User:Lexi Jenkins/Notebook/Biology 210 at AU
Lab 7: 16S Sequence- 26 February 2015
Purpose:
The purpose of this lab was to learn more about bacterial genes. Since all bacteria are unique, we have to look at their individual DNA sequences to figure out what they are made up as. Our initial hypothesis for this lab was that we would find a diverse gene sequence and a sample of bacteria that is different for the samples others had.
Materials and Methods:
To begin this lab, we first had to transfer a colony of bacteria from one each of our nutrient and nutrient plus tetracycline agar plates into 100 microliters of water in a sterile tube. We used a sterile loop to do this. Then, we incubated the tubes in a heat block for 10 minutes at 100 degrees Celsius. After that, we centrifuged the tubes for 5 minutes at 13,400 rpm. While this was happening, a PCR tube filled with a mixture of 20 microliters of primer and water was prepared. When the centrifuge was done, we transferred 5 microliters of the resulting supernatant into the PCR tube with a micropipette.The next week, we stained the PCR reaction and transferred about 10 microliters into a gel electrophoresis for a little while.
Data and Observations:
The primer was specifically designed to isolate the 16S Sequence from our bacteria in order to observe that one sequence. Here is what the gel electrophoresis looked like: 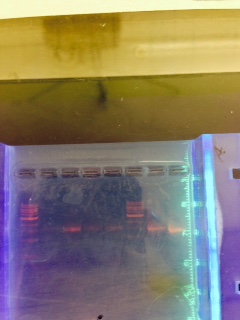 . Of the four tubes with the samples, only one really revealed anything. So, we decided to focus more on the one that ran the most through the gel.Our first gel sequence appeared to be Chryseobacterium and our second was Flavobacterium johnsoniae. Chryseobacterium is a gram negative bacteria that thrives in cold environments. Flavobacterium johnsoniae is also a gram negative bacteria commonly found in soil and freshwater.
. Of the four tubes with the samples, only one really revealed anything. So, we decided to focus more on the one that ran the most through the gel.Our first gel sequence appeared to be Chryseobacterium and our second was Flavobacterium johnsoniae. Chryseobacterium is a gram negative bacteria that thrives in cold environments. Flavobacterium johnsoniae is also a gram negative bacteria commonly found in soil and freshwater.
Conclusions and Future Directions: The 16S sequence gels yielded many different results. Our two appeared to be two different bacteria and some of the other sequences also varied. This supports our hypothesis that we would find a diverse gene sequence and have samples that varied from others in the class.
LJ
Lab 6: Zebrafish Experiment- 19 February 2015
Purpose:
This past week, we began experiments on zebrafish to see how they respond to different substances during development. These organisms are perfect for these types of experiment because zebrafish eggs are transparent, which allows us to study their embryo development. My group is using ethanol, an alcohol. We came up with the hypothesis that the control group will develop normally while the exposed group will die or develop abnormally (have varying eye spacing, delayed development).
Materials and Methods:
To conduct this experiment, we had to set up two petri dishes with 20 milliliters of water for the control group and 20 milliliters of alcohol for the test group. We labeled each petri dish and used a micropipette to measure out 20 milliliters of the substances. After that, we used pipettes to transfer approximately 20 zebrafish embryos from containers into our petri dishes. We will need to check the petri dishes every several days to document their development. Also, we will have to try to preserve a few of the fish to study later in the experiment.
Data and Observations:
After we started the experiment on Thursday the 19th, we had to come in and check the zebrafish on Friday. On the Thursday, the majority of the zebrafish were in the 20 somite stage, with one hatched fish in the water dish. In the alcohol dish, the embryos were almost split 50:50 between being in the 4 somite and 20 somite stages of development. On Friday, the water dish contained 7 dead eggs, 20 living eggs, 2 living hatchlings, and 3 dead hatchlings. Most of the embryos were in the 20 somite stage and more had hatched from the day before. In the alcohol petri dish, there were 3 dead eggs, 27 living eggs, and no living or dead hatchlings. Most of the embryos were around the 25 somite stage. We checked the zebrafish on Friday the 27th and found some pretty interesting results. Of the water dish, there was only about 1 dead egg, no living eggs or living hatchlings, and 2 dead hatchlings. Of the alcohol dish, there were about 17 dead eggs, 2 dead hatchlings, and no living eggs or living hatchlings. Though there weren't any living zebrafish, there were a bunch of motile bug-like creatures floating in both of the petri dishes. This is what the bugs looked like:  Luckily, we were able to preserve a few of our fish before they died. Of the ones preserved, they all seemed to have some sort of developmental abnormality about them. Their gills and eyes were unordinary shapes and sizes.
Luckily, we were able to preserve a few of our fish before they died. Of the ones preserved, they all seemed to have some sort of developmental abnormality about them. Their gills and eyes were unordinary shapes and sizes.
Conclusions and Future Directions:
We are able to draw the conclusion that the alcohol effected the fish in negative ways. Therefore, our initial hypothesis was supported by the experiment. LJ
Lab 5 Part II: Observing Vertebrates- 12 February 2015
Purpose:
This past week, after we observed invertebrates in our group niches, we observed visible vertebrates. The purpose of this is to complete an observation of the life within niches.
Materials and Methods:
To complete this part of the lab, we just had to make observations of the life within our niche. Because our niche is the school farmland, it is enclosed so no large animals will eat the plants. Our niche is on the other side of campus by the tennis courts and track. When we made our observations, it was approximately 9:30 in the morning during the winter. So, it was cold and the sun had been up for awhile. No one was around this niche and no cars were driving around.
Data and Observations:
Overall, we did not see a lot of evidence of vertebrates at that time. There was a bird that appeared to be a robin and a squirrel was just outside the niche. There also seemed to be a cardinal on a tree not too far from the niche. Since we did not see very many signs of life, there may be groundhogs and moles around. The robin, or Turdus migratorius, is classified as Chordata Aves Passeriformes Turdidae Turdus Turdus migratorius, and would benefit from earthworms in the niche. The Eastern grey squirrel, or Chordata Mammalia Rodentia Sciuridae Sciurius Sciurius carolinensis, would benefit from any nuts that fall from the nearby trees. The groundhog, or Chordata Mammalia Rodentia Sciuridae Marmota Marmota monax, are a type of rodent that might benefit from other rodents in the area. The cardinal, or Chordata Aves Passeriformes Cardinalidae Cardinalis cardinalis, would also benefit from the worms around the niche. Finally, the mole, or Chordata Mammalia Eulipotyphla Talpidae Scalopus aquaticus, is a creature that would benefit from almost anything living in the niche. It eats bugs, worms, and many other small living organisms. This means that these organisms make up the total population of the community (the niche) and contribute to the total carrying capacity of the niche. The squirrel, groundhog, and mole are all considered primary consumers because they eat primary producers. The robin, cardinal, and mole can also be considered secondary consumers because they consume the organisms that eat primary producers 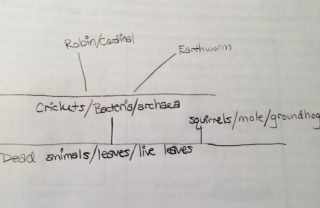 .
.
Conclusions and Future Directions:
Overall, the organisms, vertebrates and invertebrates, in this niche all provide their own elements to the area. LJ
Lab 5: Observing Invertebrates - 12 February 2015
Purpose:
This past week, we began our investigation into the world of invertebrates. From our Berlese Funnel sets we made the week before, we observed the organisms that fell into the ethanol/water mixture to see how these simple systems became more complex. Our initial hypothesis for this lab was that our Berlese Funnel sets would contain several different types of soil invertebrates.
Materials and Methods:
To start this lab, we observed prepared slides of Planaria and other coelomates. Then, we observed examples of arthropods to prepare to observe the specimen from our Berlese Funnels. After that, we broke down our funnels by tearing off the tape and only taking the 50 milliliter conical tube with the ethanol/water mixture to pour into petri dishes. We poured the first half of the mixture into one petri dish then poured the remaining part with the majority of the specimens in a second petri dish. When this was done, we first looked at them under dissecting microscopes before using concave wet mount slides to observe several drops the mixture from each petri dish with a compound microscope.Finally, we had to consider what types of vertebrates are found in our niches.
Data and Observations:
The Planaria we observed were roundworms and earthworms. Roundworms move by using long muscles in their bodies to move inside a host, since they cannot crawl  . Earthworms, on the other hand, can "crawl" by using their round, segmented bodies to extend forward and move forward
. Earthworms, on the other hand, can "crawl" by using their round, segmented bodies to extend forward and move forward  . We ended up observing different organisms (Earthworm, Fly, Springtail X (Primitive), Ground Spider LL) and finding a variety of differences between them. All of these ranged from approximately 40 millimeters long (Fly)
. We ended up observing different organisms (Earthworm, Fly, Springtail X (Primitive), Ground Spider LL) and finding a variety of differences between them. All of these ranged from approximately 40 millimeters long (Fly) 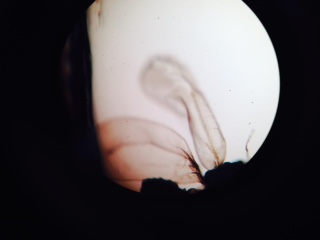 to 100 millimeters long (Springtail X (Primitive)). The most common kinds of organisms in leaf litter are probably spiders and earthworms.Here is the table of invertebrates we filled up during lab:
to 100 millimeters long (Springtail X (Primitive)). The most common kinds of organisms in leaf litter are probably spiders and earthworms.Here is the table of invertebrates we filled up during lab: 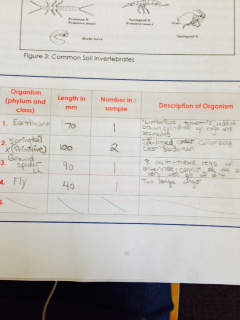
Conclusions and Future Directions:
Overall, there was a diverse assortment of invertebrates present in our Berlese Funnel set. The evidence we collected supports our hypothesis that our funnels would contain several different types of soil invertebrates. LJ
Lab 4: Plants and Fungus- 5 February 2015
Purpose:
This past week, we finally started to investigate plants and fungi. The objective of the lab was to learn about how different plant and fungi differed and the roles they play in the environment. Before we began the lab, we came up with the hypothesis that the specimen we collected would have different specialized structures and vascularizations that help them function.
Materials and Methods:
To start this lab, we first collected plant samples from our farmland transect 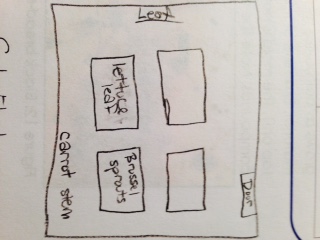 .At the same time, we started a gel electrophoresis of our PCRs from the last week
.At the same time, we started a gel electrophoresis of our PCRs from the last week  . Then, we had to identify them by their features of vascularization, specialized structures, and mechanisms of reproduction using compound microscopes and our eyesight. We also had to take note of their lengths, shape, and identity. After that, we observed a fungus sample and tried to identify which one of the three groups of fungi it belonged to. To end class, we set up Berlese Funnels to collect invertebrates from our samples. To do this, we had to first pour 25 milliliters of a solution of 50:50 ethanol and water into a 50 milliliter conical tube. Then, we fit a piece of screening material to the inside of a funnel to prevent the niche samples from falling into the ethanol. After that, we put the niche samples in the top of the funnel and parafilmed the base of the funnel and tube together to prevent the ethanol from evaporating. Finally, set the funnel on a stand under a 40 watt lamp for the next week.
. Then, we had to identify them by their features of vascularization, specialized structures, and mechanisms of reproduction using compound microscopes and our eyesight. We also had to take note of their lengths, shape, and identity. After that, we observed a fungus sample and tried to identify which one of the three groups of fungi it belonged to. To end class, we set up Berlese Funnels to collect invertebrates from our samples. To do this, we had to first pour 25 milliliters of a solution of 50:50 ethanol and water into a 50 milliliter conical tube. Then, we fit a piece of screening material to the inside of a funnel to prevent the niche samples from falling into the ethanol. After that, we put the niche samples in the top of the funnel and parafilmed the base of the funnel and tube together to prevent the ethanol from evaporating. Finally, set the funnel on a stand under a 40 watt lamp for the next week.
Data and Observations
Overall, the plant and fungus samples we chose to observe varied greatly. We observed a fallen tree leaf, a brussel sprout leaf, a lettuce leaf, a carrot stem, and fungus from one of our tetracycline agar plates. The lengths of them varied from just 2.75 centimeters to 14.5 centimeters. While three of the samples reproduced by means of seeds planted in the ground, the carrot grows flowers that produce seeds. Also, the majority of our samples proved to be dicots (tree leaf, brussel sprout leaf, and lettuce leaf) 


 . We discovered that our fungus is a zygomycota
. We discovered that our fungus is a zygomycota 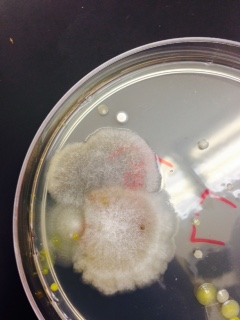 . Fungi sporangia are globelike structures of hyphae spores that turn black. They are important because they produce spores to reproduce
. Fungi sporangia are globelike structures of hyphae spores that turn black. They are important because they produce spores to reproduce 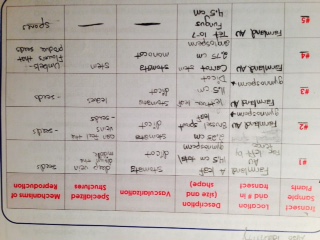 .
.
Conclusions and Future Directions:
In the end, the results of this lab did support our hypothesis that the specimens we would observe would contain various specialized structures and vascularizations. Next time, I might want to observe more species of tree leaves to compare their structures and functions. LJ
Lab 3:Microbiology: DNA and Bacteria- 29 January 2015
Purpose:
This past week, we observed bacteria in agar plates to learn about antibiotic resistance and differences between bacteria. We also started observing how bacteria can be gram positive or gram negative. We came up with the hypothesis that there will be several species of bacteria present in the agar plates.
Materials and Methods:
To start the lab, we observed our Hay Infusion Cultures and paid attention to any changes in smell and appearance. Then, we collected our eight agar plates and counted the number of bacterial colonies present on them. After that, we scraped small amounts of growth from four of the plates with sterilized loops and made wet mounts to observe the bacteria under compound microscopes. At the same time, we made four gram stains by using the sterilized loops and mixing the growth with a drop of water. We used bunsen burners to dry the samples to the slides before soaking them in crystal violet, iodine, decolorizer, and safranin for certain amounts of time over a staining tray. Between each soaking, we rinsed the chemicals off with water. After that, we blotted the slides dry with kimwipes then observed them under compound microscopes. Finally, we made PCRs to isolate DNA in the bacteria. To do this, we had to 1)transfer a colony of bacteria into 100 microliters of water in a sterile tube, 2)incubate the tube in 100 degree Celsius water for 10 minutes, 3)centrifuge the tube for 5 minutes at 13,400 rpm, and 4)transfer 5 microliters of the supernatant from the centrifuged tube into the 16S PCR reaction tubes that were prepared while the sample centrifuged. The tube contained a mixture of 20 microliters of primer and water.
Data and Observations:
From our final observation of the Hay Infusion Culture, we learned that there was about half the amount of water as the day we made the cultures. Also, the color of the water was about the same but the smell was less pungent than the week before. Here is our reult table for our serial dilutions: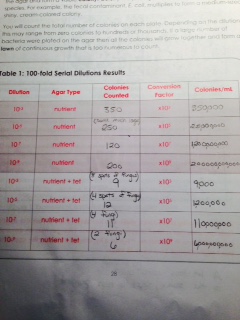 When we observed the agar plates, we realized that the plates without the tetracycline had more colonies than those with it. This means that some of the bacteria was not resistant to the tetracycline, so they were unable to grow in the agar plates. We also noticed that the agar plates with the tetracycline made more fungus grow than the plates without the tetracycline. When we observed the gram stains, only one of the slides appeared to be gram positive, while the other three were negative. Each slide yielded different bacteria. One appeared to be Filamentous and the others were Fusobacterium, Vibrio, and Coccus. None of the samples were motile. Here is our table of our findings:
When we observed the agar plates, we realized that the plates without the tetracycline had more colonies than those with it. This means that some of the bacteria was not resistant to the tetracycline, so they were unable to grow in the agar plates. We also noticed that the agar plates with the tetracycline made more fungus grow than the plates without the tetracycline. When we observed the gram stains, only one of the slides appeared to be gram positive, while the other three were negative. Each slide yielded different bacteria. One appeared to be Filamentous and the others were Fusobacterium, Vibrio, and Coccus. None of the samples were motile. Here is our table of our findings: 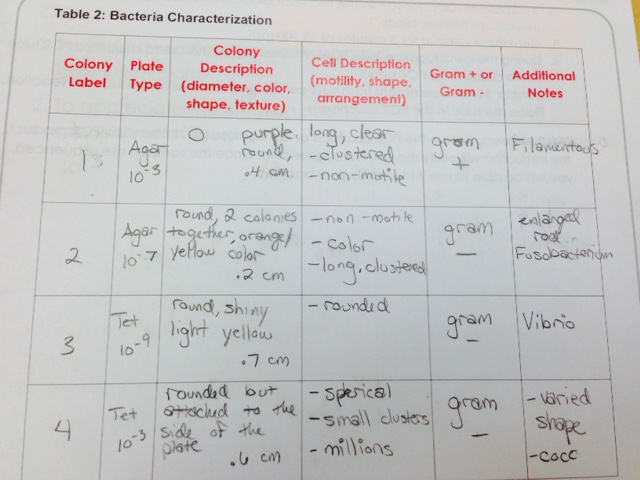
Conclusions and Future Directions:
Overall, there were species of bacteria discovered in the agar plates. This discovery supports our initial hypothesis that there would be several species of bacteria present in the agar plates.
LJ
Lab 2:Studying Algae and Protists-22 January 2015
Purpose: This past week, we learned how to identify different organisms based on a number of physical aspects. We also learned that dichotomous keys are important ways of identifying the nature of many organisms from different environments. We came up with the hypothesis that the different niches we collected samples from will contain different types of bacteria and biotic organisms.
Materials and Methods: For this lab, we continued to use the Hay Infusion Cultures we made last week. These were made by first placing 10-12 grams of the soil we collected from our niche into a plastic jar with 500 millilters of deerpark water. Then, we measured .1 gram of dried milk on a scale and mixed it into the jar by shaking it for 10 seconds. After that, we just had to remove the lid and set the labeled jar in a safe area of the lab for the week. We observed the infusions in the jar then made two wet mounts (from the base of the jar and the surface) to observe them under compound microscopes. After that, we used dichotomous keys to identify as many organisms in the samples as possible. When this was done, we went on to the next part of the lab where we made dilutions of our cultures to observe what kinds of bacteria will grow in them over the next week. To do this, we labeled four 10 milliliter tubes of sterile broth. Then, we labeled four plates each of nutrient agar and nutrient agar plus tetracycline. We added 100 micrometers of the culture to the first 10 milliliter tube of broth using micropippetors, then took 100 micrometers of this dilution and added it to the next tube. This continued for all of the tubes, causing the final tube to be the most diluted. Finally, we plated the dilutions by taking 100 micrometers of each dilution in pipettes and spreading them.
Data and Observations: The Hay Infusion Culture we made was a murky brown color with dirt settling at the bottom of the jar half full of water. It did not smell too bad, but we could detect the smell of some of the dead grass in it. We ended up identifying arcella (pale, non-motile algae), actinosphaerium (clear, spherical, non-motile algae), stentor (cone-shaped, non-motile), and gonium (greenish, many celled) from the sample collected at the base of the culture container. At the surface of the culture, there were not many organisms, but we were able to identify euglena (motile, elongated, contains flagella). This could mean that the condition It was interesting to find several algae that were completely different from each other because all of the samples we collected from out niche were from the same general area and soil.
Conclusions and Future Directions: After completing this lab, we can conclude that within each niche, there is a plethora of organisms and algae that can be found. This supports our hypothesis we made at the beginning of the lab. Next time, I might compare the results we found to those of other groups to further examine the hypothesis.
LJ
Lab 1:The Diversity of Life- 15 January 2015
Purpose: This week, we studied the concepts of natural selection and evolution. In order to show how certain traits in a population have changed over time, we needed to collect and study samples of biotic items found in nature. We came up with the hypothesis that the specimens found in different parts of the campus will contain different types of bacteria.
Materials and Methods: For this lab, we observed different types of Volvocine Line algae (Chlamydomonas, Gonium, and Volvox) under compound microscopes. We used droppers to drop the algae samples onto microscope slides and we used micrometers to measure the specimen. We used 50 milliliter sterile conical tubes to collect samples outside in specific niches. Then, we added 10-12 grams of the specimens to plastic jars and added 500 milliliters of water and .1 grams of dried milk to make Hay Infusion Cultures.
Data and Observations:
When we observed the Volvocine Line algae, we were able to find 250 Chlamydomonas, 3 Gonium, and 2 Volvox. Only the Volvox has a specialization (spiked) and only the Chlamydomonas had a mechanism of motility (flagella). The Chlamydomonas was isogamous while the other two were oogamous. The Chlamydomonas were small, egg-shaped organisms, the Gonium were spherical, and the Volvox were spiked. In the niche (the school garden), we collected brussel sprout leaves and soil samples from several areas. There were a lot of biotic factors in that area. Five biotic factors in the niche were broccoli, brussel sprouts, grass, lettuce leaves, and the bushes. Five abiotic factors were ice, the planter boxes, rocks, the fence, and fallen leaves. Here is a map of our transect: 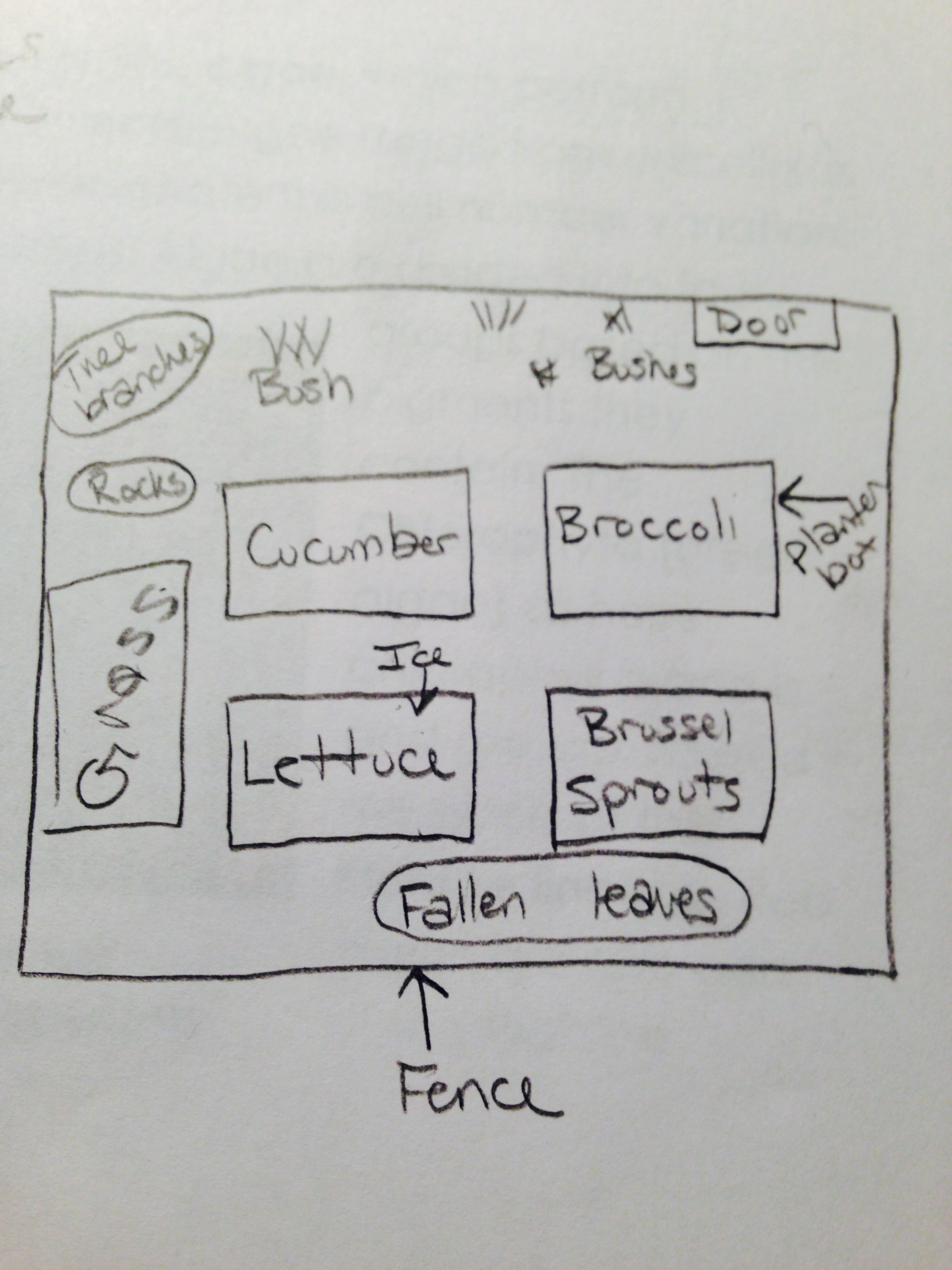
Conclusions and Future Directions: The specimens collected in each niche all seemed to include different materials. This can mean that there are different bacteria and living things in each area, which could confirm our hypothesis when we observe the Hay Infusion Cultures.
LJ
16 January 2016*Lexi Jenkins 16:35, 16 January 2015 (EST):
I am excited to work on all of the Biology 210 labs!
LJ