User:Lila Glansberg/Notebook/Biology 210 at AU
Embryology and Development of Zebrafish Study -- March 18, 2015
Purpose
The purpose of this lab is to learn about the embryology and stages of development. A lot can be learned about an organism by looking at its development: specialization of cells and structures, for example. The comparison of different organisms' development further distinguishes organisms apart; that is, the development of similar organisms can be very different from one another. This can assist in the definition of species. This lab also provides and opportunity for students to set up their own experiments to test the effects of environmental factors on the development of zebrafish. In this lab, I study the effects of salinity on zebrafish development. I hypothesize that the zebrafish in the saline environment will develop more slowly than the fish in freshwater.
Materials and Methods
-healthy, translucent zebrafish embryos -petri dishes -test variable solution (I observed the effects of salinity on development) -paramecium (fish food) -dropper -paraformaldehyde
Procedure: Select a variable (light, salinity, acidity, etc) to test on the zebrafish. Set up the control group by adding 20 mLs of water to a petri dish, and placing in 20 healthy, translucent zebrafish embryos with a dropper. For the test group, add 20 mLs of the solution with the tested variable in it, and place in 20 embryos. Organize an observation schedule for the next two weeks. On day 4-5, remove 10 mLs of water/solution and any dead embryos or empty egg cases, and add 25 mLs of water/solution. Save any dead embryos in paraformaldehyde. On day 7, remove 5 mLs of water/solution with any egg cases or dead specimen, and add 5 mLs water/solution. Preserve 1-3 embryos from both dishes in paraformaldehyde. Between day 7 and 14, remove 5 mLs and add 10 mLs. Starting after day 7, feed the fish two drops of paramecium--this can be done just when the water is changed. On day 14, make final observations on the fish. Survivors will be kept in an aquarium.
Results
Day 1: 1 specimen dead in the control group, 2 dead in the test group. None are hatched in either group.
Day 4: 1 more specimen dead in control group (total of 17 alive), 5 more dead in test group (total of 13 alive). All of control group is hatched and swimming. None of the test group are hatched.
Day 7: 4 more dead in control (13 alive), all specimens in test group dead. None of the test group had evacuated their egg cases before death. In the control group, the body and tail were just becoming pigmented. There was no strong heartbeat to be observed, though I would assume their heart was beating. Pectoral fins were just barely visible. These were hatchlings being observed, so there was no yolk sac. The swim bladder could not be observed. The fish usually move in bursts of about 2 mm. When startled or provoked, the hatchlings moved in bursts of about 2 cm.
Day 11: All fish are dead.
Day 14: All fish are dead.
Conclusion
The purpose of this lab was to observe the effects of an environmental variable, salinity in this lab, on the development of zebrafish embryos. I hypothesized that the fish in saline solution would develop more slowly than the fish in freshwater. I accept this hypothesis. The fish in freshwater came out of their cases before the experimental group fish did. In fact, the experimental group never left their egg cases at all. The experimental group died before the control group, as well, suggesting that the salt in the water may have outright stopped development, killing the specimens. However, a few errors were made with this lab, as the control group eventually died as well. The class petri dishes were stored stacked on top of each other in a Tupperware container. This meant that students with petri dishes on the bottom had to move other students' dishes out of the way to get to their own, spilling other dishes' water in the process. This caused the death of my own control group, and potentially my experimental group. Evaporation of water from the petri was also an issue.
LG
DNA Sequencing Bacteria: 16S Sequence
Purpose
The purpose of this lab was to identify the species of bacteria living on students' transects. This was done through the sequencing of the 16S gene of the bacteria.
Methods
DNA from two selected plates of bacteria were isolated and amplified through PCR technique. The 16S rRNA gene was selectively amplified and run on an agarose gel. This sequence was then used to identify the species of bacteria. The 16S gene is large (about 1500 base pairs long), has remained more or less unchanged by evolution, and is found in almost all species of bacteria, making it a great choice to use to ID bacterial species with (Janda and Abbott, 2007). The sequence specific to each group's bacteria was provided. In order to ID their bacteria, students put this sequence into "BLAST," an online program supported by the National Institute of Health in Washington, D.C.. This program is able to identify potential species when provide with a nucleotide sequence, such as the one that the students isolated from their bacteria. Genus species of the bacteria was identified in this way.
Results
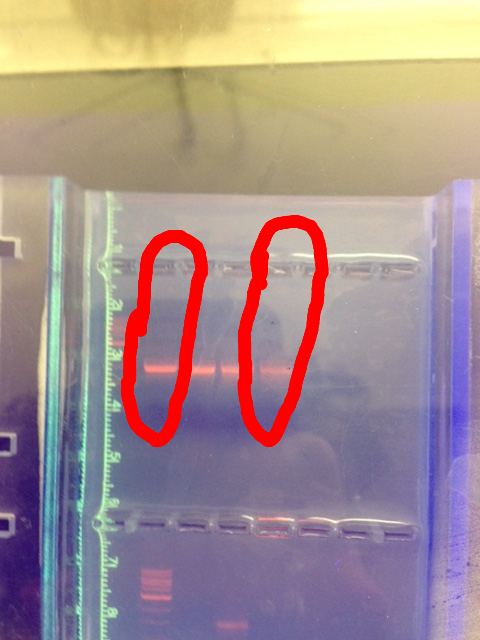
This image is of the gel electrophoresis run of the bacterial DNA. The two circled columns are the Samples A and D, from right to left, respectively. These were the ones chosen for sequencing.
The raw sequence for bacterial Sample A was NNNNNNNNNNNNGNNNTTANNNNTGCAGTCGNNCGANNGAGTAGCNCNNNNTNNCGGACGCTGACGAGTGGCGAACGGGT GAGTAATACTATCGGAACGTGCCCAGTCGTGGGGGATAACTACTCGAAAGAGTAGCTAATACCGCATACGATCTGAGGAT GAAAGCGGGGGACCTTCGGGCCTCGCGCGATTGGAGCGGCCGATGGCAGATTAGGTAGTTGGTGGGATAAAAGCTTACCA AGCCGACGATCTGTAGCTGGTCTGAGAGGACGACCAGCCACACTGGGACTGAGACACGGCCCAGACTCCTACGGGAGGCA GCAGTGGGGAATTTTGGACAATGGGCGAAAGCCTGATCCAGCAATGCCGCGTGCAGGATGAAGGCCTTCGGGTTGTAAAC TGCTTTTGTACGNAACGAAAAAGCTTCTCCTAATACNAGAGGCCCATGACGGTACCGTANNAATAAGCACCGGCTAACTA CGTGCCANCAGCCGCGGTANTACGNANGGTGCGAGCGTTNATCGCGANTTTACTGNGCGTAAAGCGTGCGCANGCNGNTA TGTNANGACANNATGTGAAATCCCCNNNNTCTNTCCTGNTNAACTGNCNTTTGTGNCTGCNNGGNTAAANTACGGNAGAG GNGTGNTGGAAT.
BLAST identifies this sequence as being from an "uncultured bacteria." In other words, the genus species of this bacteria is unknown.
The raw sequence for bacterial Sample D was NNNNNNNNNNNNNNNNNNNNNNTGCAGCCGAGCGCTANGTTTCCTTCGNGGAGGACTGNAGANANNCGCACGGGTGCGGA ACATGTGTGCAACCTGCCTTTATCAGGGGGATAGCCTTTCGAAAGGAAGATTAATACCCCATAATATTTTGAGTGGCATC ACTTGAAATTGAAAACTCCGGTGGATAAAGATGGGCACGCGCAAGATTAGATAGTTGGTGAGGTAACGGCTCACCAAGTC TACGATCTTTAGGGGGCCTGAGAGGGTGATCCCCCACACTGGTACTGAGACACGGACCAGACTNNNNCGGGAGGCAGCNG TGAGGAATATTGGACGATGNGTGAGAGCCTGATCCAGCCATCCCGCGTGAAGGACGACGGCCCTATNGGTTGTAAACTTC TTTTGTATAGGGATAAACCTTTCCACGTGTGGGANGCTGAAGGTACTATACGAATAAGCACCGGCTAACTCCGTGCCAGC NGCCGCGGTAATACGGAGGGTGCAAGCGTTATCCGGATTTATTGGGTTTAAAGGGTCCGTAGGCGGATCTGTNAGTCAGT GGTGAAATCTCACAGCTTAACTGTGAAACTGCCATTGATACTGCNGGTCTTGAGTGNTGTTGAAGTAGCTGGAANAAGTN GTGTANCGGTGAAATGCATAGATNTNACNNACAACACCANTNGCGAAGGNNGGTTACTAANCAACAACTGANGCTNNNGG ACGAAGCGTGGGGAGCGAACAGGATTAGATACCCNGGTANTCCACGCNTNNNCGANGCNNNTCCGTTTTNGNTTTCNGAT CNNAGNCNAANCGAANGTGANCAGTAGCCACTGGGNGNNTNCGNCGNANTNNNNANNNTCNNNNNATTGACNGGGGNNNN CNNNNNNNGNNNNTTNTGTCNTTANTCNNNNANNNCGCNNGGNNCNNNNNNNCNNANNNGNNNNGNNAGNNNNNNNNNCN TNNNNGNNTTNNNNGNNNNNNNNNNNNNCANCNNNCCGNNNNGNNNANTANNNNNGNNNNNNCNNNNC.
According to BLAST, the bacteria with this sequence is Chryseobacterium sp. B.
Both of these sequences are listed from the forward primer.
Conclusion
Chryseobacteria is a gram-negative bacteria. It usually grows in lawns, and has a yellow-ish cream color when plated (Vandamme, Bernardet, Segers, Kersters, and Holmes, 1994). This matches the data collected on Sample D, as can be seen in Lab 3. Sample D was a gram-negative bacteria, that was cream colored and grew in a lawn with bubble-like projections. The fact that Sample D matches the description of Chryseobacterium further emphasizes that Sample D is Chryseobacterium.
References
Janda, JM & Abbott, SL. 2007. 16S rRNA Gene Sequencing for Bacterial Identification in the Diagnostic Laboratory: Pluses, Perils, and Pitfalls. Journal of Clinical Microbiology. 45:2761-2764.
Vandamme, P, Bernardet, JF, Segers, P, Kersters, K, and Holmes, B. 1994. New Perspectives in the Classification of the Flavobacteria: Description of Chryseobacterium gen. nov., Bergeyella gen. nov., and Empedobacter norn. rev.. International Journal of Systematic Bacteriology. 44:827-831.
LG
Lab 5.2: Vertebrates
Purpose
Most of the other types of life in the transect have been identified: plant life, protists, bacteria, fungi, and invertebrates. This means that vertebrates are the only type of life left to be classified in the transect. In this lab, students will learn about what kind of vertebrates inhabit their transects, thus completing the study of biological life of their transects at American University.
Methods
The observations of the transect were made at about 4:30 on February 12, 2015. It was cloudy and windy outside, and the temperature was about 25°F.
Results
The organisms seen were:
Gray squirrel, Sciurus carolinensis
Deer (only droppings and footprints were spotted in the transect, though this is sufficient evidence to acknowledge that deer inhabit the transect), Odocoileus virginianus
Robin, Turdus migratorius
Cardinal, Cardinalis cardinalis
Dog, Canis familiaris
Discussion/Conclusion
The vegetables growing in the garden are a clear source of food for the animals listed above, with exception of the dog. This biotic factor attracts the animals. The fertile soil would likely have worms in it, which is the attraction to the robin and cardinal, in addition to the seeds produced by the plants and scattered by humans. The dog was there as a human's pet; the garden is taken care of by humans, and someone brought along their dog. As this is a highly residential area, seeing a dog is not uncommon. Squirrels would find many materials to make a nest with as there are woodchips and bushes scattered around the transect. Deer, much to the planters' dismay, are attracted by the well-grown and large vegetables as a reliable food source.
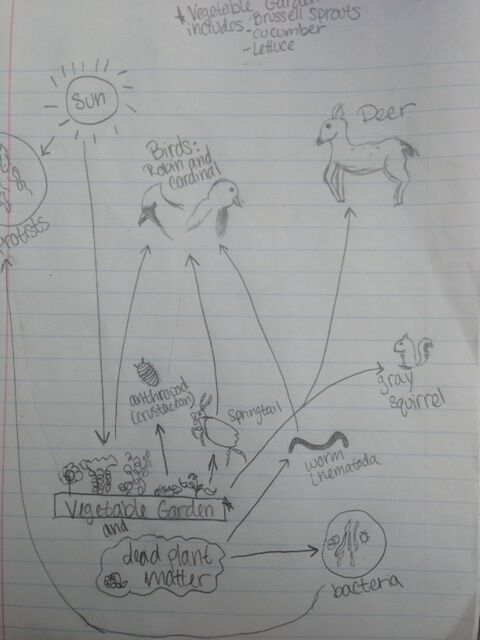
Figure 1: Food Web of Vertebrates in Transect
A community is all of the living species in an area. This includes all biotic matter: protists, bacteria, fungi, plants, animals, etc. Throughout the last five labs, we have been working to define the community of our transect. The life in the above food web is representative of the community of the transect.
Carrying capacity describes how many organisms a biological ecosystem can support, based on how many resources are available. Each of the organisms in the food web are using up resources and contributing to the area's carrying capacity.
Trophic levels describe where an organism is in the food chain in relation to producers; for example, if an organism is on the first trophic level then it feeds on producers, if an organism is on the second trophic level then it feeds on first-level organisms, and so on. The trophic levels can be clearly seen in the food web. The vegetables in the garden are the producers, as well as some protists, and first-level, or primary, consumers are worms, springtails, and anthropods, squirrels, and The secondary consumers include the robin and the cardinal. Bacteria functions as a decomposer, or an organism that feeds on, and breaks down, dead material.
LG
Lab 5.1: Invertebrates
Purpose
In this lab, students will gain an understanding of invertebrates and how they evolved. Students will learn that invertebrates come in wide varieties and can be quite complex. There is very high variation in all of these creatures (i.e. compare a sea sponge to an insect); during this lab students will learn to appreciate this by observing preserved specimens and observing invertebrates found in their own transect.
Materials and Methods
Materials needed for these procedures are: -Planaria sample -Dissecting scope -Nematodes -Annelida -Preserved example organisms -Berlese funnel prepared during Lab 4 -Dichotomous Key for Insecta class
Procedure 1: Observe Planaria with the dissecting scope. Note the type of movement these organisms use to get around. Observe a cross section of the organism with a microscope, and note the stained digestive tract. Observe the nematodes and a cross section of their structure. Note their movement. Observe the Annelida. Observe their internal organs and layers of muscle and note their type of movement.
Procedure 2: Many of the invertebrates in your transect will be arthropods, and so it is important to be able to classify the different classes of these organisms. Observe the preserved example organisms from each of the five major classes. These classes are arachnida, diplopoda, chilopoda, insect, and crustacea. These classes are defined by body parts, body segments, and number of appendages.
Procedure 3: Break down the Berlese Funnel and pour the top 10-15 mL of liquid and organisms into one petri dish, and pour the remaining liquid into another dish. Probe around with a pencil to better see the organisms, as they will be mixed with debris. Observe the dishes with a dissecting scope, and identify the class of any arthropods you observe. Most will be insects, so you may use the Dichotomous key to identify insects. Identify at least five invertebrates, and document their descriptions into a table.
Data and Observations
The Planaria specimens slide in straight across to their destination. This makes sense as their bodies are cylindrical and elongated, with no visible muscles. The nematodes are slow, wormlike, and for the most part just wriggle around in place. Their bodies start of wider, and are more tapered at the opposite end, which is why much of their movement is head-driven. The annelids wiggle and twist, with their rings of muscles bunching and unbunching. They are very long and thin, and are surrounded in pink rings of muscle with a central vein going down the middle that can be seen with a dissecting scope.
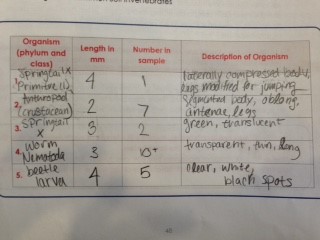
Table 1: Invertebrates in Transect #4
This table describes five invertebrates from Transect #4 (farmland). Most were arthropods, however specimen #4 belonged to the phylum nematoda.
Conclusion
The size range of the organisms was 2-4 mm. The primitive organism Springtail X and the beetle larvae were the largest at 4 mm. The smallest organism were the anthropods at 2 mm. The most common organism was the nematode worms, as there were 10+ in the sample taken.
LG
Lab 4: Observing Plants and Fungi
Purpose
The purpose of this lab is to understand the diversity of the plant world and observe plants in your transect. This lab will also focus on learning about the function and place of fungi in an ecosystem. I hypothesize that most of the plants found in my transect will have been planted and grown, and wouldn't usually grow naturally in this environment. Students will also learn how to distinguish monocots from dicots, and recognize the different structures in plants.
Materials and Methods
-Ziploc bags -phone camera (recommended) -moss -Lily plant -agar plate of black bread mold (Rhizopus stolonifer) -50:50 ethanol/water solution -Funnel -screening material -50 mL conical tube
Procedure 1: Obtain two Ziploc bags and bring them to your transect. Take a leaf litter sample by finding an area with soil and dead leaves. Put the dead leaves and plant matter into one of the Ziploc bags. This will be used later to set up the Berlese funnel.
Take representative samples from five plants in the transect. Make sure to do this in way that is minimally damaging to the plant. Choose a diversity of plants, and take a picture of each specimen that you take a sample from so you can remember it accurately when you study the sample in the lab. If you take a sample of the tree, make sure to take a picture of the entire tree.
Procedure 2: Compare the moss with the provided angiosperm (lily). Think about the differences in their xylem and phloem. Examine the cross section of a lily stem and find the xylem and phloem.
Procedure 3: Examine moss leaves with a dissection scope or a low magnification compound scope. Note the cuticle, which is indicated by the smooth surfaces.
Procedure 4: Examine Polytrichum. Identify the male and female gametophyte and sporophyte. Dissect a lily flower and identify the anther, pollen, stigma, and style. Observe and record information on the plant samples taken from your transect. Record their location in the transect, size and shape, vascularization, specialized structures, and mechanisms of reproduction.
Procedure 5: Examine the black bread mold in a petri dish with a dissecting scope, leaving the top on.
Procedure 6: Pour 25 mL of the 50:50 ethanol/water solution into the 50 mL conical tube. Place a piece of screening material into the bottom of a funnel. You may want to tape the screen to the funnel to ensure that no leaf litter falls into the preservative. Place the leaf litter sample in the funnel. Set up the funnel on a ring stand on a ring stand so that it's held in the tube with ethanol. Wrap parafilm around the spot where the tube and funnel meet to prevent the ethanol from evaporating. Put the funnel under a 40 watt lamp. Cover everything with foil. Leave the Berlese funnel on the lab bench for one week.
Data and Observations
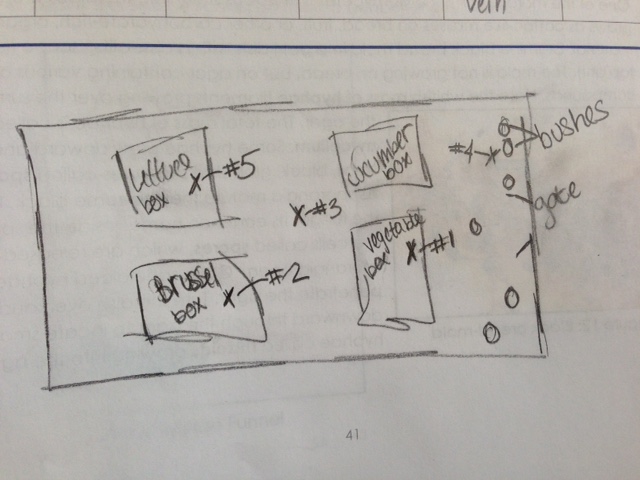
Figure 1-Transect Sample Plant Locations
This figure shows where each sample was taken from in the transect. Each X designates a spot where the transect was taken from, and the number designates which sample was taken from that spot.
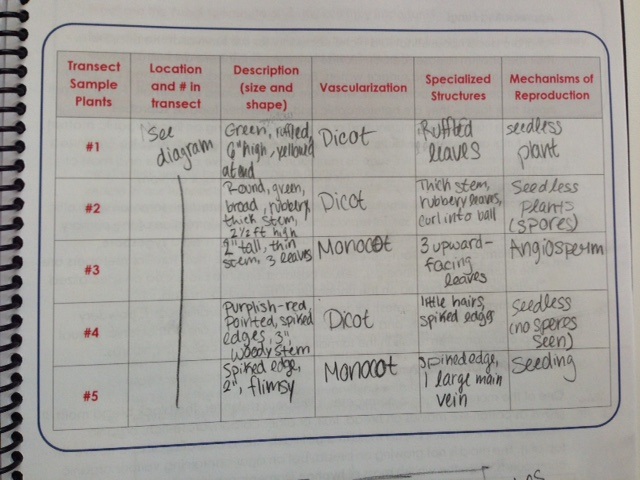
Table 1-Transect Sample Plant Observations and Information
This table shows the observations made about each plant sample taken from the transect. I believe sample #1 to be of the genus Brassica. Sample #2 is also the genus Brassica. Sample #3 is the genus Trefolium. Sample #4 is Lycophyta. Sample #5 is Lactuca.
Sample #1's leaves were ruffled and spread out in a fan, much like a lily. It was very close to the ground. Sample #2's leaves were tightly rolled up in small balls, and attached to a 2.5 foot-high stalk. This plant had spores on the underside of the leaf. Sample #3's leaves were grouped in a cluster of three, and spread out evenly from one another. The plant stood about 2 inches off the ground. Sample #4's leaves stood by themselves, apart from each other, and were dispersed sparsely on the plant's woody stem. The plant was about 2 feet tall. Sample #5's leaves were all clustered in a large, loose ball, about 5 inches off the ground.
Figure 2-Fungi Under Microscope
The darkened spots are the sporangia.
Conclusion
Fungi sporangia are the structures that contain cells called spores in the fungi. Spores are released when the sporangia open. Spores function similarly as seeds, though are structurally different. Spores help the fungi reproduce. Sporangia are therefore important; keeping the spores safe is in the fungi's best reproductive interests. Safe spores mean a new generation of fungi.
I predict that there will be different types of bugs in my transect, such as isopods. I would expect the invertebrates to thrive off of dead plant matter (particularly vegetables).
LG
Lab 3: Microbiology and Sequencing Bacterial DNA--February 4, 2015
Purpose
The purpose of this lab is to learn what types of bacteria are living in your transect, and classify about them. This involves doing a number of procedures, including sequencing the bacteria's DNA. To classify bacteria, students must understand characteristics of this domain. Students will also learn about antibiotic resistance in organisms and how DNA sequencing can be used to identify an organism. The students will also make one last observation of their hay infusion culture. I hypothesize that the hay infusion culture changes from week to week because it continues to grow; it has a supply of oxygen and nutrients and therefore the microorganisms present are still able to survive and reproduce.
Materials and Methods
Materials required for this lab are: -bacterial colonies plated last lab (8 plates) -sterile tube -light microscope -heat block -prepared slides of bacteria -centrifuge -loop -PCR tube -Bunsen burner -primer/water mixture -crystal violet stain -PCR machine -Gram's iodine mordant -95% alcohol -safranin stain
Procedure 1: Observe the growth on the plates taken from the Hay Infusion Culture. Count the total number of colonies on each plate, paying attention to which plate contains antibiotic, and what its dilution is. Record in Table 1.
Procedure 2: Looking at Table 1 and the plates of bacteria and answer the questions provided in the lab manual (See conclusion). Determine the mechanisms of tetracycline and what bacteria it affects, making sure to cite sources.
Procedure 3: Observe a prepared slide of varying types of bacteria. Make a wet mount of your own bacteria by sterilizing a loop over a flame and scraping up a tiny amount of growth from the agar. Mix this growth with a drop of water on a slide. Place a cover slip on top, and observe the wet mount at 10X and 40X.
To gram stain your bacteria, start by sterilizing a loop over a flame and scraping up a small amount of growth as you did with the wet mount. Mix the growth with a drop of water on a slide, and circle the area where the bacteria is. Make sure to label the slides. After letting it air dry, pass the slide over a flame about three times, with the bacteria side up. Using a staining tray, cover the smear with crystal violet for 1 minute. Rinse off the slide with water. Next, cover the smear with Gram's iodine mordant for 1 minute and rinse with water. Decolorize the bacterial smear by flooding it with 95% alcohol for 10-20 seconds. Gently rinse with water. Decolorization has happened when solvent flows colorlessly from the slide. Cover the slide with safranin stain for 20-30 seconds, and rinse the stain off using water. Blot excess water from the side of the slide and allow it to air dry. Focus the stained sample at 10X, and then observe under 40X and 100X. Record your observations in Table 2.
Procedure 4: Select one of each type of plate (tet or non-tet) that was used for characterization. Transfer a single colony of bacteria to 100 μL of water in a sterile tube. Incubate the tube at 100°C for 10 minutes in a heat block. Centrifuge the samples for 5 minutes a 13,400 rpm. While this is happening, add 20 μL of primer/water mixture to a labeled PCR tube. Mix to ensure that the PCR bead dissolves. Transfer 5 μL of supernatant from your centrifuged samples to the 16S PCR reaction. Place the tube in the PCR machine.
Data and Observations
The Hay Infusion Culture has changed from last week. The water contained is much lower; about 1/2 of the original water is present. The fuzzy white clump has gotten larger, and the scent of sewage has gotten stronger. There are still some brussel sprout leaves present. The thin skin and veiny network have both shrunk. The culture is more clear, and soil is still collected on the bottom.
There are differences between the plated colonies with tetracycline and without tetracycline. The colonies without the antibiotic are lighter, more numerous, and more translucent. The colonies with tetracycline are yellower, larger, and more sparse than those without it.
The colonies picked for characterization were 10^-3 with tetracycline (colony A), 10^-5 with tetracycline (colony B), 10^-7 without tetracycline(colony C), and 10^-3 without tetracycline(colony D). Drawings of each sample drawn under a microscope can be seen below. There is no drawing for colony A because it wasn't able to be found under a microscope.
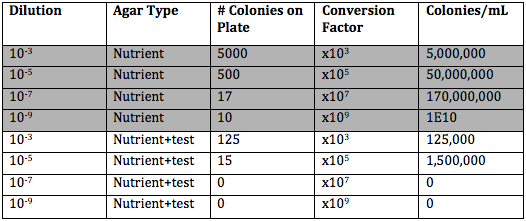
The following table, Table 1, shows the results from the serial dilution performed last lab.
The plates without tetracycline grew more than the plates with tetracycline. However, neither type of plate grew anything for a 10^-9 dilution.
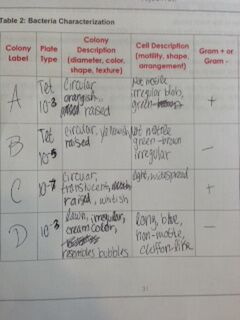
Table 2 shows the characterization of the bacteria derived from observations.
Colonies A and C are gram positive, meaning they have a thick layer of peptidoglycan in their membranes. Colonies B and D are gram negative, meaning they have very little peptidoglycan in their membranes.
Conclusion
I do not think any Archaea species have grown on my agar plates. Arechaeans only grow in extreme conditions, such as hot vents in the ocean floor. The agar plates were not kept at any extreme temperatures, just the room temperature of the lab. It seems unlikely that any would grow in these conditions. The differences between the tet and non-tet agar plates has been described in the data section. The effect of tetracycline on bacteria growth was that bacteria grew much slower and in larger, denser colonies than the plates without any antibiotic. Less bacteria grew on plates with tetracycline as opposed to the plates without tetracycline. There was no fungi growing on any of my plates, so it is hard to distinguish tetracycline's effects on fungi.
It is thought that tetracycline penetrates bacterial cells through passive diffusion. The antibiotic then inhibits bacterial growth by either destroying the bacterial membrane or interfering with protein synthesis (Schnappinger and Hillen, 1996). Tetracycline is commonly used to treat acne and rosacea today, and was important in the treatment of cholera ("WHO Model Lists of Essential Medicines," 2013).
References Schnappinger, D. & Hillen, W. (1996) Tetracyclines: Antibiotic action, uptake, and resistance mechanisms. Archives of Microbiology, 165, 359-369. Retrieved from http://american.summon.serialssolutions.com/document/show
(October 2013) WHO Model Lists of Essential Medicines Retrieved from http://www.who.int/medicines/publications/essentialmedicines/en/
Lab 2: Observing and Identifying Algae and Protists--January 28, 2015
Purpose
The purpose of this lab is to discover and identify any protists living in the transect. This is necessary to further the understanding of ecosystems in the natural world. Students must learn and understand common methods of identifying organisms, such as the use of a dichotomous key. Another purpose of the lab is to prepare for the next class: students will also dilute and plate bacteria that is found in the Hay Infusion Culture.
Materials and Methods
The materials needed for this lab are: -light microscope -dichotomous key -known organism -Hay Infusion Culture (made last class) -micropippetor -four nutrient agar plates -four nutrient agar plus tetracycline plates
Procedure 1: Observe wet mount of a known organism with a microscope. Focus on one organism and describe it and record its size. Use a dichotomous key to determine what the organism is.
Procedure 2: Note the smell and appearance of the culture after bringing it to your desk. Draw samples from two different niches, including some plant matter, and note exactly where in the culture you took the samples from. Observe wet mounts of the samples taken under a microscope, and using the dichotomous key, identify what protists and algae are present. Diagram the organisms observed and measure their size.
Procedure 3:
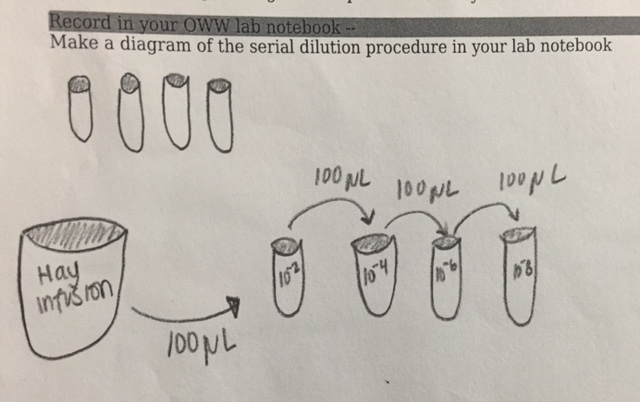
Label four tubes of 10 mL sterile broth with 10^-2, 10^-4, 10^-6, and 10^-8. Set a micropippetor to 100 microliters. Obtain four of each kind of agar plate--label all of the tetracycline plates with "tet." Label each of plate with their respective numbers, and add your lab group's info on the plate. Swirl the Hay Infusion Culture--with the lid on-- to mix up the contents. Add 100 microliters from the culture to the tube labeled 10^-2. Mix the tube thoroughly. Add 100 microliters of broth from this tube to the 10^-4 tube and swirl to mix well. Repeat this twice to make the 10^-6 and 10^-8 test tubes. Pipette 100 microliters from each tube onto their respective nutrient agar plate (10^-2 goes on the plate labeled 10^-3, etc.), and spread it around the plate carefully. Repeat this procedure for each nutrient agar plus tetracycline plate. Place the agar plates in a safe spot in the lab where they will incubate at room temperature.
The following is a diagram of the serial dilution procedure:
Data and Observations
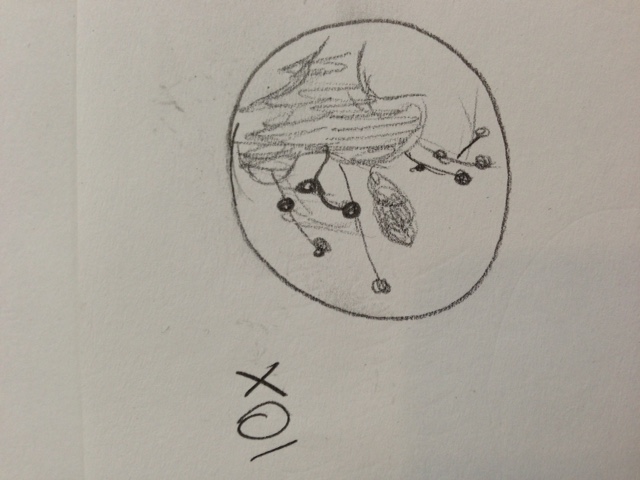
The infusion culture smelled like spoiled or rotten vegetables. Its appearance was brown and gooey, with a thin skin on top and soil resting on the bottom. On the top, there formed a network of vein-like structures, in which there floated a fuzzy white clump, about the size of a nickel, two-thirds of which was submerged.
Below is a diagram of the culture as seen from above, and each X represents where a specific sample was taken from.
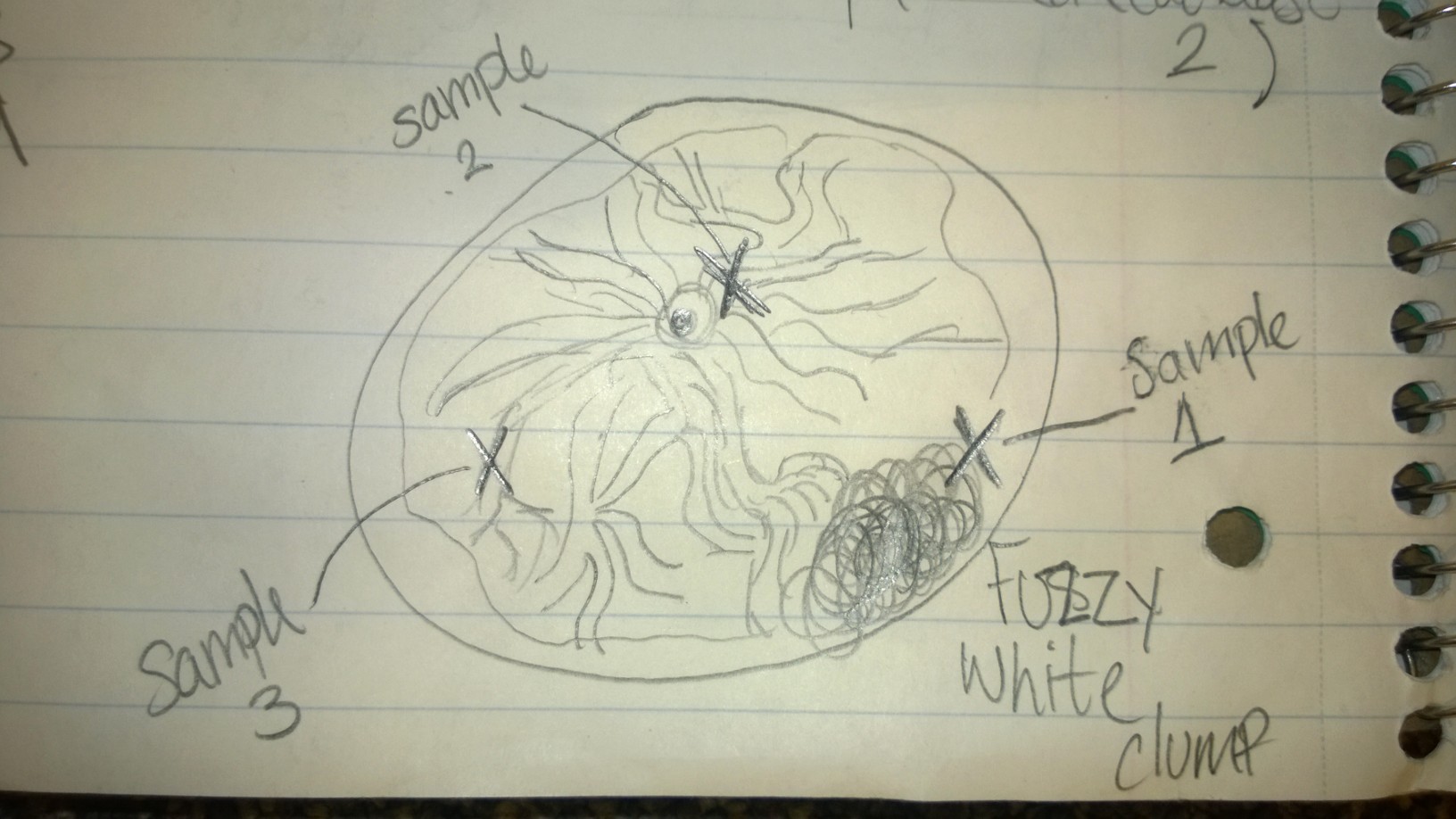
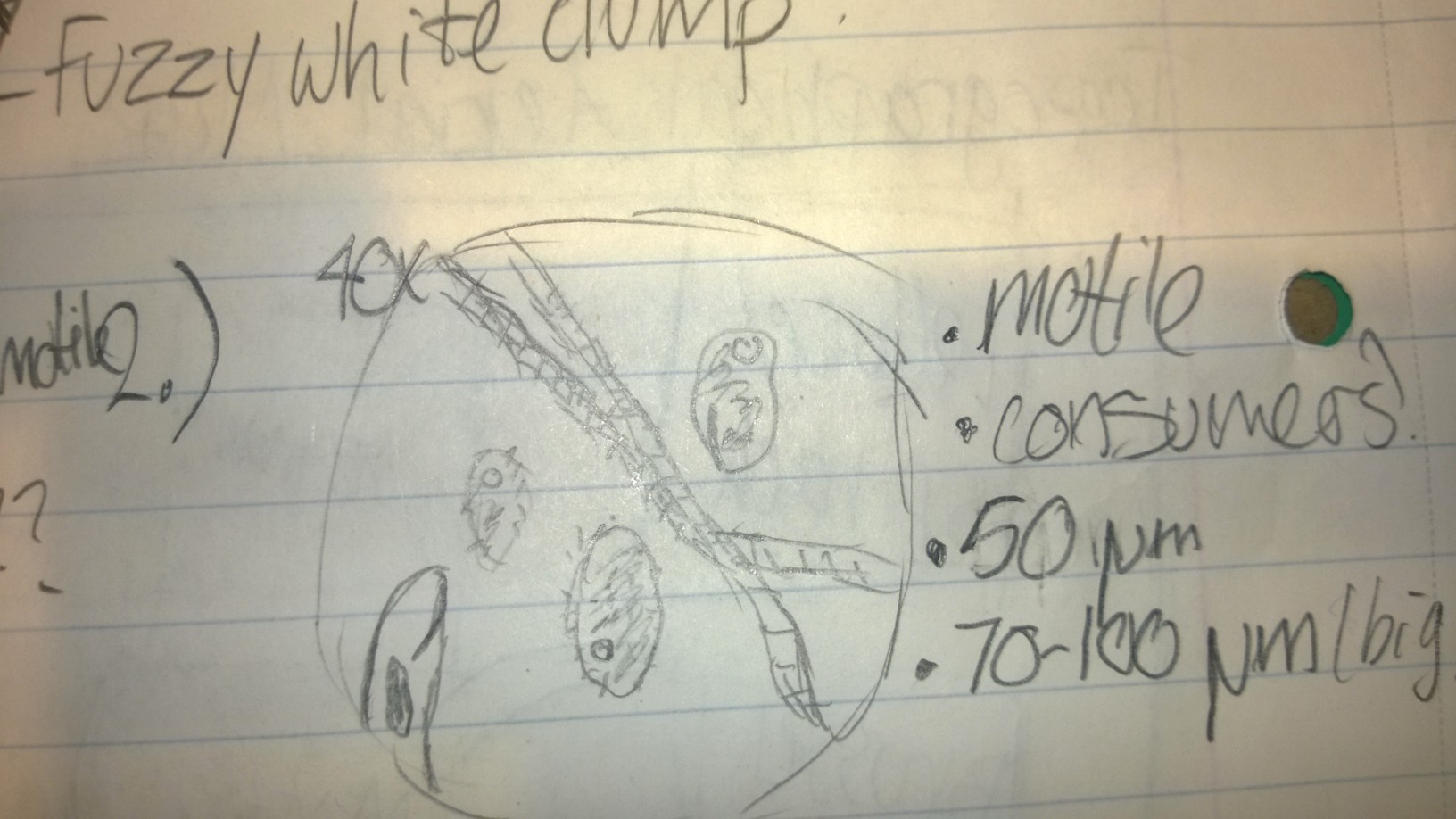
Sample 1 contained a non-motile species that I was unable to identify with the dichotomous key. It had some green parts, leading me to believe it was a species that performed photosynthesis. It was about 50 micrometers.
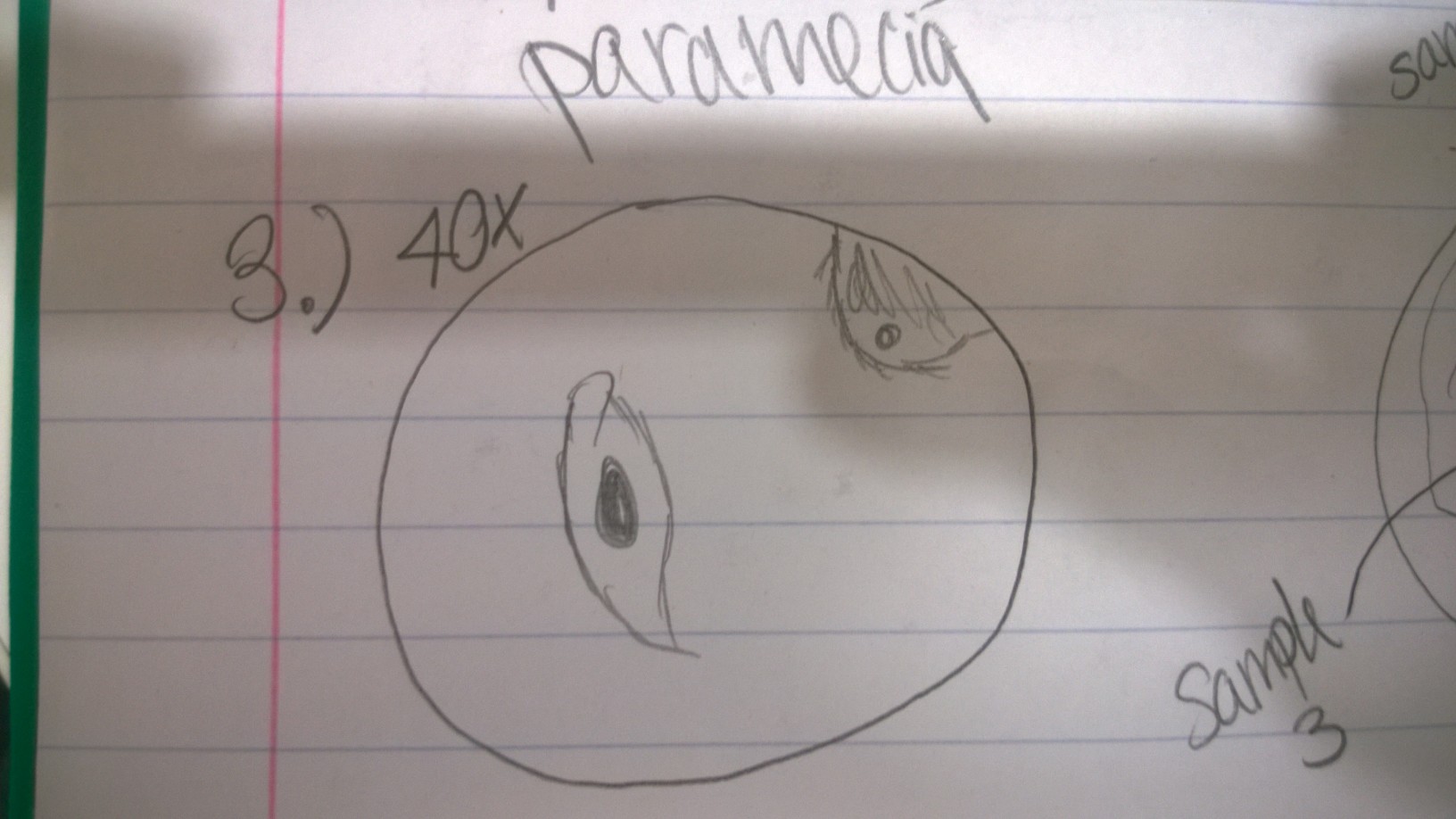
Sample 2 contained a variety of organisms, including colpidium, at least two different types of paramecia, and an organism I suspected to be some kind of worm. The colpidium were roughly 50 micrometers, and the paramecia ranged from 70-100 micrometers.
Sample 3 contained more colpidium and various paramecia.
Conclusion
The protists found in the hay infusion culture were both heterotrophs and autotrophs. The consumers are competing for food, while the autotrophs are competing for sunlight to produce their own nutrients with. If the hay infusion culture grew for another two months, I would predict the smell to become more potent, the number of protest species to approach carrying capacity of the ecosystem, and for the white clump to get larger. As each population approached carrying capacity, the selective pressures on each organism would increase: organisms must now be very selectively fit in order to survive, as there is more competition.
LG
Lab 1: A Study of Evolution and Ecosystems at AU--January 25, 2015
Purpose
The purpose of this lab is to begin to understand the different types of life that exist on Earth. It is necessary to understand the basics of evolution before one studies diversity, as all life forms evolved from a common ancestor. By examining a transect of an ecosystem close to home, students can understand the high diversity of life that exists even in small areas.
Materials and Methods
Materials needed for this lab are: -samples of Chlamydomonas, Gonium, and Volvox -light microscope -soil/vegetation sample from transect -water -dried milk
Procedure 1: Prepare wet slides of and observe the three types of algae under a microscope. Note the number of cells contained by each specimen, the colony size, any cell specialization, and motility mechanisms. Note if the organism is isogamous or oogamous. Draw a picture of the organism under the microscope, and note the magnifying power.
Procedure 2: Examine the assigned transect and make a map of the area. Make a list of five biotic and five abiotic specimens within the transect. Take a soil/vegetation sample in a tube to bring back to the lab. This should be what you believe is the most accurate representation of your transect. Back at the lab, prepare a Hay Infusion Culture: Put 10-12 grams of the transect sample in a plastic jar with 500 mL of purified water. Add 0.1 gm of dried milk, place the lid on the jar and mix gently for 10 seconds. Remove the lid and store the jar in a safe place in the lab. Label the jar with your name and transect number.
Data and Observations
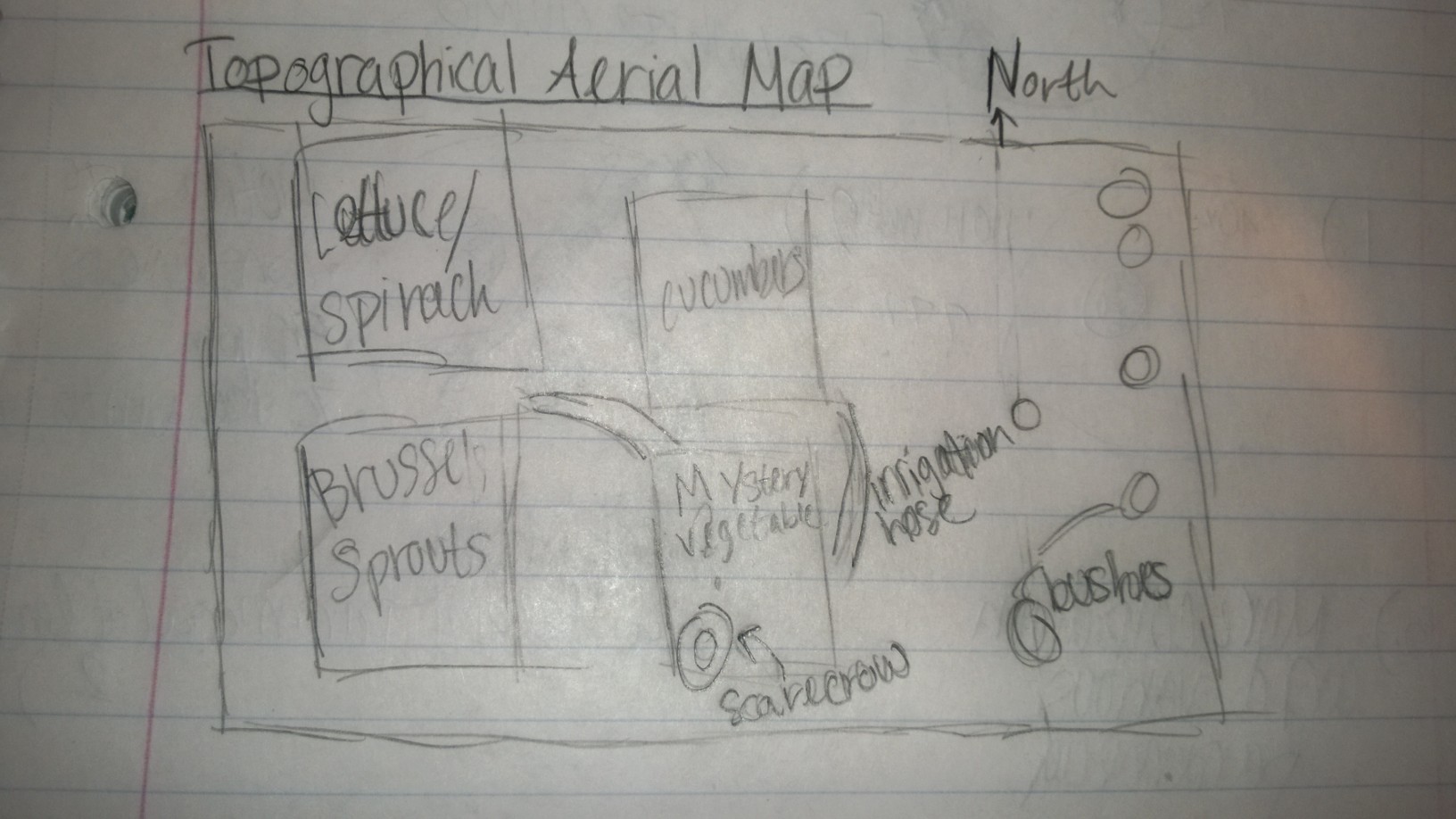
The transect assigned to me was farmland in the form of the AU community garden.
Five biotic elements: Cucumber Lettuce Brussel Sprouts Spinach Bushes
Five abiotic elements: Scarecrow Planting box Irrigation system (rubber hosing) Soil Snow
Conclusion
My transect has many plants growing there, and so I would predict the soil to be full of specimen that make it fertile. I would also expect to see organisms that feed off of plant material. The protists and bacteria I would expect to see would be ones that could survive in the varying temperatures and weather of DC, as well as survive off of the plant material (or fecal matter) that is present in the transect.
LG
January 21, 2015
Photosynthesis is the process of capturing light energy and packaging it as potential energy in the chemical bonds of glucose molecules.
LG