User:Paola Quinones Lopez/Notebook/Lab 3 Bacteria
Paola Quinones Lab 3: Microbiology & Identifying Bacteria
I. Introduction The purpose of this lab is to quantify the number of bacteria, observe the diversity of bacteria, and observe the diversity of the morphological characteristics of the bacteria found in the Hay Infusion. In this lab we also tested for naturally occurring antibiotic resistance in bacteria, identified bacteria based on their colony, morphology, motility, cell shape & gram stain and use the PCR and DNA sequencing to verify the identification of bacterial species.
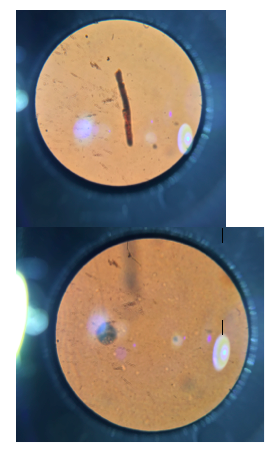
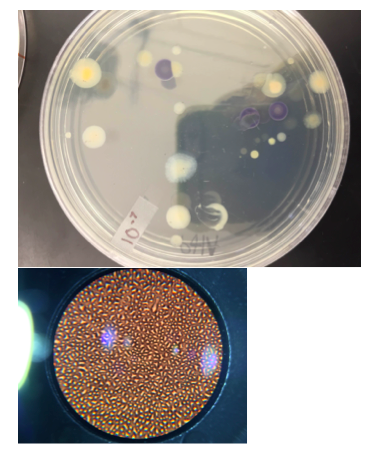
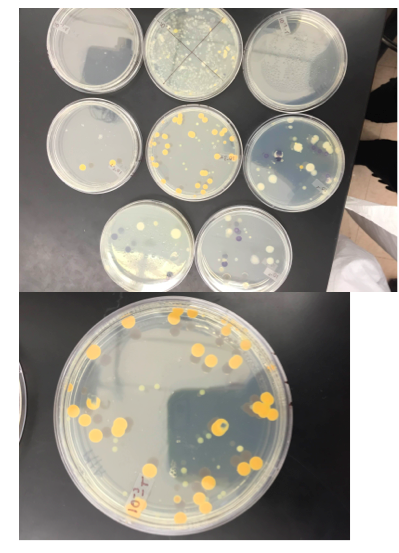
II. Methods & Materials For the Procedure I, we observed the growth on the agar plates from the Hay Infusion. We counted the number of colonies on each of the plates and annotated it on Table 1. We then converted the number of colonies to colonies/mL. For Procedure 3, we observed prepared slides of bacteria & then observed a native wet mount preparation & a gram stain of 2 defined colonies from the nutrient agar plate & from the tetracycline plate. We then prepared 1 wet mount & 1 gram stain for each of the 2 nutrient agar plates and nutrient agar plus tetracycline plates, Then we used a compound microscope and the 100x oil immersion objective to observe our prepared slides. To make the wet mount, we sterilized a loop with a flame and scraped up a tiny amount of growth from the surface of the agar. Then, a drop of water was added on the slide and a cover slip was placed over the drop. We observed our wet mount in the microscope. We observed the organisms shape & motility. For the gram stain, we sterilized a loop and scraped growth from the agar and added a drop of water like in the wet mount. Then, with a staining tray, we covered the bacterial smear with crystal violet for 1 minute and then rinsed the stain off using a bottle of water. After, we covered the bacterial smear with Gram’s iodine mordant for 1 minute and rinsed it off like before. We decolorized by flooding the bacterial smear with 95% alcohol for 10-20 sec and rinsed it gently. Then, we covered the smear with safranin stain for 30 sec and rinsed the stain with water. After the slide is dry, we observed it in the microscope. For the PCR Amplification, we selected one from two plates of the nutrient agar and one from the two tetracycline plates. We used the primer and PCR to amplify the 16S rRNA gene. We labeled the 2 PCR tubes. We added 20 ul of primer/water mixture to one of the tubes. Mix it to dissolve PCR bead. Using a toothpick, we picked a small amount of bacteria from one of the colonies and then submerged the toothpick in the primer/water mix of a PCR tube & twist for 5 seconds. The tube was then placed in the PCR machine.
III. Results
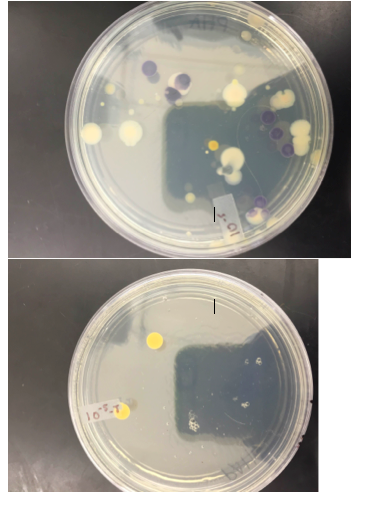
The colonies in the plates with antibiotic are less and much smaller than those without antibiotic, this is evident in 10-7 and 10-9 which have antibiotic, in which there are no colonies at all. This may indicate that bacteria that were affected by the antibiotic haven’t developed the resistance necessary to survive when antibiotic is present. Those who have colonies have developed the necessary resistance. The effect of tetracycline on the number of bacteria and fungi is that it results in them having less colonies or none. Only two species of bacteria are unaffected by tetracycline.
IV. Tables and Graphs (Data)