. Additionally, Fungi were identified from looking at the slide sample's characteristics under the microscope. The hypothesis for this experiment is that because of the vast diversity of plants that exist, all of the matter collected in the transect would display unique characteristics in terms of vascularization, size/shape, special characteristics, and reproductive elements. If the plants observed are either monocot or dicot then they will display different reproductive processes or different phenotypes such as height accordingly, and plant diversity will be shown in the transect.
Materials and Methods: In order to acquire the five plants from the transect, zip-lock bags were obtained and we proceeded to the transect. About 500g of this leaf litter was placed into one bag to be used for the Berlese funnel for next lab for which a handful of (dead) leaves and small amount of dirt was obtained from edge of the transect with softer soil and a variety of dead leaves. Then, five representative samples were collected from the transect: tall grass, a red bush, a flowering bush, a cattail, and moss. The characteristics of a Bryophyte moss, Polytrichum, and an Angiosperm, Lily were compared based on observations with the microscope and the Lilly was dissected. The five samples from the transect were also observed under the microscope where all seven of the samples were examined in terms of vascularization, specialized structures, and reproduction. The observations from the five plants were described in Table 1 where it was also determined whether the seeds were monocot or dicot. A sample of fungi, Rhizopus stolonifer, was also observed under the microscope. To prepare the Berlese Funnel for collecting invertebrates in the next lab, 25 mL of the 50:50 ethanol/water solution was poured into a flask. A piece of screening material was placed into the bottom of the funnel, and the sides were taped securely so that the leaf litter could not fall into the preservative. Then, the funnel was placed into the neck of the square-sided bottle. Following that, the leaf litter sample was put into the top of the funnel, and the entire Berlese Funnel was covered in foil. A 40 watt lamp was set in place above the funnel with an incandescent bulb 2 inches from the top of the leaf litter. The lighted set up and Berlese funnel will remain on the lab bench for one week.
Data and Observations:
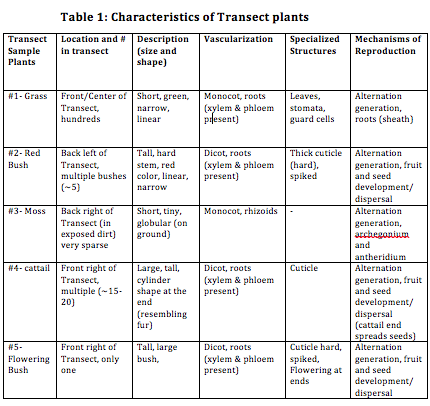
Table 1 above displays the samples obtained from the transect, their location in the transect, and other aspects of plant characterization studied during this lab.
The photo above represents the sample #1 labeled "Grass" in Table 1
The photo above represents the sample #2 labeled "Red Bush" in Table 1
The photo above represents the sample #3 labeled "Moss" in Table 1
The photo above represents the sample #4 labeled "Cattail" in Table 1
The photo above represents the sample #5 labeled "Flowering Bush" in Table 1
The photo above represents the Zygomycetes fungi, Rhizopus stolonifer observed
Conclusion: As was predicted, the grass, red bush, flowering bush, moss, and cattail samples all displayed different shape, sizes, vascularization, and specialized structures as well as reproductive mechanisms. In this way, the experiment was successful in addressing its objectives and the hypothesis was correct. We were also able to learn more about fungi by identifying Ascomyocta based on its black spores observed under the microscope. Fungi sporangi are a cell that contains spores where sporangium can be the site of meiosis or mitosis. They include hyphae that grow up and form black, globelike structures and are significant because the sporangium can enclose its spores until they are mature/ready to disperse. However, one thing I found somewhat surprising during this experiment was that most of the samples besides the grass were dicot in terms of vascularization. Monocots have one seed leaf while dicots contain two seed leaves. The other four plant samples were identified as dicots due to two cotyledon, the broad leaf or network of veins, and vascular bundles in a ring observed under the compound microscope.
The following is of a sample of a fungus identified during the experiment. It is clearly a fungus because of the black spores and sporangia linked by grey lines which are mycelium.
This image above is a drawing of the fungus identified as Ascomyocta.
2.11.15 Excellent lab book entry. I love the fact that you have included some outside sources and images. Very thorough and clear SK
Lab 3: "Microbiology and Identifying Bacteria with DNA Sequences" 02/04/2015
Purpose: The purpose of this experiment is to identify the bacteria originally taken from the and study the characteristics of such bacteria. In doing so this experiment aims to examine antibiotic resistance using tetracycline and understand the importance of DNA sequences in identifying species. After considering the inhibiting nature of tetracycline, it is likely that the agar plates with tetracycline will have very few bacterial growths.
Materials and Methods: In order to prepare a gram stain, a loop was sterilized over a flame and used to scrape up a tiny amount of an isolated growth from the surface of the agar before it was added/mixed to the water on a labeled slide. Then a red sharpie was used to circle the plate under where the sample was taken. With the bacterial smear side up, the air dried side of the agar was then heat fixed by exposing it to fire as it passed through the flame three times. The bacterial smear was covered with crystal violet using a staining tray. After exactly 1 minute the stain was rinsed off from a wash bottle filled with water. Following, the bacterial smear was covered with Gram’s iodine mordant for 1 minute before being rinsed again. The bacterial smear was flooded with 95% alcohol for 10-20 seconds and rinsed gently until decolorization occurred because the solvent flowed colorlessly from the slide. Then the smear was covered with safranin stain for 30 seconds before being rinsed with the wash bottle of water. Lastly, the excess water was blotted from the slide with a kimwipe before air drying and being observed under a low magnification, a 40x, and a 100x oil immersion objective.
So far for the DNA isolation and PCR amplification, one bacteria from a nutrient agar plate and the tetracycline plate was selected. Then the a single colony of bacteria was transferred to 100 ul of water in a sterile tube and incubated at 100oC for 10 minutes in a heat block with the tubes floating in the water. Following the heat block the samples were centrifuged for 5 minutes at 13,400 rpm while 20 ul of primer/water mixture were added to a labeled PCR tube. Then this was mized to dissolve the PCR bead. Five ul of supernant were transferred from the centrifuged samples to the 16S PCR reaction before being placed in the PCR machine.
Above is an example of a colony morphology used to characterize and help identify the bacteria present from http://www.microbiologyonline.org.uk/teachers/observing-microbes/observing-bacteria-in-a-petri-dish
The picture above was taken of our actual gram stain preparation.
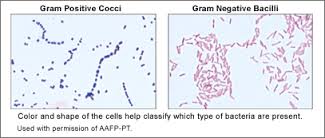
This image is an example of how gram stains were used to characterize and help identify the bacteria present from http://labtestsonline.org/understanding/analytes/gram-stain/tab/test/
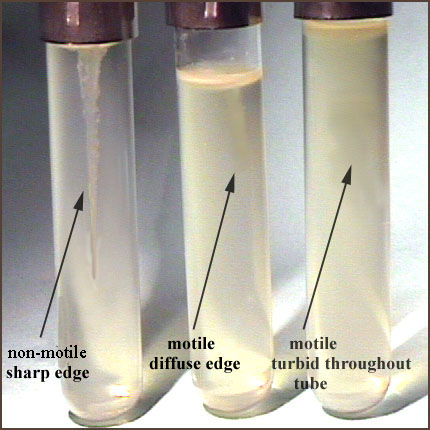
The picture above displays a motility study used for studying the characteristics of bacteria from http://faculty.lacitycollege.edu/hicksdr/motility.htm
Data and Observations:
Hay Infusion Culture: no smell, evaporation, all materials settled at the bottom of jar
It is likely the jar changes from week to week because bacteria are growing at various rates while other biotic materials are breaking down at various rates simultaneously. Both of these factors could be responsible for altering the appearance and smell of the Hay Infusion Culture.
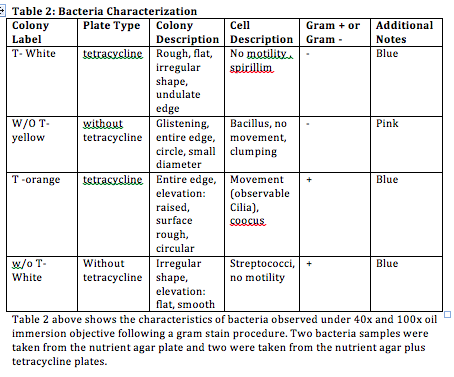
Above is an image of the agar plates after 1 week where the left column has the antibiotic.
Above is an image through the microscope of the colony labeled 'Without Tetracycline- Yellow' on Table 2
Above is an image through the microscope of the colony labeled 'With Tetracycline- Orange' on Table 2 where motility/cilia were readily observed.
Above is an image through the microscope of the colony labeled 'Without Tetracycline- White' on Table 2
Above is an image through the microscope of the colony labeled 'With Tetracycline- White' on Table 2
Conclusion:Based on the data displayed in Tables 1 and 2, the hypothesis made at the beginning of the experiment is correct where fewer bacteria colonies grew on the plates with tetracycline. The most noticeable differences in the colony types on the plates with antibiotic is that they had large growths of fungi and only a few large, orange bacteria colonies whereas the plates without antibiotic was covered more diverse colonies of bacteria (no fungi). As indicated by table 1, in the presence of tetracycline the total number of bacteria is significantly decreased as compared to the plates without tetracycline. The plates with the antibiotic allow fungi and only select bacteria with DNA that allows them to be resistant to tetracycline to flourish. Tetracycline is a protein inhibitor whose mechanism of action is by stopping the binding of aminoacyl-tRNA to the mRNA-ribosome complex. More specifically, tetracycline binds to microbial ribosomes’ 30S subunit. In the future, the identities of the bacteria will finally be determined using DNA isolation and PCR amplification.
Reference:
Chopra, Ian, and Marilyn Roberts. "Tetracycline Antibiotics: Mode of Action, Applications, Molecular Biology, and Epidemiology of Bacterial Resistance." Microbiology and Molecular Biology Reviews. American Society for Microbiology, June 2001. Web. 03 Feb. 2015. <http://www.ncbi.nlm.nih.gov/pmc/articles/PMC99026/>.
2.5.15 Excellent notebook entry. Well organized, lots of detailed observations. Pictures are a bit large. Try saving images as smaller sized files and then uploading them. SK
Lab 2: “Identifying Algae and Protists” 1/22/2015 Italic text Purpose: The purpose of this experiment was to study the protists that were living in the Hay Infusion Culture as well as identify those protists/algae using a dichotomous key based on their observed characteristics. After setting up a Hay Infusion the experiment intended to better understand the species of organisms living in a particular niche on the American University campus and how they interact with their environment. The hypothesis for this experiment is that if samples are taken from different sections of the Hay Infusion with different environmental properties, then the organisms that live in each will have different characteristics as well based on which section they are obtained from.
Materials and Methods: One week prior to the experiment a Hay Infusion Culture was set up by filling a 50 mL conical tube with a soil and ground vegetation sample from the groups designated 20 meter by 20 meter transect. Then 12 grams of the soil/ground vegetation was placed in a plastic jar with 500 mLs of deerpark water. After 0.1 gm of dried milk was added to the jar and a lid was secured, the jar’s contents were mixed thoroughly. The lid was removed and the jar was left in lab for a week. Upon returning to lab initial observations of the culture were made followed by samples being taken for microscopic observation. One sample was taken from the niche at the top of the liquid while the second was taken from the bottom where the soil/ground vegetation settled. Then a wet mount from the two different niches was made and studied under a microscope during which the sizes, shape, and method of motility of the protists were measured/recorded with the ocular micrometer. Lastly, the protists/algae present were determined using a dichotomous key.
Data and Observations:
Initial Observations of Hay Infusion Culture: extremely foul smell, solid contents (especially soil) settled to the bottom, some water evaporated, no observable indication of life on top of the liquid
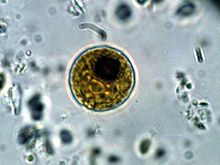
Sample from Top of Hay Infusion Culture: Top of culture was mainly opaque liquid and free of observable particles, Very little abiotic particles to naked eye and when under the microscope, No other observable protists, At first less # of algae (only 2) but over time more emerged (4), Algae were moving around rapidly (motile), Unicellular, Algae were small with a circular shape (12 um, 13 um, 15 um, 20 um), Identified as Chlamydomonas based on size and shape
 The photo above is of a drawing of the algae observed under the microscope, which was identified as Chlamydomonas.
The photo above is of a drawing of the algae observed under the microscope, which was identified as Chlamydomonas.
The photo above is an image of an actual Chlamydomonas under a microscope taken from http://en.wikipedia.org/wiki/Chlamydomonas#mediaviewer/File:Chlamydomonas_EPA.jpg
Sample from Bottom of Hay Infusion Culture: Bottom of culture had settled soil and vegetation, Dirt clumps and other abiotic particles in sample observable to naked eye & under microscope, 4 protists were observed, Protists were moving around rapidly (motile), Protists were larger with an oval shape (~50 um), Observed tiny circular organelles inside cell, Identified as Colpidium, a protozoa,
 The photo above is an image of the ocular view of the microscope when steadying the bottom of the Hay Infusion Culture in which one can see the protozoa and other abiotic materials
The photo above is an image of the ocular view of the microscope when steadying the bottom of the Hay Infusion Culture in which one can see the protozoa and other abiotic materials
 The photo above is of a drawing of the protist observed under the microscope along with some of its organelles, which was identified as Colpidium.
The photo above is of a drawing of the protist observed under the microscope along with some of its organelles, which was identified as Colpidium.
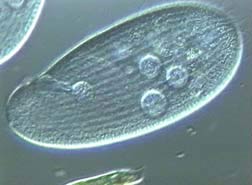 The photo above is an actual image of a Colpidium taken from https://www.ebiomedia.com/colpidium-rabbit-of-the-microcosm.html
The photo above is an actual image of a Colpidium taken from https://www.ebiomedia.com/colpidium-rabbit-of-the-microcosm.html
Conclusions and Future directions: The experimental results supported the hypothesis because organisms differed based on the section/environment of the hay infusion culture from which they were obtained. However, there was only one identifiable organism in each of the two sample sections of the hay infusion culture. The Chlamydomonas was at the top of the culture likely because algae need to be closer to the sunlight to undergo photosynthesis effectively via their light-absorbing chloroplasts. However, the protozoa, Colpidium were living at the bottom of the culture. This is likely because organisms that are closer to the bottom of the culture are closer to the plants and therefore consume nutrients from breaking down the plants as they are catabolic. Also, it is likely the protozoa were closer to the plants in order to meet their oxygen needs as plants release oxygen during photosynthesis. According to Freeman on page 2, organisms such as Chlamydomonas must meet all the needs of life. Chlamydomonas contains all its genetic information in its single cell and acquires the energy it needs to function from photosynthesis. Furthermore, there is diversity in the species such as the size of the organism that allows for its continual evolution. If the Hay Infusion Culture grew for another two months I predict the plant materials would decompose so maybe there would be less observable protozoa because they need to feed off the plant nutrients (selective pressure) and more species of algae or fungi protists instead. Another selective pressure reducing the number of protozoa would be a decline in oxygen as plants die. However, in order to continue this study of protists in their environment, the Hay Infusion Culture could be left for months and samples could be taken each week to record any difference in the species of organisms.