User:Patrick Hampson/Notebook/Biology 210 at AU
January 20, 2015 Created my notebook -P.H.
January 20, 2015 Biological Life Found in Sections of American University Performed: January 14, 2017.
Purpose:
Life on Earth is magnificently diverse. Today's life forms are specialized and equipped for where they live, as they inherit the most fit characteristics from organism many generations before their own. Due to their adaptations and their ancestor's migrations, not many life forms found on Earth today exist in numerous environments throughout the world. Surely, there are many classifications of animals that live in various environments throughout the world, such as bats, mice, and foxes, to name a few, but there are even variations in characteristics between similar species due to inherited adaptations. In this lab, small, sample sized ecosystems, also referred to as transects, will be studied (Bentley, Walters-Conte, Zeller, 2015). Transects contain both biotic (living) and abiotic (non-living) factors that interact with one other and create a web of complex relationships that are essential to balance life within the ecosystem.
Materials and Methods:
Each group was assigned a different transect to thoroughly survey and investigate. The assigned transects were 20 x 20 meters, except for one transect, the manicured grassland (Quad), which was a triangle. Groups went out and examined their transects thoroughly. Once every square meter of the transect was examined, the groups created their own topographical, or aerial maps which depicted their transect in full detail, down to the placement of seemingly small abiotic factors such as patches of snow. A representative dirt and vegetation sample was then retrieved from a random location in the transect and placed in a 50 mL tube, until the tube was filled. Anywhere between 10 and 12 grams of the dirt and vegetation sample were placed into a plastic jar which also contained 500 mLs of purified water. 0.1 grams of dried milk was added to the mixture, and also mixed in very gently, to not kill the living organisms from the sample. The lid of the jar was left off, to minimize the impact on the pint-sized ecosystem. The jars were left alone for one week, until the next lab time.
Group 5 transect:
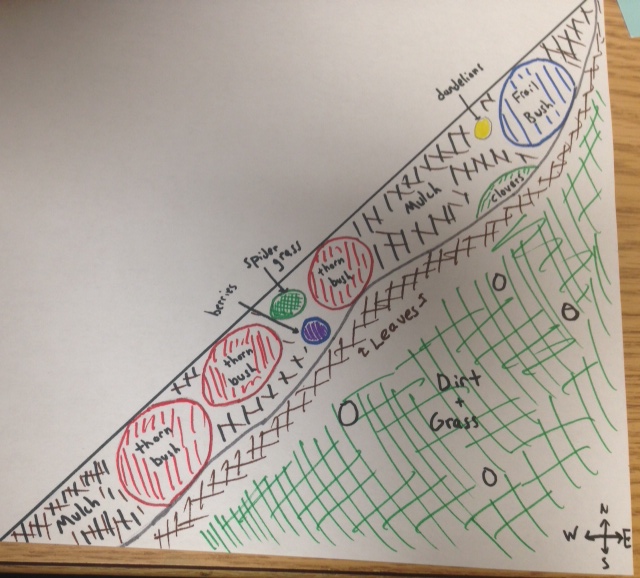
The transect assigned to the group was a triangular (right) portion of the quad which was located in front of Hurst hall, right next to a walkway and a sign for "Friedheim Quadrangle". The transect was comprised mainly of a level, grassy plain, with a portion of it including an area of bark mulch with potted and wild plants. The transect contained seasonal abiotic factors, such as decaying dead leaves, proving fall was some time ago, as well as snow, illustrating the presence of winter. Abiotic factors included: snow, dirt, mulch, and dead leaves. Biotic factors included: grass, spider grass, clovers, wild berries, dandelions, bushes equipped with thorns (rose bush?), and a population of another type of bush that looked rather frail. The mulch area was a constant on the western side of the transect. Leaves loosely blanketed the mulch area, and even spilled onto the closest edges of the grass. Thorn bushes ranged from the bottom south west portion of the triangle, to just above the western north-south midpoint of the triangle. Berries and spider grass were located right next to each other, in between two thorn bushes, near the north-south midpoint of the transect. The clovers and dandelions were located more towards the northern most bush, which did not have any thorns and looked rather frail. Dirt was a constant base throughout the transect, being located underneath the mulch, as well as a base for the grass. Grass accounted for most of the area of the transect, ranging from most of the east to west distance, as well as most of the north to south distance. snow was scattered though out the transect.
Reference Sources:
Bentley, M., Walters-Conte, K., Zeller, N. 2015. A Laboratory Manual to Accompany General Biology II. American University Department of Biology: Washington, DC. 14-17.
January 27, 2015
Examining Algae and Protists
Performed January 21, 2015
Purpose:
The classification of organisms has always been an ongoing project, as new information is brought to the forefront with discoveries, and can make us refute what we accept today. Only recently has the scientific world been able to open up a whole new world of discoveries to examine with the invention of the microscope, which allows them to observe microscopic organisms (Bentley, Walters-Conte, Zeller, 2015). These microscopic organisms can be categorized into two classifications, prokaryotes and eukaryotes. Eukaryotes will be examined in this lab, specifically, the two large groups which are classified as algae and protists (Bentley, Walters-Conte, Zeller, 2015). It should be predicted that these two groups will be present when samples are drawn from the Hay Infusion Culture, insinuating that they live in the assigned transect.
Materials and Methods:
The groups retrieved their jars which contained their Hay Infusion Culture. The culture was carefully examined and observations regarding appearance and smell were recorded. Two samples were drawn from different niches, also known as positions or locations, inside of the jar. One sample was drawn directly below the seemingly moldy surface, near leaf debris, and the other sample was drawn directly from the dirt covered bottom. Wet mounts were made from both of these samples, and they were observed under the microscope. Many different organisms were observed under the microscope, but only three organisms were recorded from each sample. These organisms were identified with the use of a dichotomous key. After the examination of these organisms was complete, the group prepared for the next lab, in which prokaryotic organisms would be observed. Four test tubes, each containing 10 mLs of sterile broth, were retrieved, and labeled 10^-2, 10^-4, 10^-6, and 10^-8. Eight petri dishes were retrieved, four of which contained only a nutrient agar base, whereas the other four contained the nutrient agar base plus tetracycline (an antibiotic). The dishes containing the antibiotic were marked, and plates from both groups were labeled 10^-3, 10^-5, 10^-7, and 10^-9, and each plate was initialed. The Hay Infusion Culture was gently mixed, stirring up all the organisms within. A micropippetor set at 100 microliters was used to draw up a sample from the culture, and transfer it to the test tube labeled 10^-2. The test tube was mixed via swirling. A new tip was put on the micropippetor, and 100 microliters was taken from the 10^-2 tube and placed in the tube labeled 10^-4. This tube was also swirled in order to mix up the contents thoroughly. The previous steps were followed while transferring 100 microliters from the tube labeled 10^-4 to the tube labeled 10^-6, and so on from the tube labeled 10^-6 to the tube labeled 10^-8. 100 microliters were transferred from the tube labeled 10^-2 to the tube labeled 10^-3, as the additional substance provided by the agar plate acts as a further dilution. The 100 microliters was thoroughly spread throughout the dish using a sterilized spreader. These steps were repeated for adding 100 microliters from test tube 10^-4 to plates labeled 10^-5, test tube 10^-6 to plates labeled 10^-7, and test tube 10^-8 to plates labeled 10^-9, each time using a new micropippetor tip and a sterilized spreader. The dishes were left out at room temperature on a rack for a week. The point of growing the prokaryotic organisms in a petri dish is to make viewing the organisms easier, as viewing them directly from the culture would be very challenging (Bentley, Walters-Conte, Zeller, 2015).
Data and Observations:
The Hay Infusion Culture: The culture had a peculiar smell to it, nothing like I have smelt before, the only way to describe it was rancid. There was mold at the top of the culture, which resembled burned hot chocolate, as it appeared rugged.
The samples:
Top (Near the plant matter, directly under the mold covered surface): We observed what appeared to be:
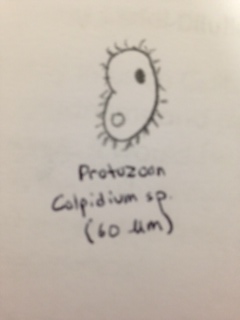
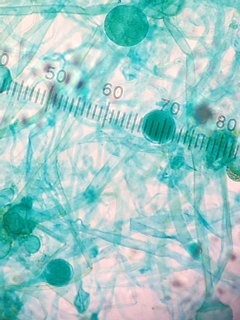
Colpidium sp. (60 micrometers), (Protozoa) appeared to be relatively translucent with a greenish hue, motile in the sense that it appeared to glide.
Paramecium aurelia (120 micrometers), (Protozoa) appeared to be relatively translucent with a greenish hue, motile in the sense that it appeared to glide. Seemed to simply be larger and proportionally thinner than Colpidium sp..
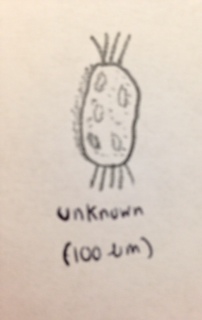
and an unknown (not identifiable from the dichotomous key) organism, (Protozoa) which had cilia on both ends, and some on one side (100 micrometers). This organism was also relatively translucent, but had a slightly dirty hue to it.
These organisms could be near the top of the culture for multiple and various reasons, such as they were eating the decomposing leaves, are comfortable in the structure that the mold brings to their environment, or even that they wish to get more sunlight than the organisms living at the bottom of the culture. Neither of the identified organisms undergo photosynthesis, but the third very well could have. Paramecium do tend to feed of of photosynthesizing algae, so it would not be preposterous for them to be feeding off the leaves.
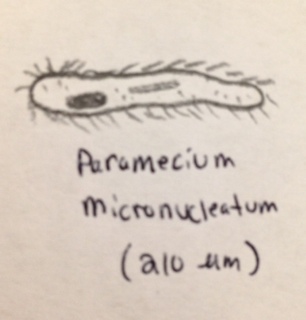
Bottom (Right from the dirt-settled bottom): we observed what appeared to be Paramecium multimicronucleatum (210 micrometers), (protozoa) this organism was translucent and had dirty brown hue to it. It motile motion makes it appear to glide.
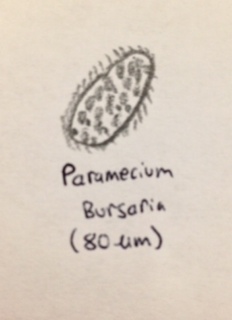
Paramecium bursaria (80 micrometers), (protozoa) this organism was almost seemingly clear (very translucent), with hints of green throughout. Its motile motion also made it appear to glide.
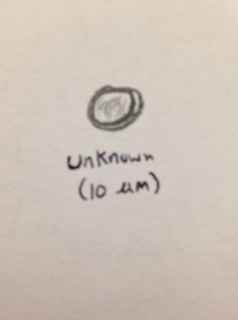
and an unknown (not identifiable from the dichotomous key) organism that was circular and vibrating (10 micrometers), this organism was also translucent and had hints of green throughout. This organisms mode of motility simply seemed to be vibrating. No visible cilia of flagella.
These organisms could have been living in the bottom of the sample for various reasons, such as they prefer not to experience too much sunlight, or just do not need to undergo photosynthesis in any sense, or they could have liked the structure and comfort that the dirt-settled bottom brought to them.
If the Hay Infusion Culture continued on for another two months, I would expect one of two things. Either all of the organisms in the culture would die, and this is because there is no plant life to create oxygen for the organisms to breathe. Although the lid of the culture is wide open, the mold will have completely taken over the surface, preventing any way for the protozoa to get any oxygen from the surface. OR there will be an abundance of one particular organism that has adapted to its environment and found a way to outlast/ out-compete all the other organisms in the culture. This organism would be the most fit for the environment, and therefore do the best in competing for survival. Factors that could change the community in the sample include, but are not limited to: the mold completely taking over the surface, creating an impenetrable barrier for the protozoa, plant life coming into existence, the elimination of oxygen in the water, lack of resources/essentials of life, and of course time.
Conclusion and Future Directions:
Protozoa were the only eukaryotic organisms that were observed from the Hay Infusion Culture samples, there was not any algae observed in either sample. This could simply be human error, where the eye is drawn to the moving organisms like the protozoa, or sampling error, where the group drew samples from niches where algae would not be expected to be abundant. That being said, just because there was no algae observed from the samples, does not mean that there was none present in the Hay Infusion Culture.
Fulfilling the five requirements of life: Paramecium aurelia: This organism fulfills the five needs of life while living at the top of the culture. Requirement 1: Obtaining Energy. Paramecium are known to have a beneficial relationship with algae, where the Paramecium can gain energy from the photosynthesis of algae (Naturforsche). This relationship is constant, and should be expected to exist within the culture. Requirement 2: Comprised of Cells. Paramecium are uni-cellular organisms, which means that their entire body is comprised of a cell. Whether it be one cell or many, all living organisms are comprised of cells, and Paramecium aurelia exemplify this fact.The survival of the Paramecium in the culture is, in itself, an example of the third requirement. The Paramecium was able to adapt to a new environment that it was placed in, even though it is not an aboriginal species to that environment. This shows the organism's ability to adapt based on information that it received and processed, in order to give itself the best chance of survival. Requirement 4: Replication. Although no replication was witnessed under the microscope, it is known that Paramecium replicate asexually and sexually, depending on environmental circumstances (Russell). Requirement 5: Evolution. No evolution was witnessed under the microscope, and this is logical. Evolution takes a lot of time to make even the slightest of changes. It makes sense that no changes would occur after only observing the organisms for a few minutes. (Freeman, Quillin, Allison, 2014).
Reference Sources:
Bentley, M., Walters-Conte, K., Zeller, N. 2015. A Laboratory Manual to Accompany General Biology II. American University Department of Biology: Washington, DC. 19-23.
Freeman, S., Quillin, K., Allison, L. 2014. Biological science (5th ed.). Pearson: Glenview, IL. 2
Naturforsch, Z. 2004. Green paramecia as an evolutionary winner of oxidative symbiosis: a hypothesis and supportive data. Journal of Biosciences. The National Institute of Health: Bethesda, MD.
Russell, E. The Biology Classics: Paramecium - Reproduction. BioMedia Associates.
February 3, 2015
Investigating the characteristics of bacteria
Performed: January 28, 2015
Purpose:
Unicellular organisms are an essential part of the balance of nature, and can be found in all three domains of Bacteria, Archaea, and Eukarya (Freeman, Quillin, Allison, 2014). In the previous lab, unicellular organisms from the Eukarya domain were examined and studied, but for this week, Prokaryotes will be observed, particularly those from the Bacteria domain. Prokaryotes are organisms which lack a nucleus, and all other forms of membrane-bound organelles (Freeman, Quillin, Allison, 2014). The reason why prokaryotes from the Bacteria domain will be studied in particular this week is simply based off of their abundance. Prokaryotes from the Bacteria domain are everywhere! Which is why it is no surprise that they exist in the transect samples. Archaea, on the other hand, tend to live and thrive in extreme environments, making them “extremophiles” (Bentley, Walters-Conte, Zeller, 2015). The class transects were not located in any extreme environments such as the bottom of the sea, or in hot springs, so it is not likely that any Archaea will appear in observations in the course of this lab. The bacteria that have grown on agar plates for the past week are not likely to all be similar, they will be diverse in shape, size, motility, and gram type. Their colonies will vary in numerous characteristics too such as in color, shape, size, texture, elevation, and edge characteristics. By observing all of their characteristics thoroughly, these bacteria can be categorized and their specific identity can be determined.
Materials and Methods:
Procedure for part 1: The Hay Infusion Culture was examined, and changes were recorded for one last time. The agar plates prepared from the previous lab were then observed, and the number of colonies located on each plate was determined and recorded. From these numbers, a conversion factor corresponding with the dilution rate (ex. 10^-3 dilution rate corresponds with 10^3 conversion rate) was used to establish the number of colonies that could be found per mL in the Hay Infusion Culture.
Procedure for part 2: Comparisons were made between the number of colonies on the agar plates and those on the agar and tetracycline plates.
Procedure for part 3: Four selected colonies, two from the agar plates and two from the agar and tetracycline plates, were selected to be observed further under the microscope. The colonies characteristics were recorded. Four separate wet mounts, each containing a sample from one colony, were made by removing a portion of the colony using a sterilized loop device. The slides were carefully labeled to ensure that there would be no mixing up of the slides, and each colony’s plate of origin would not be a mystery. The wet mount slides were observed with 10x magnification, 40x magnification, and then with 100x magnification with oil immersion. Then a gram stain was made of the selected colonies. A portion of the previously selected colonies were scraped off of the plates with a sterile loop device and placed on different slides, each being mixed with a drop of water. Each slide was clearly labeled with the colonies plate of origin. The slides were passed over a bunsen burner, bacteria side up, until the slide was dry. The slides were placed over a staining tray and covered with crystal violet for one minute. The stain was then rinsed off with distilled water. The slides were then covered with Gram’s iodine mordant for one minute. The stain was then rinsed off with distilled water. The slides were then rinsed with 95% alcohol for 10-20 seconds. The slides were then covered with safranin stain for 20-30 seconds. The slides were then rinsed off with distilled water. The slides were then dried using kimwipes and a dabbing technique. The slides were then observed with 40x magnification and 100x magnification with oil immersion technique.
Procedure 4: PCR procedure was followed to prepare for the next lab where DNA sequencing is explored. A single colony was transferred to 100 microliters of water in a sterilized tube. This was done four times, as four colonies were observed. The tubes were then heated at 100 degrees Celsius for ten minutes. The samples were then centrifuged for five minutes at 13,400 rmp. While waiting for the centrifuge to finish, 20 microliters of a primer/water mix was added to a labeled PCR tube. The tubes were then mixed to thoroughly dissolve the PCR bead located within the tube. Five microliters of supernatant were obtained from the centrifuge samples and transferred to the 16S PCR reaction. The tube was then placed in the PCR machine.
Data and Observations: There were numerous changes that occurred to the Hay Infusion Culture during the course of the week since previous observations were made. The rugged film substance that had previously covered the surface of the water had dissipated, and the dirt bottom could be seen from an aerial view. This could be due to whatever comprised the film to die due to lack of nutrients, or because it was eaten away at by other organisms located in the sample. A significant amount of the water had disappeared, it is logical to suspect that evaporation was the cause of this. The previous water level could be estimated based on the ring of debris stuck to the side of the jar. The smell of the jar was just as rancid as ever.
Table 1
| Dilution | Agar Type | Colonies Counted | Conversion Factor | Colonies/mL |
| 10^-3 | Nutrient | 1500 | 10^3 | 1500000 |
| 10^-5 | Nutrient | 300 | 10^5 | 30000000 |
| 10^-7 | Nutrient | 150 | 10^7 | 1500000000 |
| 10^-9 | Nutrient | 60 | 10^9 | 60000000000 |
| 10^-3 | Nutrient + Tet. | 700 | 10^3 | 700000 |
| 10^-5 | Nutrient + Tet. | 150 | 10^5 | 15000000 |
| 10^-7 | Nutrient + Tet. | 250 | 10^7 | 2500000000 |
| 10^-9 | Nutrient + Tet. | 5 | 10^9 | 5000000000 |
In comparison, the number of colonies found on the agar plates was larger than the number of colonies found on the plates that also contained tetracycline. This was predictable, because tetracycline inhibits bacterial growth. Only a few colonies were able to demonstrate antibiotic resistance and grow on the tetracycline plate. Of the colonies that grew on the tetracycline plates, all of them were colored orange.
The colony observed from the nutrient plate derived from 10-9 dilution (Figure 12).
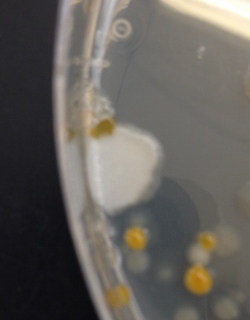 Figure 13
Figure 13
The colony observed from the nutrient plate derived from 10-7 dilution (Figure 13).
The colony observed from the nutrient plate with tetracycline from 10-7 dilution (Figure 14).
The colony observed from the nutrient plate with tetracycline from 10-5 dilution (Figure 15).
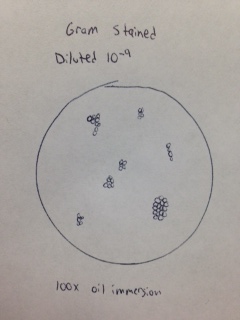
Gram stain of colony from 10-9 dilution (Figure 16). The bacteria observed appear to be Gram negative cocci.
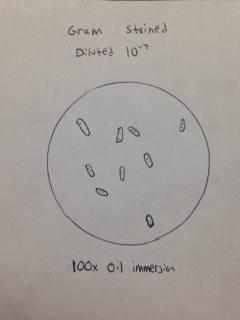
Gram stain of colony from 10-7 dilution (Figure 17). The bacteria observed appear to be Gram positive bacillus.
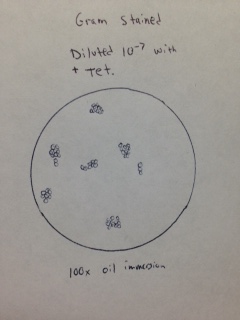
Gram stain of colony from 10-7 dilution with tetracycline (Figure 18). The bacteria observed appear to be Gram positive cocci.
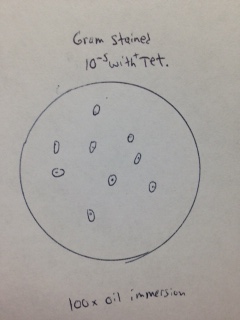
Gram stain of colony from 10-5 dilution with tetracycline (Figure 19). The bacteria observed appear to be Gram positive cocci-bacillus.
Table 2
| Colony Label | Plate Type | Colony Description | Cell Description | Gram +/- |
| 10^-9 | Nutrient | Orange, Medium-Large, circular, smooth, glistening, convex, entire edge | Cocci, Non-motile | Gram - |
| 10^-7 | Nutrient | White, Large, undulate, raised, glistening, irregular edge | Bacillus rods, motile, but not flagella seen, no chains or clusters | Gram + |
| 10^-5 | Nutrient + Tet. | Cloudy white, medium, dry surface, convex, circular, entire edge | Cocci-bacillus, non-motile, some dividing, no chains or clusters | Gram + |
| 10^-7 | Nutrient + Tet. | Orange, Large, smooth, glistening, round, convex, entire edge | Staphylococci, non-motile, clustered | Gram + |
Conclusions and Future Directions:
Overall, it is amazing to see these prokaryotes replicate. Given the right nutrition, the tiny, microscopic organisms are able to multiply to the point where their colonies are easily seem with the naked eye. Though the sheer number of colonies grown on the agar plates outweighed the number grown on the agar and tetracycline plates, it is still impressive that the bacteria are able to replicate when facing resistance, such as antibiotics. The number of colonies which were observed is microscopic in comparison to those that can be found in standard dilution, as the colonies observed were diluted at a minimum of one thousand times the standard concentration. The fact that some plates observed contained colony numbers that were too high to count accurately illustrates the seemingly infinite number of bacteria that exist in our world.
Tetracycline inhibited the growth of colonies on plates, as can be seen by the smaller number of colonies found on the tetracycline plates. Tetracycline is a relatively inexpensive antibiotic that is effective against both gram positive and gram negative microorganisms. It works by preventing protein synthesis, and it does this by averting the aminoacyl-tRNA’s attachment with ribosomal acceptors (Chopra, Roberts, 2001).The lack of white or cream colored colonies on the tetracycline plated could mean that that bacteria is heavily influenced by the presence of tetracycline, and cannot form an antibiotic resistance. Orange colonies were able to form on the tetracycline plates, but not as frequently as they were able to on the plates that contained just agar. This means that the tetracycline did play a role in limiting the number of orange colonies, but some of the bacteria were able to form a resistance, thus forming some colonies.
In the next lab, the DNA sequencing of the bacteria will be explored with the help of the PCR reaction.
References:
Bentley, M., Walters-Conte, K., Zeller, N. 2015. A Laboratory Manual to Accompany General Biology II. American University Department of Biology: Washington, DC. 26-32.
Chopra, I., Roberts, M. 2001. Tetracycline Antibiotics: Mode of Action, Applications, Molecular Biology, and Epidemiology of Bacterial Resistance. Microbiology and Molecular Biology Reviews. 65(2): 232-260.
Freeman, S., Quillin, K., Allison, L. 2014. Biological science (5th ed.). Pearson: Glenview, IL. 531.
February 10, 2015
Observing the Characteristics of Plants and Fungi
Performed: February 4, 2015
Purpose:
Plants and Fungi are both key species with important roles in sustaining of Earth's atmosphere. Plants fix CO2, converting it into oxygen through the process of photosynthesis. Through this process, plants continually emit an element that is essential to the survival of many forms of life. Fungi, while they do not create oxygen, act as decomposers, releasing CO2 into the atmosphere and injecting forms of Nitrogen into the ground and water supplies, which will be later utilized by plants (Bentley, Walters-Conte, Zeller, 2015). Plants have evolved over time to become a very diverse group or organisms, with different methods of sustaining the species as a whole (reproductively speaking), and functions for sustaining life. Over time, it is accepted that plants have adapted from their aquatic origins to become terrestrial organisms, which have since developed vascular systems, seeds, and most recently, pollen, as well as numerous other features and functions that has allowed them to survive and develop (Bentley, Walters-Conte, Zeller, 2015). Fungi have also undergone a series of adaptations and development as well, creating their own distinct functions for reproduction and survival. The purpose of this lab is to examine the distinct characteristics of plants and fungi, and form an understanding for how these characteristics play a key role in the species' survival.
Materials and Methods:
Procedure 1: Groups ventured to their respective transects and gathered vegetation, which were placed into ziploc bags. In one bag, a leaf litter sample was added, which was comprised of dead leaves and soil clusters found around the transect. Live vegetation samples were collected and placed in another bag, as well as some additional samples from the transect such as acorns, seeds, flowers, or pine cones for example.
Procedure 2: Individuals observed the differences between a Bryophyte, moss, and a portion of an Angiosperm, the lily plant stem. The height differences were observed. Then the lily stem was further examined so groups could carefully observe the xylem and the phloem layers of the stem.
Procedure 3: Leaves of moss were examined further under a dissection scope. Observations were made and recorded. Then the leaves of the Angiosperm were further examined so that the stomata and guard cells of the leaves could be observed.
Procedure 4: Moss was examined under a microscope again, with focus on identifying the female and male gametophytes and sporophytes. Diploid and haploid portions of the moss life cycle were noted. Then a lily flower was dissected, in order to have students attempt to identify portions of the plant that are necessary for reproduction, such as the anther, style, stigma, and ovary.
Procedure 5: Molds were examined in petri dishes. The top was observed to see the majority of the mold, which contained the mycelium, sporangia, and spores. The petri dishes were then flipped over to observe rhizoids that had grown through the agar foundation of the dishes. A mushroom was placed under a dissection scope and observed. Prepared slides were then examined under microscopes. The two prepared slides observed were of Rhizopus and Rhizopus nigricans.
Procedure 6: In preparation for next week, where invertebrates will be observed, Berlese Funnels were constructed. 25 mL of a solution containing a ratio of 50:50 water/ethanol was poured into a tube. A piece of screen was placed at the bottom of a funnel, and was taped into place. The tube containing the water/ethanol solution was attached to the bottom of the funnel with layers of wax-film to prevent evaporation. The device and parafilm were secured with tape. The funnel device was attached to a ring stand with a clamp. The leaves and trace amount of soil gathered from the transect in procedure 1 were placed into the top of the funnel device. The device was placed under a 40 watt lamp. The device was then covered with foil. The Berlese funnels were left in the labs for a one week period of time.
Data and Observations:
Plant Samples Gathered and Observed:
Table 3
| Transect Sample Plants | Location and count | Description | Vascularization | Specialized Structures | ' | Mechanisms of Reproduction | Genus |
| 1. Grass | Field portion, countless | Short, green blade, rooted, 8.5 cm | Yes | Stomata, guard cells, cuticle | Seeds | Poaceae | |
| 2. Clover | Mulch bed, 200 stems | Green, rooted, white roots, 3.25 cm | Yes | Stomata, guard cells, cuticle | Pollen | Trifolium | |
| 3. Spider Grass | Mulch bed, 15 plants | Green blade, rooted, white root, 28 cm | Yes | Stomata, guard cells, cuticle | Seeds | Poaceae | |
| 4. Rose Buds | Mulch Bed, 7 bushes | Brown, dead, half-eaten | Yes | Stamen, anther, stigma, style, ovary, ovule, filament, microsporangium | Pollen | Rosa | |
| 5. Leaf | Mulch bed, 3 plants | Green, broad, spiny shaped, soft | Yes | Stomata, guard cells, cuticle | Pollen | Unknown | |
Arrangement of Leaves for transect plants:
1. Grass: Leaves comprise the majority of the length of the plant. Leaf appeared to be a common grass leaf. Leaves are the only visible (above ground) portion of the plant. Number of leaves depends on the size of the clustered root system.
2. Clover: Multiple (2-3) circular shaped leaves are attached to the top of a proportionally long stem.
3. Spider grass: Leaves comprise the majority of the length of the plant. Leaf appeared to be much larger and elongated that common grass. Leaves are the only visible (above ground) portion of the plant. There appeared to be an extensive (deep) root system for each blade
4. Rose bud: On adult bushes, leaves are scattered throughout the branch structures and stems. This bud however did not contain any leaves.
5. Leaf: This leaf was protruding from the ground, attached to another similar leaf. The two leaves were the only visible (above ground) portions of the plant.
Monocot or Dicot seeds:
The only undeveloped plant that was brought back from the transect was a dead rose bud. It is not possible to distinguish between monocot and dicot by looking at this pathetic looking bud, but roses in general are dicot. It appeared as though flower petals could be seen within the bud.
The fungi observed under the dissecting scope and microscope in procedure 5 were mushroom (Basidiomycota), and Rhizopus nigricans (Zygomycota).
Rhizopus nigricans:
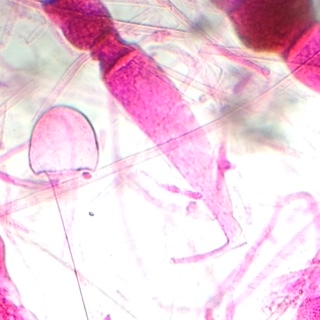
This sample is clearly fungus due to the presence of sporangia, the black clusters. The sample shows what is unmistakably a complex formation of hyphae, also known as the mycelium (the complex network of filaments).
Rhizopus:
This sample is clearly fungus, more specifically of the Zygomycota group, due to the presence of formed zygosporangia.
Conclusions and Further Directions:
Through evolutionary development, plants have become diverse, showing significant variation in vascularity, specialized structure, and reproductive mechanisms. These evolutionary developments are apparent when observing and comparing organisms such as the moss and lily studied in this lab. While writing this lab, I even discovered that even something as simple as grass can be broken down into almost 2,000 species! Fungi have also become more diverse as a result of time and evolutionary development, and clearly have their own unique methods and functions. Such significant differences were seen in this lab when comparing a mushroom and a black bread mold grown in culture. These evolutionary developments continuously allow for plants and fungi to adapt to new and ever changing environments, ensuring their survival for the future. Next week, the invertebrates collected from the Berlese funnel will be examined.
References:
Bentley, M., Walters-Conte, K., Zeller, N. 2015. A Laboratory Manual to Accompany General Biology II. American University Department of Biology: Washington, DC. 34-43.
February 17, 2015
The Examination of Transect Invertebrates
Performed: 2/11/2015
Purpose:
Invertebrates are not simple, malleable blobs that fail to hold any organized structure, in fact, their distinguishing characteristic is simply that they have no vertebral column, also known as a backbone. All animals, invertebrates included, have an organized tissue structure, but the characteristics of this tissue structure are widely diverse. The classification of invertebrates can be broken down into 14 unique phyla (Bentley, Walters-Conte, Zeller, 2015). The complexity of an organism can also be broken down into the categories, radial symmetry, in which the organism is comprised of essentially just a top and a bottom, and bilateral symmetry, which is more complex and developed, containing a top, a bottom, and two sides (Bentley, Walters-Conte, Zeller, 2015). Another way an organism can be categorized is based on the number of different types of germ tissue an organism is comprised of. The three germ tissues each provide the organism with a certain feature. Ectoderm comprises the skin and nervous system, Endoderm comprises the lining of the digestive tract, and mesoderm is responsible for the formation of muscle, bones, organs, and circulatory system (Bentley, Walters-Conte, Zeller, 2015). Invertebrates can broken down even further, in regards to the development of their coelom, a fluid filled cavity which lines their digestive tracts. Acoelomates completely lack a coelom, pseudocoeloms have an incomplete coelom lining, and coelomates have a fully developed coelom (Bentley, Walters-Conte, Zeller, 2015). By observing the specimens found in the transect, we should be able to witness these numerous distinctive features that make invertebrates so diverse.
Materials and Methods:
Procedure 1: A prepared sample of Planaria were placed under a dissection scope, and their characteristics and movement were observed. A prepared sample of nematodes were placed under a dissection scope, and their characteristics and movement were observed. A sample of Annelida, earthworms, were observed, their outer layers of muscle and position of their internal organs were examined. These three specimens collectively account for all three levels of coelom development, acoelomates (Planaria), pseudocoelomates (nematodes), and coelomates (Annelida).
Procedure 2: Samples of each of the five major classes of arthropods were observed, which included insect, arachnida, crustacea, diplopoda, and chilopoda.
Procedure 3: The Berlese tunnels which were set up in last week’s lab were disassembled. The tubes at the bottom of the funnels were retained. The top 10-15 mLs of liquid was poured into a small petri dish. The remaining liquid was poured into a second petri dish. Both samples were examined under a dissection scope. Sometimes the positioning of the organisms were inconvenient, making them difficult to examine, and so they were influenced with the help of a rolled up shred of paper. The arthropods observed were then identified based on keys of images provided to us in the lab manual and on the projector.
Procedure 4: Vertebrates that could pass through or inhabit the transect were considered. Observations from previous experience also provided information for this consideration.
Data and Observations:
Derived from Procedure 1: The Planaria appear to move with ease through the water in a gliding fashion. They appear to be faster than the nematodes that they are enclosed with. The nematodes also appear to move through the water with ease. They seem to glide with less effort than the Planaria, as they do not scrunch up nearly as much. They are noticeably slower than the Planaria. The Annelida were the only specimens observed in soil, and not water. They appeared to have more difficulty travelling through and above the soil, which is understandable, as soil does not provide the same advantages as water in regards to movement (lighter apparent weight, less resistance of motion). The Annelida had to squirm to make any motion at all.
Derived from Procedure 2: Samples of the five major classes of arthropods were examined, along with samples of other invertebrates. There was a tarantula from the class arachnida, a centipede from the class chilopoda, a millipede from the class diplopoda, a crab from the class crustacea, and a few sample of insects. In addition to these arthropod samples, there were also samples that contained an octopus, a squid, starfish, sand dollars, and sponges.
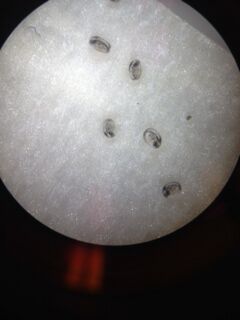
Derived from Procedure 3: From the two samples collected from the berlese funnel (sample from the top and bottom), five invertebrates were identified. There were several springtails found in the sample from the top, as well as a single Thrips. The sample from the bottom contained a soil mite, a couple of ants, and a couple of nematode worms.
Table 4
| Organism | Length (mm) | Number in Sample | Description |
| Springtail Phylum: Arthropoda Class: Insecta | 1 to 6 | 4 | Two antenae, narrow, long structure, six legs, pointed tail |
| Thrips Phylum: Arthropoda Class: Insecta | 3 to 4 | 1 | Two long antenae, long and narrow wings, broad body structure. |
| Soil Mite Phylum: Arthropoda Class: Arachnida | 1 | 1 | Eight short legs, round body |
| Ant Phylum: Arthropoda Class: Insecta | 2 | 1 | Segmented into three sections, six legs, two antenae, black |
| Nematode Worm Phylum: Nematoda Class: Secernentea | 2 | 1 | elongated, round, dark |
Derived from Procedure 4:
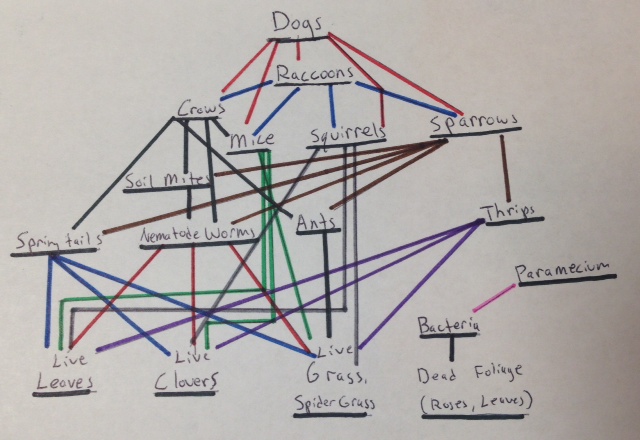
After observing the transect, several vertebrates had been identified, including crows, sparrows, squirrels, mice, dogs, and raccoons. Information about most of these vertebrates can be found in table 5. A food web of the transect can be found in figure 27.
Table 5
| Vertabrate Organism | Classification | Abiotic Factors beneficial to survival | Biotic Factors beneficial to survival |
| Sparrow | Phylum: Chordata Class: Aves Order: Passeriformes Family: Emberizidae Genus: Passer Species: P. domesticus (House Sparrow) | Sunlight, water, dirt, temperature, oxygen | Trees, invertebrates, fruit, seeds, nuts |
| Crow | Phylum: Chordata Class: Aves Order: Passeriformes Family: Corvidae Genus: Corvus Species: C. brachyrhynchos (American Crow) | Sunlight, water, dirt, temperature, oxygen, carrion | Trees, invertebrates, small vertebrates, fruit, seeds, nuts |
| Mouse | Phylum: Chordata Class: Mammalia Order: Rodentia Family: Muridae Genus: Mus Species: A. flavicollis (Yellow-necked field mouse) | Sunlight, water, dirt, temperature, oxygen | plants, grass, fruit, nuts, seeds |
| Raccoon | Phylum: Chordata Class: Mammalia Order: Carnivora Family: Procyonidae Genus: Procyon Species: P. lotor (Raccoon) | Sunlight, water, dirt, temperature, oxygen, garbage | Trees, plants, grass, fruits, seeds, nuts, fish, small vertebrates, invertebrates, eggs, organic wastes |
| Dog | Phylum: Chordata Class: Mammalia Order: Carnivora Family: Canidae Genus: Canis Species: C. lupus (Dog) | Sunlight, water, dirt, temperature, oxygen | Vertebrates, invertebrates, fruit, plants, grass |
Conclusion and Future Directions:
From Procedure 1: The three different invertebrates observed in this lab represent a small sample of invertebrate locomotion. The three specimens observed in this lab each had their own distinct means of motion, even if they did appear similar in some regards. The flat, elongated body structure of the Planaria allows it to move seemingly effortlessly through an aquatic environment. The moments in which it appears to be scrunched up is more than likely due to an undulating motion in movement. The nematode worm is also elongated, but not as flat, as its circumference has some girth to it. With this extra girth, the nematode worm does not appear to rely on an undulating motion in its movement. The Annelida appears to utilize some of its developed muscular system, wiggling its way through the soil in a much less efficient way than Planaria or nematode worms did in the water.
From Procedure 3: The size of the invertebrates observed from the transect in this lab varied, but none were anywhere near a whole centimeter, the largest was 6 mm. The largest invertebrate observed was a Springtail (6 mm), whereas the smallest observed were the soil mite, and another springtail, that were merely 1 mm. All of these invertebrates are arthropods, which is not surprising, as arthropods are the most abundant soil invertebrates (Bentley, Walters-Conte, Zeller, 2015).
From Procedure 4: In reflection to the food web created from all the organisms from the transect observed in lab thus far, the web depicts the complex relationships between all the organisms found within an ecosystem. As members of the same ecosystem, all of these organisms are part of a unique community. A community is a collection of interacting species living in the same approximate vicinity (Bentley, Walters-Conte, Zeller, 2015). As a community, these organisms must share a limited amount of the resources found in the ecosystem. Carrying capacity is the number of organisms that can live within an area (Bentley, Walters-Conte, Zeller, 2015). Carrying capacity varies between species, and typically the larger the organism, the lower the carrying capacity. If the population numbers exceed the carrying capacity, some of the species will die off due to lack environmental resources. There is simply not enough resources to go around. Trophic levels describe the role each organism plays in the ecosystem, more specifically, in regards to that organism’s consumption habits. Trophic levels describe an organism’s place on the food chain, the higher the level, the higher that organism sits on its ecosystem’s food chain (Freeman, Quillin, Allison, 2014). Organisms that share the same trophic levels are likely to consume the same sources of energy, organisms with a lower trophic level, who are lower on the food chain.
Overall, invertebrates, as a collective group, are vastly different. These vast differences were observed in lab this week, whether it was under a microscope or comparing samples that were preserved in jars. The collection of invertebrates includes seemingly everything ranging from simple sponges to malleable jellyfish to hard shelled crabs and lobsters, and even tarantulas. Due to their significant variations, invertebrates can be found in nearly every environment, meaning that they play a crucial role in the global ecosystem. We are able to get a slight grasp of their significance to the world by first observing their significance in our 20 by 20 meter transect.
References:
“American Crow” Wikipedia, The Free Encyclopedia. 16 Feb. 2015. Wikipedia. (18 Feb. 2015).
Bentley, M., Walters-Conte, K., Zeller, N. 2015. A Laboratory Manual to Accompany General Biology II. American University Department of Biology: Washington, DC. 45-49.
“Dog” Wikipedia, The Free Encyclopedia. 17 Feb. 2015. Wikipedia. (18 Feb. 2015).
Freeman, S., Quillin, K., Allison, L. 2014. Biological science (5th ed.). Pearson: Glenview, IL. 1150.
“House Sparrow” Wikipedia, The Free Encyclopedia. 16 Jan. 2015. Wikipedia. (18 Feb. 2015).
“Raccoon” Wikipedia, The Free Encyclopedia. 14 Feb. 2015. Wikipedia. (18 Feb. 2015).
“Yellow-necked mouse” Wikipedia, The Free Encyclopedia. 18 Apr. 2014. Wikipedia. (18 Feb. 2015).
March 3, 2015 16S Sequencing
16S sequence
Purpose: In the week three lab, we examined bacteria colonies which we obtained from our transect samples and grew on petri dishes with agar and agar plus tetracycline bases. We categorized these colonies based on cell and colony morphology. We were stuck with simple, generic classifications, which left us with a very broad realm of possibilities as to what we could have grown on our sample plates. Luckily for us, we were able to prepare a PCR for the bacterial 16S sequencing (via PCR amplification). PCR amplification is a process which utilizes DNA or RNA polymerase, and cuts DNA into sections of varying length, which are run through an agarose gel, with some help of an electrical current (Bentley, Laslo, Walters-Conte, Zeller, 2014). Larger pieces of DNA fragments have a more difficult time moving through the gel, and do not travel very far from their original well, whereas smaller pieces of DNA fragments can move through the gel with more ease, and travel further away from the gel. Scientists are able to determine the length of the DNA fragments produced by comparing the fragment’s location at the end of the run to a ladder, which is placed right beside the sample. The DNA amplification process allows scientists to determine where the DNA under observation came from, in this case, as specific as species. By identifying the species of bacteria which can be found within our transect, we will gain a deeper understanding of our transect, and the complex web of relationships that can be found within it.
Materials and Methods:
Procedure 1 (Set up) (During lab 3): Four bacterial colonies were selected, two from nutrient plates, and two from nutrient plus tetracycline plates. The four colonies selected were then placed into four separate, sterile tubes, which contained 100 microliters of water. The tubes were clearly marked to indicate which contained they contained. The tubes were then capped, and heated on a heating block for ten minutes at 100 degrees Celsius. The samples were then centrifuged for five minutes at 13,400 rpms. While waiting for the centrifuge to complete its cycle, 20 microtimes of a primer/water mixture was added to a PCR tube, which was clearly labeled to indicate what bacteria sample it would hold. The PCR tube was then shaken in order to dissolve the PCR bead inside. Five microliters of supernatant were then removed from the centrifuge tubes and placed in the PCR tubes. The PCR tubes were then placed in the PCR machine.
Procedure 2 (analysis):
A few lab meeting later, the PCR results were ready. Groups retrieved the codes from the TA page of Openwetware.com, Biology 210 at AU, Lab Section 6. The retrieved codes were then run through a BLAST program, which identifies species based on genetic data input. The codes provided to us were from the bacteria colonies retrieved from the 10^-7 nutrient plate and the 10^-7 plus tetracycline plate. Species identification and characteristics was provided as a result of the BLAST, and comparisons were made to the bacteria morphologies and bacteria colony morphologies which were made back in week three lab.
Results:
Figure 28 shows that three of the four samples which we ran had thick bars similar to the bars indicated in the PCR guide sheet which was in the lab. From these three respectable samples, we were allowed to pick two more for further analysis. We chose to run the samples from the 10^-7 nutrient plate and the 10^-7 nutrient plus tetracycline plate.
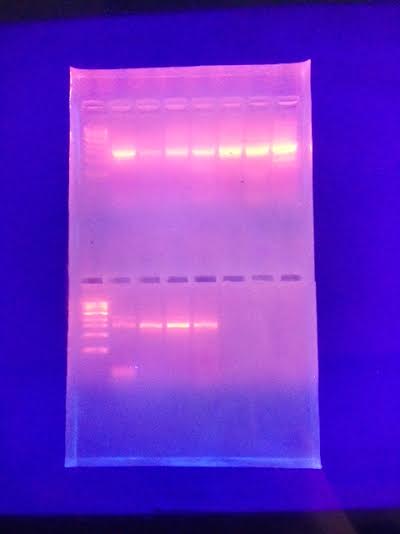 (Figure 28)
(Figure 28)
The following two series of letters are the forward sequences identified from the two samples selected for further analysis.
Forward sequence of 10^-7 nutrient NNNNNNNNNNNNNNCNNNNNNTGCAGTCGAGCGAATGGATTGAGAGCTTGCTCTCAAGAAGTTAGCGGCGGACGGGTGAG TAACACGTGGGTAACCTGCCCATAAGACTGGGATAACTCCGGGAAACCGGGGCTAATACCGGATAATATTTTGAACTGCA TGGTTCGAAATTGAAAGGCGGCTTCGGCTGTCACTTATGGATGGACCCGCGTCGCATTAGCTAGTTGGTGAGGTAACGGC TCACCAAGGCAACGATGCGTAGCCGACCTGAGAGGGTGATCGGCCACACTGGGACTGAGACACGGCCCAGACTCCTACGG GAGGCAGCAGTAGGGAATCTTCCGCAATGGACGAAAGTCTGACGGAGCAACGCCGCGTGAGTGATGAAGGCTTTCGGGTC GTAAAACTCTGTTGTTAGGGAAGAACAAGTGCTAGTTGAATAAGCTGGCACCTTGACGGTACCTAACCAGAAAGCCACGG CTAACTACGTGCCAGCAGCCGCGGTAATACGTAGGTGGCAAGCGTTATCCGGAATTATTGGGCGTAAAGCGCGCGCAGGT GGTTTCTTAAGTCTGATGTGAAAGCCCACGGCTCAACCGTGGAGGGTCATTGGAAACTGGGAGACTTGAGTGCAGAAGAG GAAAGTGGAATTCCATGTGTAGCGGTGAAATGCGTAGAGATATGGAGGAACACCAGTGGCGAAGGCGACTTTCTGGTCTG TAACTGACACTGAGGCGCGAAAGCGTGGGGAGCAAACAGGATTANATACCCTGGTAGTCCACNCCGTANCNATGANTGCT AANTGTTAGAGGGTTTCCGCCCTTTANTGCTGAANTTAACGCATTAAGCACTCCGCCTGGGGAGTACNGGCCNNNANGGC TGAANCTCAAANGAATTGACNGGGGCCCGCACNAGCGGTGGNANCATGTGGTTTAATTCNAANCANCGCNAANANCCTTN NCNNGTCGTGNCNTGCGTCNGAAAACCCTNANATNGGCTTCTNNNTCCGNAGCNNANTNGACNNNTGGTGCATGNTGTNN NNNTCNNNTCNNNNNATNTTGGNNNANTCNNNCANNNNNNCAACCNTGNNNTTNNNTGN
Forward sequence of 10^-7 nutrient plus tetracycline NNNNNNNNNNNNNCNANNNATGCNAGCCGAGCGGTAGAGATTCTTCGGAATCTTGAGAGCGGCGTACGGGTGCGGAACAC GTGTGCAACCTGCCTTTATCAGGGGGATAGCCTTTCGAAAGGAAGATTAATACCCCATAATATATTGAATGGCATCATTT AATATTGAAAACTCCGGTGGATAGAGATGGGCACGCGCAAGATTAGATAGTTGGTGAGGTAACGGCTCACCAAGTCTACG ATCTTTAGGGGGCCTGAGAGGGTGATCCCCCACACTGGTACTGAGACACGGACCAGACTCCTACGGGAGGCAGCAGTGAG GAATATTGGACAATGGGTGCGAGCCTGATCCAGCCATCCCGCGTGAAGGACGACGGCCCTATGGGTTGTAAACTTCTTTT GTGTAGGGATAAACCTAGATACGTGTATCTAGCTGAAGGTACTATACGAATAAGCACCGGCTAACTCCGTGCCAGCAGCC GCGGTAATACGGAGGGTGCAAGCGTTATCCGGATTTATTGGGTTTAAAGGGTCCGTAGGCGGATTTGTAAGTCAGTGGTG AAATCTCACAGCTTAACTGTGAAACTGCCATTGATACTGCAAGTCTTGAGTGTTGTTGAAGTAGCTGGAATAAGTAGTGT AGCGGTGAAATGCATAGATATTACTTAGAACACCAATTGCGAAGGCAGGTTACTAAGCAACAACTGACNCTGATGGACGA AAGCGTGGGGAGCGAACAGGATTANATACCCTGGTNNTNNNNGCCGTNNCNATGCTAACTCNTTTTTGGAGCGCAANCTT CANAGACTAAGCGAAAGTGATAAGTTANCCACCTGGGGANTANNANCGCAAGTTTGAAACTCANAGGAATTGACGGGGGG CCCGCACAANCNGNTGGATTATGTNGTTTAATTCNATGATACGCGAGNANNCNGTACCNANGGCTTAANNGGGNAATNAC GGGGTNNANNAANNNACTTNTCTTNNACATTTTTNNANGTGCTNNANGGGTNNNTNNNTCNNCTCNTGCCCNTCAGNNNT ANGNTGANNTCCNNNNACNANNNNNNANCCCCTNNNCNNNNNNNCGATCNNNANGNNNNNGGNNNNCNNNNNNNNNNNNN CCTACGGNNNNNNNNN
The bacteria from the colony on the 10-7 nutrient plate was identified as (best matched to) Bacillus cereus. The bacteria from the colony on the 10-7 nutrient plus tetracycline plate were identified as (best matched to) Chryseobacterium indologenes. The observed bacteria and colony morphology observed in week three were are juxtaposed to known characteristics of the bacteria identified (Table 6)
(Table 6)
| Species | Colony Description | Cell Description | Gram +/- |
| 10^-7 nutrient | White, Undulate, Raised, Glistening, Circular, Irregular edge | Bacillus, Rods, Motile, No flagella seen, no chains or clusters. | Gram + |
| Bacillus cereus | White or Gray, Undulate, Raised, Smooth or Rough, Irregular Edge | Bacillus, Rods, Motile, Small chains. | Gram + |
| 10^-7 nutrient + tet. | Orange, Smooth, Glistening, Round, Convex, Entire edge | Cocci, Staphylococci. | Gram + |
| Chryseobacterium indologenes | Deep Yellow, Smooth, Round, Convex, Entire Edge | Rod, Non-motile | Gram - |
(Bottone, 2010) (Cunniff, 2012)
Conclusion:
Reference Table 6 for details. For the majority of both cell and colony morphology, our BLAST result was accurate in comparison to the observations we made and recorded. Despite having a majority of characteristics shared by the bacteria in our sample and the BLAST result, there were still some differences upon further research. Why don't they all match? In the case of the comparison of our 10^-7 nutrient plus tetracycline plate to the Chryseobacterium indologenes, there were a few differences. In the case of cell description, we observed a cocci, or a staphylococci, whereas the source describes a rod. In images of the bacteria, they do indeed look round. The difference could have been as simple as a misclassification on our part and not fully understanding the inclusive range of bacillus’ rod shape. The bacteria observed was also Gram positive, as opposed to the source which described the cell to be Gram negative. In this case, it was our first gram stain. We made conclusions about the cell’s nature based on observations of our first Gram stain. Mistakes during this process are not unlikely. Especially when we are using judgement to see how much dye the cell retained.
References:
Bentley, Meg. Laslo, Mara. Walters-Conte, Kathryn. Zeller, Nancy. 2014. American University Lab Manual for General Biology I. American University: Washington, D.C.
Bottone, E. 2010. Bacillus cereus, a Volatile Human Pathogen. Clinical Microbiology Reviews, 23(2): 382–398.
Cunniff, J. 2012. Chryseobacterium indologenes. Kenyon College, Gambier,OH.
ZEBRA FISH
PURPOSE
Drugs and chemicals can greatly affect development and quality of life experienced by an individual, each with different effects.This experiment will measure the effects of an estrogen mimic, Ethisterone. Estrogen is the primary female sex hormone. Despite stimulating natural estrogen, estrogen mimics can cause side effects. In order to observe the effects that Ethisterone has on development, we will be observing the development of Zebrafish. Zebrafish are considered model organisms because of their clear egg sacs, translucent bodies in their embryonic to juvenile stages, and a short amount of time required to reach adulthood. In this experiment, we will administer Ethisterone at a low concentration.
Variable being tested: Estrogen Mimic (25 mg/L concentration)
March 3, 2015
Visitation 1 (2/18/2015):
Materials and Methods
To start, 20 mLs of Deer Park water was removed from a large volumetric flask using a automated pipette, and was then added to a petri dish. This petri dish would serve as the control group, as it contained strictly Deer Park water, and no other additives. The petri dish was then labeled “PHBT Control” using tape. Next, 20 mLs of Deer Park water which contained 25 mg of Ethisterone per Liter was removed from the large volumetric flask labeled “Ethisterone 25 mg/L” using an automated pipette, and was placed in another petri dish. This dish was labeled “PHBT Estrogen mimic” using tape. Once the petri dishes were prepared, it was time to select the Zebrafish embryos. The selection process was strictly based on convenience and appearance of the eggs. We were advised to stay away from eggs that appeared to be encompassed in “fuzz”. Due to the nature of this selection process, the groups cannot be considered random samples of the Zebrafish embryo population as a whole, just ones that appear healthy. First, 20 seemingly healthy embryos were removed from their original container using a transfer pipette and placed in the petri dish containing the control water. Then, 20 more seemingly healthy embryos were removed from the same container via a transfer pipette and placed in the petri dish which contained the low concentration of Ethisterone. The embryos were then observed under a dissection microscope, and their developmental stages were recorded. These results can be seen in Table 1. The lids were then put on their respective petri dishes, and secured with multiple pieces of tape. The dishes were then carefully placed in the lab section’s tub.
Hypothesis: The estrogen mimic will increase the growth rate and final size of the specimens which are grown in its presence.
Prediction: If the embryos are exposed to an estrogen mimic, then the embryo development will be quicker and the final specimen samples will be larger.
Results
Upon further observation, under a dissection microscope, all of the eggs appeared to be in the development stage somewhere around somite stage, with one exception, which appeared to be be more developed, possibly the 33 hour stage. These results can be seen in table 1.
Table 1: Developmental Stages Witnessed Post Selection and Distribution
| Stages | Ethisterone (count) | Control (count) |
| Somite Stages | 19 | 21 |
| 33 hour stage | 1 | 0 |
Visitation 2 (2/20/2015):
Materials and Methods
The petri dishes were retrieved from the section’s tub, and brought to a table for observation. In this observation, counts were recorded of how many embryos were still in their eggs, along with their stages of development, how many embryos had hatched, how many bad eggs there were, and how many hatched individuals had died. These results, as well as survival rates, can be seen in Table 2. After observations were complete, a clean transfer pipette was used to remove debris from the water in both samples. This process was done in a careful way to minimize the volume of water taken from each sample. Each petri dish was then replenished with 5 mL of their respective solutions via an automated pipette. Pictures were then taken of the embryos, and can be seen in figures 1 and 2. The petri dishes were then covered, secured with tape, and placed in the section’s tub.
Results
Table 2: Stages and Hatching Rate Observed on Visitation 2
| ' | Estrogen Mimic | Control |
| Still in eggs, stage | 20, 36/48 hour stages | 19, 36/48 hour stages |
| Hatched | 1 | 1 |
| Bad Eggs | 0 | 1 |
| Dead Hatched | 0 | 0 |
-Notified that development might take longer, due to the lab not being 80 degrees or so.
Visitation 3 (2/23/2015):
Materials and Methods
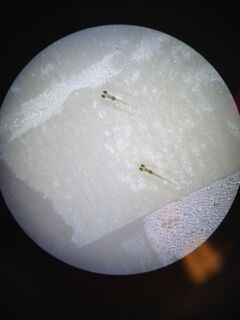
The petri dishes were retrieved from the section’s tub, and placed under a dissection microscope for initial observations, which entailed recording how many embryos were still in their eggs, how many embryos had hatched, how many dead eggs there were, and how many hatched individuals had died. These observations, and survival rates, can be seen in Table 3. Further observations, which focused on the development of characteristics and behavior, were then made. These results can be seen in Table 4. Once the observations were complete, and information was recorded, the petri dishes were replenished with 5 mLs of their respective liquid using an automated pipette. Any visible debris was removed from the dishes via a transfer pipette. Pictures were taken of both groups, as can be seen in figures 3 and 4. The petri dishes were then covered, secured with tape, and placed back in the section’s bin.
Results
Table 3: Counts Collected Under the Dissection Microscope on Visitation 3
| ' | Estrogen Mimic | Control |
| Still in eggs | 1 | 3 |
| Hatched | 19 | 17 |
| Bad Eggs | 0 | 0 |
| Dead Hatched | 0 | 0 |
| Survival rate from previous trip | 100% | 100% |
Observations: Ethisterone: -light green, light, speckled black -eye size = 10 micrometers under 4x magnification -size: full ocular lens -Rapid movement, long pauses at edge of plates, sudden turns -good body…?
Control: -one alive, but shows signs of illness, moves slowly when touched, barely responsive to touch. -skinnier, larger head, smaller tail -light green/ darker than Ethisterone -rapid movement, would not remain still in dissection glass.
Visitation 4 (2/25/2015):
Materials and Methods
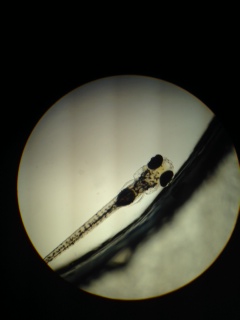
The petri dishes were removed from the section’s bin and placed under a dissection microscope for initial observations. Results of these observations can be seen in Table 4. Further observations were then made of one selected specimen from each group, using of a microscope outfitted with an ocular micrometer. Selected specimens were withdrawn with the use of a transfer pipette, and were placed in a depression slide. The results of these observations can be seen in Table 6, and pictures can be seen in figures 5,6 and 7. These specimens were then transferred to a capsule, and exposed to an anesthetic, Tricaine solution at a concentration of 1 drop per mL of water present. The samples were left for the TA to add paraformaldehyde, for preservation purposes. Then, an automated pipette was used to remove 10 mLs of liquid from each petri dish, and transfer it to a waste-containing bowl. During this procedure, the flashlight on my phone was used to attract the fish to one side of the container, so water could be withdrawn from the other side without abducting any fish. 13 mLs of respective liquid was then added to each dish via an automated pipette. Once the water had been changed, it was time to feed the fish for the first time. The fish had not been fed prior to this visitation, due to the fact that for Zebrafish, eating is a learned habit, they would not know what to do with the food otherwise. One drop of solution containing the microorganism, Paramecium, which would serve as food for the Zebrafish for duration of this experiment, was added to each petri dish using the pipette which was found in the food container. The petri dishes were then covered, secured with tape, and put back in the section’s bin.
Results
Table 4: Counts Collected Under the Dissection Microscope on Visitation 4
| ' | Estrogen Mimic | Control |
| Still in eggs | 0 | 0 |
| Hatched | 20 | 19 |
| Dead Eggs | 0 | 1 |
| Dead Hatched | 0 | 0 |
| Survival rate from previous trip | 100% | 95% |
| Removed for observation | 1 | 1 |
Observations (of the samples selected for staging): Estrogen mimic: Measurements (under 4x magnification): 160 x 25 = 4,000 micrometers -clear space in heart?
Control: Measurements (under 4x magnification): 180 x 25 = 6,300 micrometers -full heart?
Collective samples are attracted to the light of the microscope. Noticeable migration to stay under the light.
Visitation 5 (2/27/2015):
Materials and Methods
Both dishes were retrieved from the section’s bin and brought to a dissection microscope for observation. Results of these observations can be seen in Table 5. Pictures were then taken and can be seen in figures 8 and 9. Then 10 mLs of liquid was removed from each petri dish. The flashlight tactic used on visitation 4 was duplicated. 10 mLs of respective solutions were added to each petri dish using an automated pipette. Visible debris was removed from both dishes using a transfer pipette. No food was given on this day. The dishes were then closed, secured with tape, and placed back in the section’s bin.
Results
Table 5: Counts Collected Under the Dissection Microscope on Visitation 5
| ' | Estrogen mimic | Control |
| Still in eggs | 0 | 0 |
| Hatched | 19 | 18 |
| Dead Eggs | 0 | 0 |
| Dead Hatched | 0 | 0 |
| Survival rate from previous trip | 100% | 100% |
Visitation 6 (3/2/2015):
Materials and Methods
Upon removing the dishes from the section’s bin, it was noticed that the water in each dish was grotesque. Observations were made using a dissection microscope, and the results of these observations can be seen in Table 6. 12 mLs of liquid were then removed from each petri dish via an automated pipette, and was transferred to a waste-containing bowl. The dishes were then replenished with 12 mLs of respective liquids using an automated pipette. Any remaining debris was removed with a transfer pipette. The dishes were then covered, secured with tape, and put back into the section’s container.
Results
Table 6: Counts Collected Under the Dissection Microscope on Visitation 6
| ' | Estrogen Mimic | Control |
| Hatched | 19 | 18 |
| Dead Hatched | 6 | 7 |
| Survival rate from previous trip | 68.40% | 61.10% |
-Water was noticeably disgusting
Observations: -No noticeable differences observed between two plates
(Final day of observations)
Visitation 7 (3/4/2015):
Materials and Methods
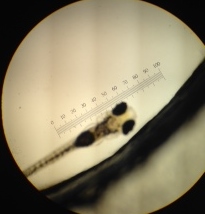
The dishes were retrieved from the section’s bin. Initial observations were made under a dissection scope, and can be seen in Table 7. Two specimens from each group were then transferred to two separate depression slides using a transfer pipette, and were observed under a microscope outfitted with an ocular lense. Data from these observations can be seen in Table 10. Pictures were taken of both groups under the microscope., and can be seen in figure 10, 11 and 12. We were then given back our fixed specimens which were preserved from visitation 4. These specimens were also observed under the microscope outfitted with an ocular lense, and the results can be seen in Table 11. Both dishes were emptied out into their properly labeled bowls.
Results
Fish counts to bee seen in Table 6.
Table 6
| ' | Estrogen Mimic | Control |
| Alive Hatched | 0 | 0 |
| Dead Hatched | 13 | 11 |
| Survival rate from previous trip | 0.00% | 0.00% |
All of the fish died.
Observations: The mimic group appears to be less decomposed than control group.
-Control are only fain outlines with black dots swarming them.
-Mimic- darker pigmentation is still present, and are more easy to see and make out. Being swarmed by black and gray dots. Some characteristics such as eyes can be made out in a couple of specimens.
Body measurements of fish taken during visitation 7 can be seen in Table 7, where the body size of two fish from each group were recorded. (Used the bodies that could still be identified, recorded, and measured).
Table 7: Counts Collected Under the Dissection Microscope on Visitation 7
| ' | Etrogen Mimic | Control |
| Body size: micrometers | 3,750 | 3,750 |
| 3,500 | 3,750 | |
Measurements regarding the fish taken for observation in week 1 and placed in paraformaldehyde lab can be seen in Table 8.
Table 8: Final Measurements of Fixed Specimens from Visitation 4
| ' | Etstrogen Mimic | Control |
| Right Eye Size: micrometers | 400 | 375 |
| Left Eye Size: micrometers | 375 | 425 |
| Tail Width: micrometers | 75 | 100 |
Notes: The estrogen mimic specimen looks like a typical fish, where are the control specimen looks deformed with an oddly shaped body. This could just simple be from the paraformaldehyde process though.
Conclusions:
The purpose of this experiment was to identify the effects that the estrogen mimic, Ethisterone, plays on embryos during the developmental stages.
It was hypothesized that the presence of the estrogen mimic, Ethisterone, will increase the growth rate and final size of the specimens exposed to it during development. If embryos are exposed to the estrogen mimic, Ethisterone, then the embryo development will be quicker and the final specimens will be larger.
Using Table 1 and Table 2, we see that the speedy development is not wholly true, as they were at similar stages entering their groups, and were the same stages 2 days later. There was more eggs left to hatch in the control group on visit 3, but only by a count of 3 to 1, and one of those 3 turned out to be a bad egg. 2 is not a significantly larger number than 1. Overall the control group had 2 bad eggs, whereas the Ethisterone group had none. Maybe Ethisterone reduces the number of bad eggs by nurturing them during development. In conclusion in regards to development rate, there is a lack of a trend in the data, which does does not support the idea that Ethisterone increases the rate of development.
As far as developmental characteristics goes, there was no visible difference between the two groups after visit 3, where there was a visible difference in pigmentation, head size, and tail length between the two groups. In the last visit, all specimens from both groups were relatively the same size. The Ethisterone group was less decomposed than the Control group, however. This could be an indicator for time of death. Decmposition levels make it appear as though the control group died first, as they were more decomposed. It is a possibility that Ethisterone kept the fish alive a little longer.
There was no significant difference in sizes of features measured from the fixed specimens. The lack of differences in size measurements does not support the idea that Ethisterone increases the size of the embryos. This does not support the proposed hypothesis.