User:Marie-Eve Val


Marie-Eve VAL
INSTITUT PASTEUR
Unité "Plasticité du Génome Bactérien"
Département Génomes et Génétique
CNRS UMR3525
28 rue du Dr. Roux
75724 PARIS cedex 15
FRANCE
Email me through OpenWetWare
Research interests
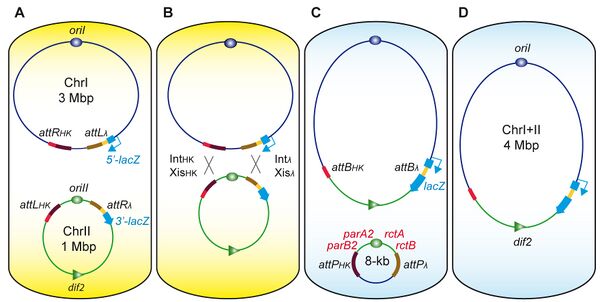
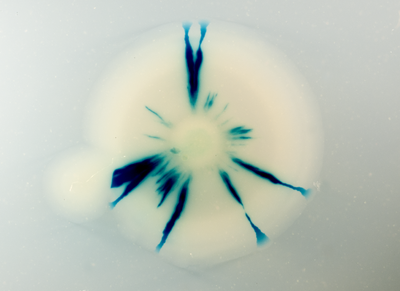
I am a research associate in Didier Mazel's lab at the Institut Pasteur of Paris. I have a great interest in bacterial genome structuration, organization and maintenance ... and a special interest in multipartite genomes. Owing to its bi-chromosomal genome architecture and its importance in public health, Vibrio cholerae, the causative agent of cholera, has become a preferred model to study bacteria with multipartite genomes. My approach is to drastically alter V. cholerae’s genome structure to gain more insight into multipartite genomes. To do so, we developed a site-specific recombination-based engineering tool, which provides us with a powerful mean to massively reorganize in principle any prokaryotic genome.
Education
- 2008 - PhD
Centre de Génétique Moléculaire (Gif-sur-Yvette) , Francois-Xavier Barre's lab
Study of the system of chromosome dimer resolution in Vibrio cholerae and its implication in the control of the lysogeny of the phage CTX coding for the choleric toxin - 2003 - Engineering degree in biotechnology and Master in cellular and molecular biology
ESBS (Strasbourg, Freiburg, Karlsruhe and Basel) - 1999 - BS in Biotechnology
LVC (Gif-sur-Yvette)
Professional Training
- Since 2013 - INSERM Research Associate
Institut Pasteur (Paris) - Didier Mazel's lab - 2009 / 2013 - Post-Doctoral research fellow
Institut Pasteur (Paris) - Didier Mazel's lab - 2004 / 2008 - PhD Thesis
Centre de Génétique Moléculaire (Gif-sur-Yvette) , Francois-Xavier Barre's lab - 2004 - Engineer position
Génoscope - Consortium National de Recherche en Génomique (Evry) - Génolevures Consortium - 2003 - Master thesis
Center of Marine Biotechnology, UMBI (Baltimore) , Hal Schreier's lab - 2002 - Engineer training
IGBMC (Strasbourg) - Olivier POCH's lab - 2001 - Engineer training
Biozentrum (Basel) - Peter Philippsen's lab - 1999 / 2000 and summer 2002 - Technician position
AGROGENE(Moissy-Cramayel) - 1999 - Internship
Centre de Génétique Moléculaire (Gif-sur-Yvette) ,Fred Boccard's lab
Publications
- Val ME, Marbouty M, de Lemos Martins F, Kennedy SP, Kemble H, Bland MJ, Possoz C, Koszul R, Skovgaard O, and Mazel D. A checkpoint control orchestrates the replication of the two chromosomes of Vibrio cholerae. Sci Adv. 2016 Apr;2(4):e1501914. DOI:10.1126/sciadv.1501914 |
A checkpoint control orchestrates the replication of the two chromosomes of Vibrio cholerae
- Soler-Bistué A, Mondotte JA, Bland MJ, Val ME, Saleh MC, and Mazel D. Genomic location of the major ribosomal protein gene locus determines Vibrio cholerae global growth and infectivity. PLoS Genet. 2015 Apr;11(4):e1005156. DOI:10.1371/journal.pgen.1005156 |
Genomic location of the major ribosomal protein gene locus determines Vibrio cholerae global growth and infectivity
- Val ME, Soler-Bistué A, Bland MJ, and Mazel D. Management of multipartite genomes: the Vibrio cholerae model. Curr Opin Microbiol. 2014 Dec;22:120-6. DOI:10.1016/j.mib.2014.10.003 |
Management of multipartite genomes: the Vibrio cholerae model
- Val ME, Kennedy SP, Soler-Bistué AJ, Barbe V, Bouchier C, Ducos-Galand M, Skovgaard O, and Mazel D. Fuse or die: how to survive the loss of Dam in Vibrio cholerae. Mol Microbiol. 2014 Feb;91(4):665-78. DOI:10.1111/mmi.12483 |
Fuse or die: how to survive the loss of Dam in Vibrio cholerae
- Val ME, Skovgaard O, Ducos-Galand M, Bland MJ, and Mazel D. Genome engineering in Vibrio cholerae: a feasible approach to address biological issues. PLoS Genet. 2012 Jan;8(1):e1002472. DOI:10.1371/journal.pgen.1002472 |
Genome Engineering in Vibrio cholerae: A Feasible Approach to Address Biological Issues
- Das B, Bischerour J, Val ME, and Barre FX. Molecular keys of the tropism of integration of the cholera toxin phage. Proc Natl Acad Sci U S A. 2010 Mar 2;107(9):4377-82. DOI:10.1073/pnas.0910212107 |
Molecular keys of the tropism of integration of the cholera toxin phage
- Génolevures Consortium, Souciet JL, Dujon B, Gaillardin C, Johnston M, Baret PV, Cliften P, Sherman DJ, Weissenbach J, Westhof E, Wincker P, Jubin C, Poulain J, Barbe V, Ségurens B, Artiguenave F, Anthouard V, Vacherie B, Val ME, Fulton RS, Minx P, Wilson R, Durrens P, Jean G, Marck C, Martin T, Nikolski M, Rolland T, Seret ML, Casarégola S, Despons L, Fairhead C, Fischer G, Lafontaine I, Leh V, Lemaire M, de Montigny J, Neuvéglise C, Thierry A, Blanc-Lenfle I, Bleykasten C, Diffels J, Fritsch E, Frangeul L, Goëffon A, Jauniaux N, Kachouri-Lafond R, Payen C, Potier S, Pribylova L, Ozanne C, Richard GF, Sacerdot C, Straub ML, and Talla E. Comparative genomics of protoploid Saccharomycetaceae. Genome Res. 2009 Oct;19(10):1696-709. DOI:10.1101/gr.091546.109 |
Comparative genomics of protoploid Saccharomycetaceae
- Val ME, Kennedy SP, El Karoui M, Bonné L, Chevalier F, and Barre FX. FtsK-dependent dimer resolution on multiple chromosomes in the pathogen Vibrio cholerae. PLoS Genet. 2008 Sep 26;4(9):e1000201. DOI:10.1371/journal.pgen.1000201 |
FtsK-dependent dimer resolution on multiple chromosomes in the pathogen Vibrio cholerae
- Val ME, Bouvier M, Campos J, Sherratt D, Cornet F, Mazel D, and Barre FX. The single-stranded genome of phage CTX is the form used for integration into the genome of Vibrio cholerae. Mol Cell. 2005 Aug 19;19(4):559-66. DOI:10.1016/j.molcel.2005.07.002 |
The single-stranded genome of phage CTX is the form used for integration into the genome of Vibrio cholerae