User:Perry/Spring 2007
2/12/07
dissolved 500mg ampicillin salt into 10ml water to prepare 50mg/ml ampicillin stock; aliquotted 1ml into tubes, left in -20dC.
overnight 1ml LB-amp cultures of JLO159, W, H, S, ~4:30pm.
2/13/07
took overnight cultures out ~7:30pm. diluted 1:10, measured OD600.
159- .335, W- .245, H- .231, S- .279
made liquid cultures of 5ml LB-amp + 250ul overnight cultures. incubated 3h, added 5mM IPTG (53ul 0.5M), incubated 2h, diluted 1:10, measured OD600.
159- .106, W- .044, H- .023, S- .057
Aliquotted 1.5mL of H into each tube and proportional volumes of the rest. aliquotted volumes into tubes. centrifuged 8000rpm, 3min. aspirated. reconstituted in 10ul streptavidin aptamer, 10ul B/F hybrid, or 10ul B0/F hybrid.
4uM streptavidin aptamer: 48ul PBS + 2ul streptavidin aptamer (100uM)
4uM B/F hybrid: 46ul PBS + 2ul Bside oligo (100uM) + 2ul Iside oligo (100uM)
4uM B0/F hybrid: 46ul PBS + 2ul DispI oligo (100uM0 + 2ul Iside oligo (100uM)
Wrapped tubes in a napkin, rotated on rotisserie at room temperature for 1h. washed in PBS 3X, reconstituted in 50ul PBS, loaded into clear 96-well plate.
2/28/07
I'm going to shift the project away from streptavidin for now, until I learn more about membrane fractionation or until I get a full streptavidin clone from Takeshi Sano. I'm going towards expressing PDZ repeats on the surface.
Prepared 150ml cultures of JLO66 and JLO159.
3/1/07
Hi-Speed Midiprep of JLO66 and JLO159.
3/2/07
Received primers PDZ1F, PDZ2F, PDZ1RS, PDZ2RS. Dissolved in (10x#nmol)ul for 100uM. Diluted 1:10 for 10uM used in PCR.
Received F11/Na template from Alain; nanodropped at ~10ng/ul. Made a 1:50 dilution.
Prepared six 50ul PCR reactions, adding the 1ul of each of the following to 47ul Platinum PCR Supermix.
PDZ1: F11, PDZ1F, PDZ1RS
PDZ2: F11, PDZ2F, PDZ2RS
PDZ1+2: F11, PDZ1F, PDZ2RS
PDZ1/2/1+2 (1:50): same, using 1:50 F11.
Incubated 95dC 10min, (95dC 30sec, 55dC 30sec, 72dC 60sec)x30, 72dC 10min.
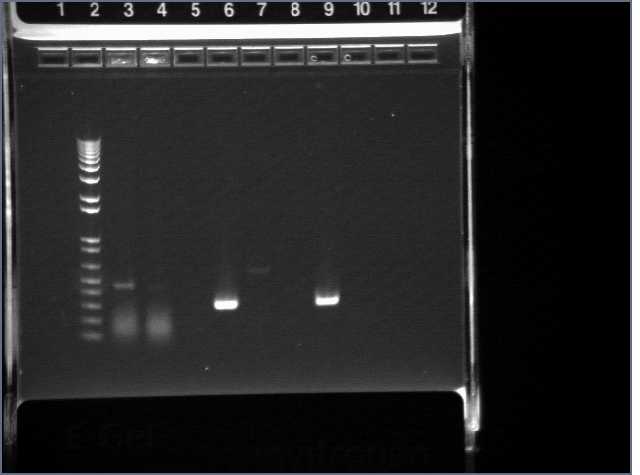
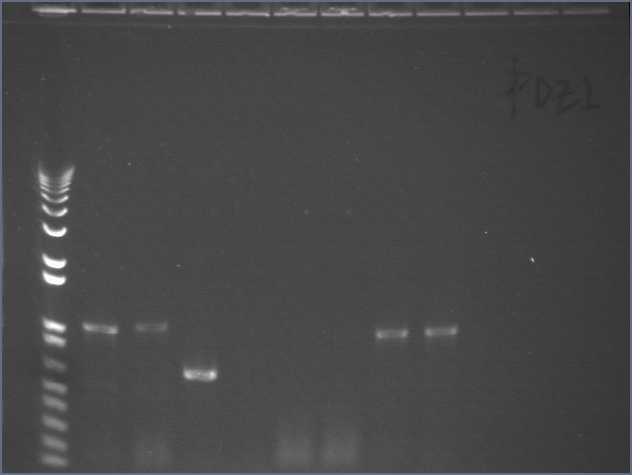
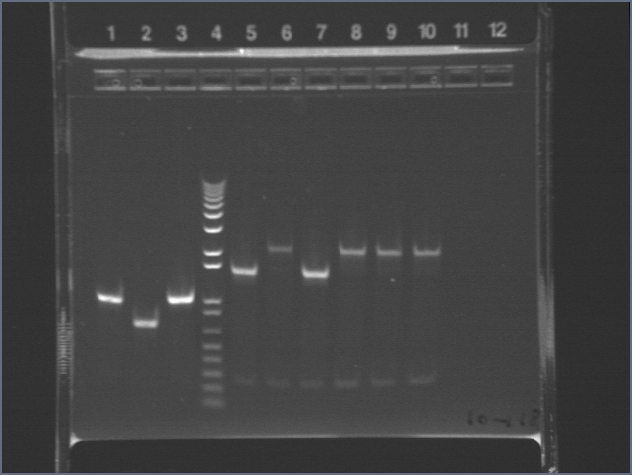
Walker ran 10ul of each PCR reaction on an e-gel.
Lane 1: 1kb+
Lane 5: PDZ1
Lane 6: PDZ2
Lane 7: PDZ1+2
Lane 8: PDZ1 (1:50)
Lane : PDZ2 (1:50)
Lane 10: PDZ1+2 (1:50)
3/5/07
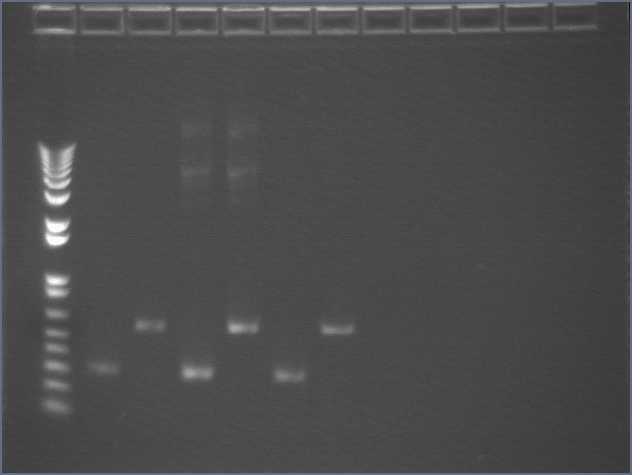
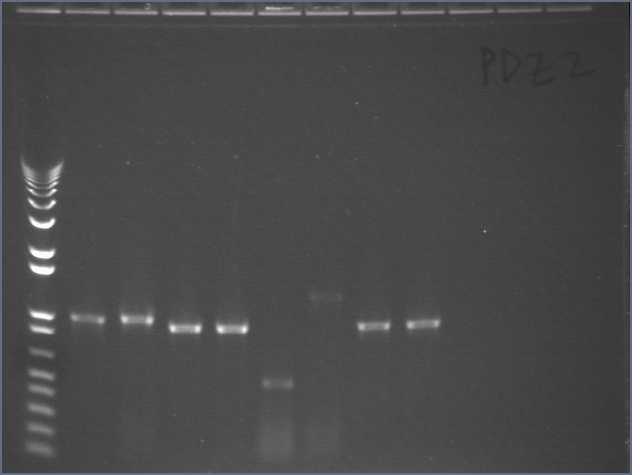
Repeated PDZ1 and PDZ1+2 PCRs (50ul) with F11, DC1-2/Na (~200ng/ul), and DC1-2/Na(1:20). Changed annealing temperature to 45dC. Ran 10ul in an e-gel.
Lane 1: 1kb plus
Lanes 2, 3: F11 PDZ1 and PDZ1+2
Lanes 4, 5: DC1-2 PDZ1 and PDZ1+2
Lanes 6, 7: DZ1-2 (1:20) PDZ1 and PDZ1+2
Performed PCR purification with yesterday's F11 PDZ2 and today's F11 PDZ1 and PDZ1+2. Eluted in 30ul water.
Performed Topo cloning/transformations with PCR-purified PDZ1, 2, 1+2.
3/6/07
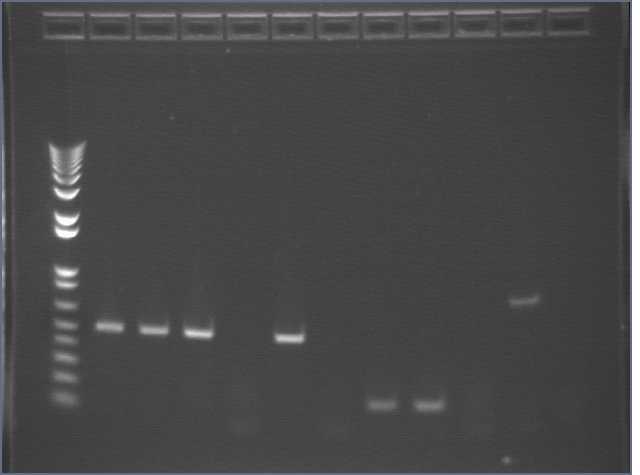
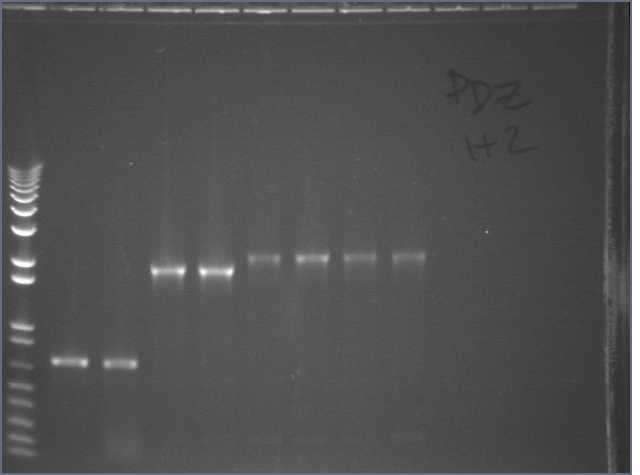
Made streaks and M13F/R colony PCRs of PDZ1 (1, 2, 3, 4), PDZ2 (1, 2, 3, 4), and PDZ1+2 (2, 3, 4).
Lane 1: 1kb+
Lanes 2-5: PDZ1 colony PCR #1, 2, 3, 4
Lanes 6-9: PDZ2 colony PCR #1, 2, 3, 4
Lanes 10-12: PDZ1+2 colony PCR #2, 3, 4
Prepared two 3ml cultures of PDZ1 #1, PDZ2#1, PDZ1+2 #3.
3/7/07
Performed minipreps, sent out sequencing, prepared XbaI/PstI digests of PDZ1, 2, 1+2. also, SpeI/PstI digests of JLO66 and JLO159.
3/8/07
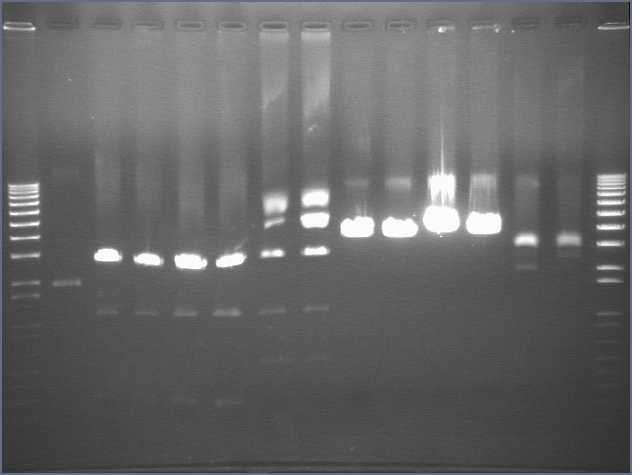
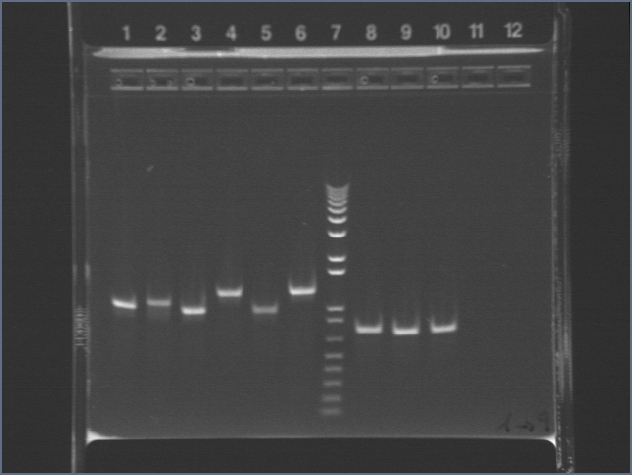
Ran 1% 100ml agarose gel (used TAE, and 3ul ethidium bromide), 140V, 50 minutes.
Lane 1: 1kb+
(Lane 2: undigested LppOmpA/pET29b)
Lanes 3, 4: PDZ1 X/P digest
Lanes 5, 6: PDZ2 X/P digest
Lanes 7, 8: PDZ1+2 X/P digest
Lanes 9, 10: JLO66 S/P digest
Lanes 11, 12: JLO159 S/P digest
(Lanes 13, 14: Lpp-OmpA/pET29b NheI/PstI digest)
Cut out 300bp band from Lanes 3 and 5, 500bp band from lane 7, and 4kb bands from Lanes 9 and 11. Performed Qiaquick gel extraction, added the sodium acetate step, eluted in 50dC water, 30ul. nanodropped then and got all 15-20 ng/ul, except for JLO159, ~35ng/ul.
Cut out same bands from Lanes 4, 6, 8, 10, 12, performed Montage gel extraction, and got all 2-3ng/ul.
Performed Quick Ligation with 4ul vector and 8ul insert, plus 12ul ligase buffer and 1ul quick ligase. Transformed into 18ul Top10 cells.
- JLO66 and PDZ1
- JLO66 and PDZ2
- JLO66 and PDZ1+2
- JLO159 and PDZ1
- JLO159 and PDZ2
- JLO159 adn PDZ1+2
3/9/07
Ug, none of the ligations worked. Onto the Clonewell!
3/11/07
Prepared 150ml LBamp cultures of PDZ1, 2, 1+2. Incubated 200rpm, 37dC, 8pm.
3/12/07
Performed midipreps. Set up 50ul, 2ug XbaI/PstI digests of PDZ1, 2, 1+2, and SpeI/PstI of JLO66, JLO159.
3/13/07
Ran digests on Clonewell e-gel, split each digest into 2 wells, 25ul each.
Nanodropping collections all yielded ~20ng/ul; however, nanodropping a collection from a blank lane yielded ~16ng/ul. Running 20ul of JLO66, JLO159 collections on a 1.2% e-gel yielded strong, sharp bands. Tried out ligation/transformations 2.5:7.5 vector:insert.
Set up new digests, same as yesterday.
3/14/07
Ligation/transformations failed. Retried Clonewell e-gel. Vacufuged samples to up concentration to about 50ng/ul.
3/15/07
Got colonies! Variation in colony sizes, possibly due to the fact that I transformed in Top10 cells, not Top10F'.
Prepared VF2/VR (200nM) PCRs and plate streaks from two big colonies (#1 and 2) and two smaller colonies (#3 and 4). Incubated 1m-1m-1m30s.
3/16/07
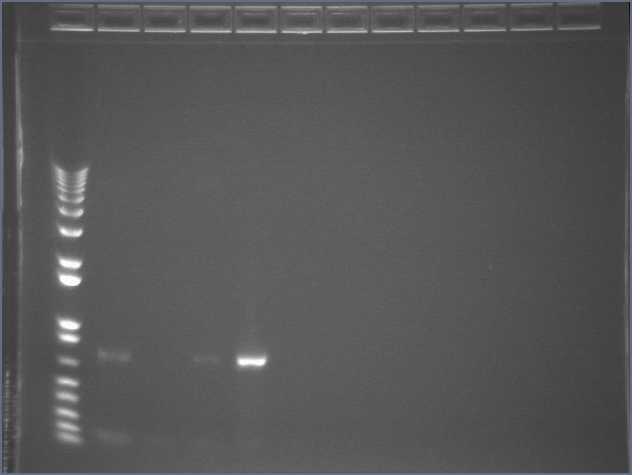
Ran 1.2% egels of all PCRs.
Lane 1: 1kb+
Lanes 2-5: JLO66PDZ1 PCRs #1-4
Lanes 6-9: JLO159PDZ1 PCRs #1-4
Lane 1: 1kb+
Lanes 2-5: JLO66PDZ2 PCRs #1-4
Lanes 6-9: JLO159PDZ2 PCRs #1-4
Lane 1: 1kb+
Lanes 2-5: JLO66PDZ1+2 PCRs #1-4
Lanes 6-9: JLO159PDZ1+2 PCRs #1-4
Parts sizes are as follows...
- J04500 - 220
- Lpp29 - 87
- OmpA66 - 63
- OmpA159 - 339
- PDZ1 - 261
- PDZ2 - 261
- PDZ1+2 - 546
Construct sizes are as follows (sum of parts plus 6bp in between parts)...
- JLO66PDZ1 - 649
- JLO66PDZ2 - 649
- JLO66PDZ1+2 - 934
- JLO159PDZ1 - 925
- JLO159PDZ2 - 925
- JLO159PDZ1+2 - 1210
VF2 primer anneals ~140bp upstream, and VR primer anneals ~100bp downstream of the cloning site.
3/23/07
After I retried digests, this time with ~3ug in 2x25ul digests, and clonewelled, I redid ligation-transformations.
3/30/07
Alain inoculated cultures from colonies, performed minipreps, diluted 1:100, and performed VF2-VR PCRs.
Lanes 1 to 6: PCR, JLO159-PDZ1 #1-6
Lane 7: 1kb+
Lanes 8 to 10: JLO66-PDZ1 #1-3
Lanes 1 to 3: JLO66-PDZ1 #4-6
Lane 4: 1kb+
Lanes 5 to 10: JLO159-PDZ1+2 #1-6
Approximate PCR sizes are as follows (sum of parts plus 6bp in between parts plus 240bp VF2/VR)...
- JLO66PDZ1 - 889
- JLO66PDZ2 - 889
- JLO66PDZ1+2 - 1174
- JLO159PDZ1 - 1165
- JLO159PDZ2 - 1165
- JLO159PDZ1+2 - 1450
4/10/07
Wow, okay, back in lab. Let's look at Alain's sequences.
- JLO66PDZ1 #6 is correct, beginning to end.
- JLO159PDZ1 #6 is correct from middle of PDZ1 domain to end. The PDZ1SR reverse sequencing reaction failed after <200bp.
- JLO159-PDZ1+2 #1 is correct from middle of JLO159 to near end of PDZ1+2.
- JLO159-PDZ1+2 #3 is correct from middle of JLO159 (a little farther upstream than #1) to near end of PDZ1+2. The orientations of the sequence alignments was weird.
Sent out the following sequencing reactions.
- 113: JLO159PDZ1 #6 OmpA46F
- 114: ...VR
- 115: JLO159PDZ2 #3 PDZ2F
- 116: ...PDZ2RS
- 117: JLO159PDZ2 #6 PDZ2F
- 118: ...PDZ2RS
- 119: JLO66PDZ2 #1 PDZ2F
- 120: ...PDZ2RS
- 121: JLO66PDZ2 #5 PDZ2F
- 122: ...PDZ2RS
- 123: JLO66PDZ1+2 #5 PDZ1F
- 124: ...PDZ2RS
4/16/07
Checked the sequencing results from 4/10/07, and they all check out. So along with 3/23 sequencing, the following have been confirmed.
- JLO66PDZ1 #6
- JLO66PDZ2 #1 or #5
- JLO66PDZ1+2 #5
- JLO159PDZ1 #6
- JLO159PDZ2 #6
- JLO159PDZ1+2 #1
4/18/07
Ordered GFP_F and GFPmek_R primers to PCR GFP out of Clontech's pEGFP vector.
Reconstituted primers in water to 100uM, then diluted to 10uM. Diluted 500ng/ul pEGFP vector 1:100 to 5ng/ul.
Setup two PCRs with 47ul Platinum PCR Supermix, 1ul GFP_F (10uM), 1ul GFPmek_R (10uM), 1ul pEGFP vector (5ng/ul). Incubated 95dC 10min, 30 cycles of (95dC 30s, 55dC 30s, 72dC 60s), 72dC 10min.
Took out 10ul from each PCR, mixed with 10ul water, and loaded onto 1.2% e-gel, ran for 30min.
PCR purified remaining 40ul into 30ul water.
No bands showed up on the e-gel.
4/19/07
Repeated PCR reaction with some new templates: (GFP in Topo), (AP-GFP in pGEX, GFP in pGEX, pEGFP, 1:100) using GFP_F and GFPmek_R primers, and (GFP in pSB1A2, 1:100) using bGFP_F and bGFPmek_R primers, which I had designed to amplify the GFP BioBrick. Also, changed PCR program to include 10 cycles with annealing temperature at 45dC and 20 cycles with annealing temperature at 55dC.
Ran 1.2% e-gel with 10ul of PCR product.
Lane 1: 1kb+
Lane 2: GFP in Topo PCR product
Lane 3: AP-GFP in pGEX PCR product
Lane 4: GFP in pGEX PCR product
Lane 5: pEGFP PCR product
LAne 6: GFP in pSB1A2 PCR product
PCR purified pEGFP PCR product, and Topo-cloned 4ul into 1ul pTrcHis vector, into 60ul Top10F' cells.
4/20/07
Got too many colonies. Difficult to pick individual clones. Picked 6, streaked, and colony PCR with GFP_F and GFP_mekR: 95dC 30s, 55dC 30s, 72dC 60s. Ran 1.2% e-gel.
Repeated PCR, picking from streaks #2 and 5, using GFP_F/TrcHis_R and GFPmek_R/Xpress_F. Also picked 7 more colonies (again, difficult to get single clones), streaked, and colony PCR with GFP_F and GFP_mekR.
4/21/07
Ran 1.2% e-gel.
Lane 1: 1kb+
Lane 2: Streak 2, XprF/GFPmek_R
Lane 3: Streak 2, TrcHisR/GFP_F
Lane 4: Streak 5, TrcHisR/GFP_F
Lane 5: Streak 5, XprF/GFPmek_R
Lane 6-12: Colonies 7-13, GFP_F/GFPmek_R
4/23/07
Ug, protein purification didn't work. It's probable that the pEGFP template is still in there after the PCR purification, and the GFP being expressed is from the pEGFP vector.
Redid PCR, 47ul PCR Supermix + 1ul GFP_F (10uM) + 1ul GFPmek_R (10uM) + 1ul pEGFP, 1:100 or 1:1000 of the 500ng/ul stock. Both PCRs yielded good bands in the Clonewell.
Initial Clonewell collections from four wells (split up 50ul PCR into two wells from each PCR) yielded ~30ng/ul. Second Clonewell collections yielded ~7ng/ul. Vacufuged initial collections down to ~30ul, for ~90ng/ul.
Used 4ul in a 5ul pTrcHis Topo reaction, split up into 1 and 4ul for 30ul Top10 transformation. Used 0.5ul in a 5ul pTrcHis Topo reaction, split up into 1 and 4ul for 30ul Top10F' transformation.
4/24/07
Transformations yielded very small colonies after 14h, so I left them out at room temperature.
Made two more PCRs with 1:1000 and 1:5000 pEGFP template. On an e-gel, the 1:5000 PCR yielded a lighter band. PCR-purified both samples, and used 4ul of purified PCR with 1:100, 1:1000, 1:5000 pEGFP template in a transformation into 20ul Top10 cells.