User:Perry/Fall 2006 Harvard iGEM work
10/4/06
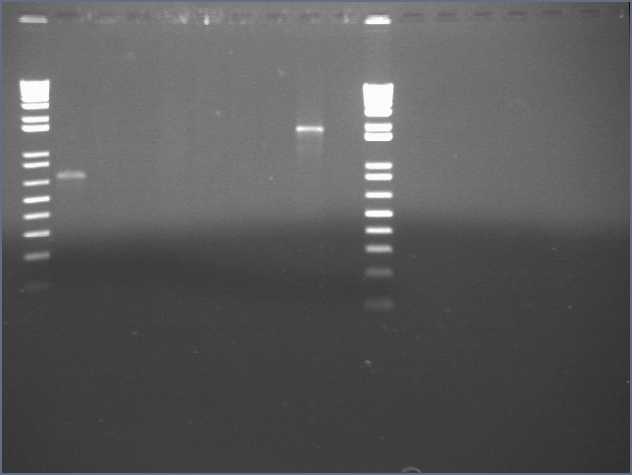
VF2/VR PCRs and carb streaks from the 8/30/06 streaks.
Lanes 1,11: 1kb+
Lanes 2-5: JLO66, W, H, S
Lanes 6-9: JLO159, W, H, S
Lane 10: E0241
10/5/06
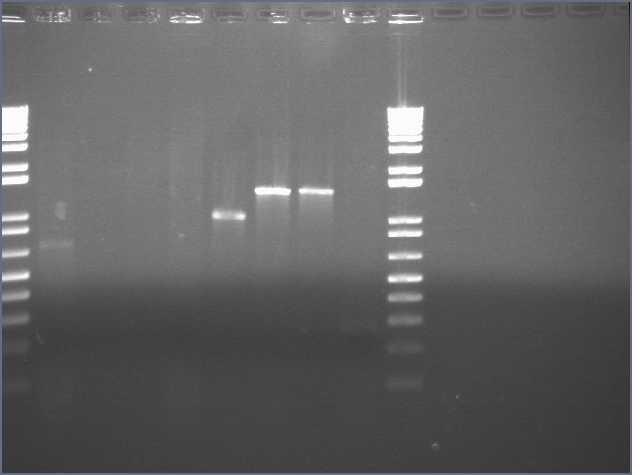
VF2/VR PCRs and carb streaks, picked from streaks made on the designated date.
Lanes 1,10: 1kb+
Lanes 2-5: JLO66(10/4), W, H (8/25), S(8/31)
Lanes 6-9: JLO159, W, H (8/31), S (10/4)
Ug, I give up trying to salvage.
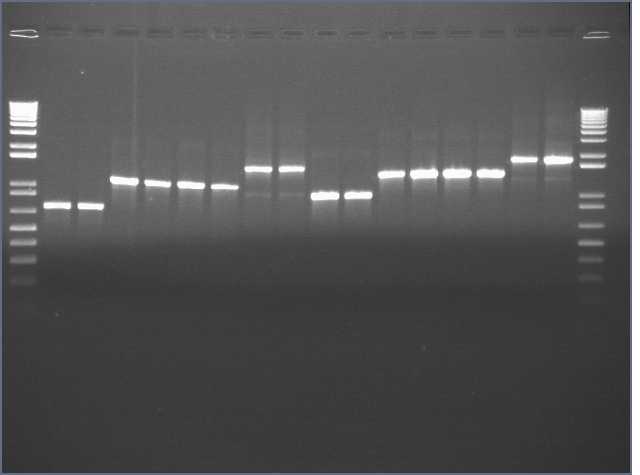
I went back to the transformation plates from 8/23/06 and made colony PCRs/carb streaks from those.
Lane 1,18: 1kb+
Lanes 2,3: JLO66 1,2
Lanes 4,5: JLO66W 1,2
Lanes 6,7: JLO66H 1,2
Lanes 8,9: JLO66S 1,2
Lanes 10,11: JLO159 1,2
Lanes 12,13: JLO159W 1,2
Lanes 14,15: JLO159H 1,2
Lanes 16,17: JLO159S 1,2
10/10/06
Prepared 4ml LB+amp liquid cultures.
10/11/06
Refrigerated 800ul from each liquid culture, and miniprepped the rest. Sent out for sequencing, PT094-PT114, in same series as PT073-PT093.
10/18/06
Inoculated 1.5ml LB amp cultures: JLO66, W, H, S, JLO159, W, H, S at 4pm
10/19/06
Took 40ul of cultures at 9am and reseeded into 2ml LB amp. At 1pm, took 1ml and continued to incubate to JLO66W, JLO66S, JLO159W, JLO159S, with 1mM IPTG; JLO159S(-IPTG); and JLO159S(+IPTG,-DNA). I also reseeded a new 2mL culture of JLO159S with 4ul from this morning's cultures with 1mM IPTG added to check proliferation of cells with IPTG present. At 4pm, the JLO159S with IPTG added at seeding grew. I spun down the (JLO66W, JLO66S, JLO159W, JLO159S)+IPTG, JLO159S(- IPTG), JLO159S(+IPTG,-DNA). There was only about 0.5ml of JLO159S(+IPTG,-DNA). I resuspended in 1mL PBS, and spun down again, and resuspended in 40ul DPBS. I reconstituted BF probein 865ul water for 200uM, then diluted to 50uM. I then added 10ul of 50uM BF to the 40ul resuspensions. I accidentally added DNA probe to JLO159S(+IPTG,-DNA), so I took the JLO159S(+IPTG at reseeding), spun down 1mL, resuspended in 1mL PBS, spun down, resuspended in 50ul PBS.
After 30min rotating, I spun down, washed 3x in 100ul PBS, resuspended in 50ul PBS, pipetted 5ul onto glass slide and covered with poly-L-lysine treated coverslip.
10/20/06
Arnaud helped me out with the microscope today. Things I learned:
- Turn on mercury lamp on first, and let it warm up for a few minutes.
- Dilute cells in PBS. We tried 1 in 2, but it will probably need a greater dilution.
- After you drop coverslip on the slide, take a kimwipe, cover the entire slip, and press down with the heel of your hand. This helps in creating a single layer of cells.
- Put on drop of immersion oil on top of the slide. Lower 100X objective until it makes contact with the oil. Then continue to lower the objective slowly until you see the cells come into focus. Try to get this first time, because lowering and raising the objective causes the coverslip to lower and rise, allowing bubbles to form.
- After you're finished, remember to clean the oil from the objective with lens paper.
10/21/06
Transformed JLO66,W,H,S, JLO159,W,H,S into Top10 cells, using minipreps 10/11/06.
Inoculated cultures for miniprep: R0010-B0032, Lpp29, OmpA66, OmpA159, LO66, LO159, StrepW, StrepH, StrepD, StrepSCD-NM, Strep E2, Strep13.
Inoculated 1ml JLO66,W,H,S, JLO159,W,H,S.
10/22/06
Diluted 20ul of JLO66,W,H,S, JLO159W,H,S cultures into 1ml LB, grew 2 hours, added 5mM IPTG, grew 3 hours, pelleted. I also reseeded a JLO66H culture without IPTG induction. Took rest of JLO66,W,H,S, JLO159,W,H,S cultures and pelleted, resuspended in LB+20% glycerol, froze in -80dC freezer.
Pelleted R0010-B0032, Lpp29, OmpA66, OmpA159, LO66, LO159, StrepW, StrepH, StrepD, StrepSCD-NM, Strep E2, Strep13 cultures for miniprep later.
10/23/06
Reattempted transformation into Top10 cells using 5ul of 1:100 diluted minipreps 10/11/06.
Resuspended JLO66,W,H,S, JLO159W,H,S, and JLO66h(-IPTG) pellets in 100ul water. Took 5ul of resuspension, added 11ul water and 4ul 5X sample buffer. For JLO66W, I noticed a smaller pellet than the rest, so I used 10ul of resuspension of 6ul water.
SDS-PAGE, loaded 15ul into each well, 10ul protein ladder, and 5ul of a His6-tagged purified protein Alain provided.
10/24/06
Antibody staining and visualization of membrane. Cut it in the middle of lane 7 and incubated each half with a different antibody.
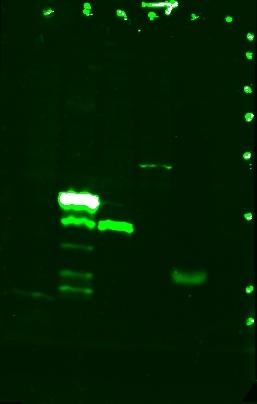 Stained with mouse monoclonal anti-His6.
Stained with mouse monoclonal anti-His6.
Lane 1: JLO66H, monomeric streptavidin with His6 tag on OmpA(46-66)
Lane 2: JLO66S, dimeric streptavidin with His6 tag on OmpA(46-66)
Lane 3: JLO159H, monomeric streptavidin with His6 tag on OmpA(46-159)
Lane 4: JLO159S, dimeric streptavidin with His6 tag on OmpA(46-159)
Lane 5: His6 (+)
Lane 6: JLO66, no streptavidin on OmpA(46-66)
The bands for JLO66H, JLO159H, and JLO159S and the top band for JLO66S are expected sizes. I think the "ladder" pattern with JLO66S may be due to breakdown of the protein.
(Lane 7: protein ladder, denoted by dots)
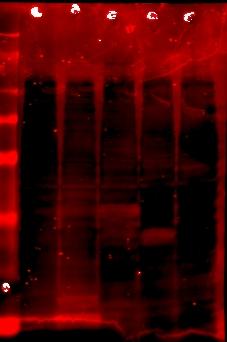 Stained with rabbit polyclonal anti-streptavidin
Stained with rabbit polyclonal anti-streptavidin
Lane 7: protein ladder
Lane 8: JLO66
Lane 9: JLO66W, monomeric streptavidin on OmpA(46-66)
Lane 10: JLO66S
Lane 11: JLO159W, monomeric streptavidin on OmpA(46-159)
Lane 12: JLO159S
The gel was contaminated with residual coomassie blue in the tray, so visualizing the red anti-rabbit secondary antibody is difficult due to the high background from coomassie blue. But you can almost see bands of the same sizes in lanes 9-12 as from lanes 1-4 in the His6 stain. Woot.
Plans to redo the SDS-PAGE with adjusted volumes of cells, and avoid coomassie blue contamination.
10/25/06
Repeated SDS-PAGE. I added JLO66H(-IPTG) as another negative control, and I adjusted volumes of resuspended pellet used. From JLO66W, JLO66H, and JLO159W, I used 20ul + 5ul sample buffer. For JLO66H, I used 2.5ul + 13.5ul water + 4ul buffer. For JLO159H and JLO66H(-) and His6(+), I used 5ul + 11ul water + 4ul buffer. For JLO66, I used 10ul + 6ul water + 4ul buffer.
SDS-PAGE. I loaded 20ul into each well, and 10ul protein ladder.
Lane 1: ladder
Lanes 2-4: JLO66W, H, S
Lanes 5-7: JLO159W, H, S
Lane 8: JLO66
Lane 9: JLO66H (-IPTG)
Lane 10: His6+ protein