User:Kirk Blackmoore/Notebook/Biology 210 at AU
01/23/14
01/16/14
Embryology & Zebrafish Development
Procedure 1: Day 1
1) The control group, as well as, the test group (salt) were set up.
2) 20 mls of Deerpark water and 20 mls of healthy translucent embryos were added to the control petri dish.
3) 20 mls of healthy translucent embryos and three varying concentrations of salt solution were added to the respective petri dishes.
4) The dishes were labeled "control" "3% NaCl", "0.3% NaCl", "0.03% NaCl"
5) Zebrafish embryos were carefully observed and the current developmental stage was determined.
- 18 hours post-fertilization
- just entering organogenesis
- eyes & tail starting to come off yoke

Procedure 2: Day 4/5
1) The petri dishes were observed for surviving embryos.
2) The dead embryos were discarded and the solutions (control or drugged) were replenished.
3) The surviving embryos were counted and observed under a disecting microscope.
| !- | Unhatched/Dead | Unhatched/Alive | Hatched/Dead | Hatched/Alive |
|---|---|---|---|---|
| Control | 4 | 0 | 5 | 38 |
| 0.03% | 2 | 0 | 4 | 20-21 |
| 0.3% | 1 | 0 | 1-2 | 28-30 |
| 3% | 1 | 0 | 25 | 0 |
Procedure 3: Day 7
1) The same as day 4/5 was done including the heart rate.
2) 1 drop of paramecium was added
| - | 0.03% Concentration | 0.3% Concentration | 3% Concentration | Control |
|---|---|---|---|---|
| Heart Rate | 78/min | 114/min | 0 | 54/min |
| Unhatched/Dead | 0 | 0 | 0 | 0 |
| Unhatched/Alive | 0 | 0 | 0 | 0 |
| Hatched/Dead | 0 | 8 | 23 | 0 |
| Hatched/Alive | 27-30 | 18 | 0 | 30-40 |
- the were active and swam often
- a few fish were preserved in
Procedure 4: Day 11
1) Similar process was continued.
- all zebra fish were found to be dead so the preserved ones were observed
| Size of Eyes (μm) | Size of body (μm) | Solution |
|---|---|---|
| 28 | 400 | Control |
| 28 | 338 | 0.03% |
| 30 | 400 | 0.3% |
| 28 | 180 | 3% |
-Kirk Blackmoore 3/10/14
Invertebrates
Procedure I: Observing Acoelomates, Pseudocoelomates, and Coelomates
- Three different types of worms were observed
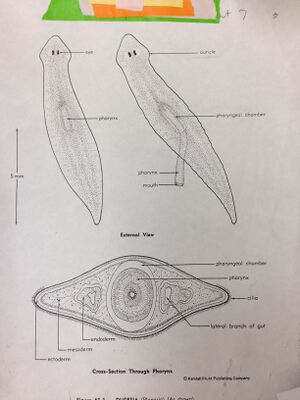 Figure 1. Diagram of Flatworm observed
Figure 1. Diagram of Flatworm observed
Procedure II: Analyzing the Invertebrates Collected with the Berlese Funnel
- The invertebrates found in the Berlese Funnel prepared from the previous lab were observed under dissecting microscope.
- Five were found and able to be identified.
 Figure 3. Naked eye view of invertebrates found
Figure 3. Naked eye view of invertebrates found
| Kind of Organism | Length in mm | Brief description of Organism |
|---|---|---|
| 1) Ant (closest) | <1 | 6 legs, what appears to be a tail, 1 pair antennae, 2 body segments |
| 2) Beetle | ~2 | 1 pair antennae, 2 body segments, wings on back, dark brown |
| 3) Fly (Diptera) | <1 | black, 1 pair of wings, small |
| 4) Spider | <1 | 8 legs, held food in legs, whitish |
| 5) Centipede | 10 | long, relatively balled up, many legs |
Procedure III: Vertebrates and Niches
Possible Vertebrates which may interact with transect:
| Vertebrate | Phylum | Class | Order | Family | Genus | Species | Benefit |
|---|---|---|---|---|---|---|---|
| 1) Brown Rat | Chordata | Mammalia | Rodentia | Muridae | Rattus | R. norvegicus | protection from rosebush, feed on |
| 2) Dog | Chordata | Mammalia | Carnivora | Canidae | Canis | C. lupus | area to excrete waste |
| 3) Eastern Gray Squirrel | Chordata | Mammalia | Rodentia | Sciuridae | Sciurus | S. carolinensis | feed on the acorns |
| 4) Rock Pigeon | Chordata | Aves | Columbiformes | Columbidae | Columba | C. livia | feed from the seeds produced by flowers |
| 5) Common Starling | Chordata | Aves | Passeriformes | Sturnidae | Sturnus | S. vulgaris | feed on seeds produced by flowers |
-Kirk Blackmoore 2/26/14
Plantae and Fungi
Procedure I: Collecting five plant samples from the transect
- Three bags were taken to the transect
- A leaf litter sample was collected, followed by some soil and added to one bag. This collection was used to set up the Berlese funnel for collecting invertebrates.
- Five different plant samples were then collected. Additionally, old leaves and branches were collected as well as other plant matter.
 Figure 1. Landscape view of Transect
Figure 1. Landscape view of Transect
Procedure II: Plant Vascularization
- The cross section slide of the lily stem was observed and the height of the plant stem was measured and found to be 24 cm
 Figure 3. Cross Section of Lily
Figure 3. Cross Section of Lily
Procedure III: Plant Specialization
- The leaves of the moss were observed
 Figure 4. Dicot vasularization under microscope
Figure 4. Dicot vasularization under microscope
Procedure IV: Plant Reproduction
| Transect: | Location & Number in Transect | Description | Vascularization | Leaves & Special Characteristics | Seeds, evidence of flowers or other reproductive parts |
|---|---|---|---|---|---|
| #1 group: Angiosperm genus: Rosa | beneath rose bush in soil | green leaf | xylem & phloem | simple leave with vascularization, oval with pointy edge. | Dicot based on xylem and phloem. |
| #2 group: Angiosperm genus: Quercus | in grass | leaf with shape of black oak leaf | xylem & phloem | 6 lobes with spines | dicot. Acorn was found suggesting that it was black oak. |
| #3 group: Angiosperm genus: Quercus | in grass. | small, brown acorn. | no vascularization | Suggests the presence of black oak based on leaf evidence. | Suggests plant is dicot. |
| #4 group: Angiosperm genus: Rosa | beneath rosebush in soil. | flower which resembled those on the rosebush. | none visible. | same as previous leaf from rosebush | Its 5 flowers shows plant is dicot. |
| #5 group: Angiosperm genus: Agrostis | removed from grass. | short, thin grass | short veins throughout the blade. | simple leaves, approx 15 cm in length. | Monocot based on vascularization. |
Procedure V: Observing Fungi
- Black bread mold was observed under a dissecting microscope.

Fungi Sporangia is important because it allows fungi to reproduce asexually, simply, by producing spores (http://website.nbm-mnb.ca/mycologywebpages/NaturalHistoryOfFungi/AsexualSporangia.html)
Procedure VI: Setting up the Berlese Funnel to Collect Invertebrates
- About 25 mL of 50:50 ethanol and water solution was added to a tube.
- A piece of filter was placed at the bottom of the funnel. A light was placed about 2 inches from the top of the leaf litter added to the funnel. It was all covered with foil.
- This was set up and left for a week.
-Kirk Blackmoore 2/26/14
Microbiology and Identifying Bacteria with DNA
Procedure I: Quantifying and Observing Microorganisms
- The total number of colonies which developed on each of the plates which were prepared the week before was counted and added to Table 1.
| Dilution (Plate label) | Agar | Colonies Counted | Conversion Factor | Colonies/ml |
|---|---|---|---|---|
| 10E-3 | nutrient | 100+ | x10E3 | |
| 10E-5 | nutrient | 50+ | x10E5 | |
| 10E-7 | nutrient | 12 | x10E7 | |
| 10E-9 | nutrient | 3 | x10E9 | |
| 10E-3 | nutrient + tet | 53 | x10E3 | |
| 10E-5 | nutrient + tet | 0 | x10E5 | |
| 10E-7 | nutrient + tet | 1 | x10E7 | |
| 10E-9 | nutrient + tet | 0 | x10E9 |
Procedure II: Antibiotic Resistance
- After observing the plates, the following questions were answered:
- 1)Do you see any differences in the colony types between the plates with vs without antibiotic? What does this indicate? Yes. It shows how the antibiotic did its job in reducing bacterial growth.
- 2)What is the effect of tetracycline on the total number of bacteria? Fungi? Tetracycline reduces the growth of bacteria and fungi.
- 3)How many species of bacteria are unaffected by tetracycline? three
How Tetracycline Works
Tetracycline disrupts many of the essential enzyme reactions for the vital processes of bacteria. Tetracycline binds specifically to the 30S ribosome of bacteria hindering the attachment of the aminoacyl tRNA to the RNA-ribosome complex. At the same time, it prevents the other steps of the protein biosynthesis. (http://www.chm.bris.ac.uk/motm/tetracycline/antimicr.htm)
Procedure III: Bacteria Cell Morphology Observations
- Prokaryotes were observed through a microscope.
1)A prepared slide containing different types and shapes of bacteria was obtained. These were then observed with 40 X objective. When they were located, they were then observed through oil immersion (100 x).
2)A wet mount was made of a tiny amount of bacterial growth from a plate added to a drop of water on a slide; a cover slip was added to it before being observed under microscope. A second slide was made. This one was allowed to air dry so that it could be stained and observed under the microscope.
3)The organisms were observed under microscope at 10x and 40x.
4)The stained slide was prepared in the following steps:
- It was labeled
- The smeared slide was heat fixed
- The smear was covered with crystal violet for one minute
- The smear was then covered with Gram's iodine mordant for 1 minute then gently rinsed
- The smear was then decolorized by running 95% alcohol on it for 10-20 seconds. It was then rinsed until if flowed colorlessly from the slide
- The smear was covored with safranin stain for 20-30 seconds then gently rinsed
- Excess water was blotted out.
- The sample was focused on at a low magnification then observed at 40x after the gram stain was done
Procedure IV: Start PCR Preparation for DNA Sequence Identification
- Bacteria was characterized from Hay infusion, some resistant to tetracycline and some that are not. The DNA for each bacteria was isolated in the colonies and the 16S rRNA gene was amplified using two primer sequences (27F and 519R).
- 1) a single colony of bacteria was transferred to 100 µL of water in a sterile tube. It was then incubated at 100°C for 10 minutes then centrifuged. 5µL of the supernatant was used in the PCR reaction.
Hay Infusion:
- darker but still slightly transparent
- some more of it was evaporated
- now has a distinct smell (more concentrated?)
-Kirk Blackmoore 2/26/14
Identifying Algae and Protists January 23, 2014
Procedure 1: How to Use a Dichotomous Key
1) a wet mount was made of the sample with known organisms and was observed with the microscope in 4X and 10X lens. Many of the organisms were motile and difficult to be seen.
2) One organism was focused on and characterized. The size was determined with the ocular micrometer.
3)A key that describes eight known organisms was used to determine the organism. The diagram was used to confirm.
4) The process was repeated with a second organism.
Procedure 2: Hay infusion Culture Observations
1) The jar was carefully taken to the workstation and the smell, and appearance were observed:
~It appeared to be brown and moldy but transparent. The smell was not strong enough to really be determined; there was no smell.
2) A few samples were taken for microscopic observation. Organisms from two different niches were observed.
3) Pictures of the observed organisms were drawn.

- Most likely Paramecium Aurelia. Observed at 10x magnification. Mobile-Very quick. Has cilia. Approximately 100 micrometers.
- Does not photosynthesize-protist.
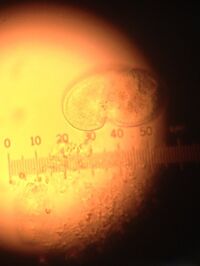
- Most likely Bursaria Trunctella. Approximately 2000 micrometers. Has a large vacuole an is covered in cilia. Mobile.
- Does not photosynthesize-protist

- This organism was unidentified but closest to Dindinium. Observed at 40x magnification. Has cilia. Mobile.
- Does not photosynthesize-possibly protozoa
- There was not much variation observed in the two niches. The organisms expressed above were the only ones which were observed in the both niches.
- If this hay infusion is to be observed for another two months, I expect there to be many more of the same organisms present. There may be less variation than what already exists based on which organisms are able to thrive in the environment created. However, there will be an increase in whatever organism survives.
- Due to the fact that the transect from which the hay infusion was created is not very large, it was expected that there wouldn't have been a whole lot of variation in the organisms found. This can be attributed to ecological selection.
Procedure 3: Preparing and Plating Serial Dilutions
1) Four tubes of 10 mL of sterile broth were obtained and labeled 2,4,6,8.
2) Four nutrient agar and four agar plus tetracycline plates were found. All tetracycline plates were labeled with "tet". One plate from each of the two groups were labeled 10^-3, 10^-5, 10^-7, and 10^-9.
3) The Hay Infusion culture was swirled and the organisms were mixed up. 100 microliters from the mix were added to the 10 mL of the broth in the tube labeled 2 for 10^-2 dilution or a 1:100 dilution. The inoculated tube was swirled well.
4) 100 microliters of broth from tube 2 was used to inoculate tube 4 (10^-4 dilution of 1:10,000). It was swirled once again to mix well.
5) The above was repeated two more times to make 10^-6 and 10^-8 solutions.
6) 100 microliters from the 10^-2 tube were taken and aeseptically placed on the surface of nutrient agar plate labeled 10^-3. The sample was carefully spread on the plate. The exact procedure was repeated on the tet plate labeled 10^-3.
7) The exact procedure with the number 4 tube on the 10^-5 plates, again with number 6 on the 10^-7 plates and, also, the number 8 on the 10^-9 plates.
8) The agar plates were incubated at room temperature over the next week.
-Kirk Blackmoore 02/09/14 2:50 pm
Biological Life at AU January 16th, 2014
Data for the memebers of the Volvocine Line
Procedure 1: The Volvocine Line
1) A slide of Chlamydomas was prepared.
2) The slide was then observed for various characteristics and living culture. Attention was placed on how the previous organism differed from the following one. (in terms of complexity etc.)
3) The process was then repeated for Gonium, then Volvox.
Number of Cells ~Chlamydomonas: 1 ~Gonium: 1 ~Volvox: Many
Colony Size ~Chlamydomonas: >50 ~Gonium: 4-14 ~Volvox: Thousands
Describe any functional Specialization of cells ~Chlamydomonas: Asexual Reproduction/egg-shaped cells/flagella ~Gonium: Held together in gelatinous mix ~Volvox: Spherical colony
Describe any reproductive specialization (isogamy vs oogamy) ~Chlamydomonas: Isogamy ~Gonium: Oogamy ~Volvox: Oogamy
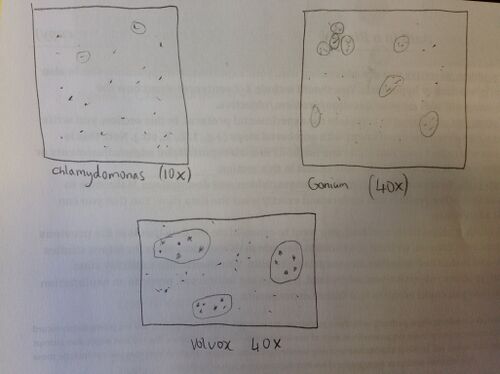 Figure 1. Hand drawing of the Volvocine Line
Figure 1. Hand drawing of the Volvocine Line
It can be seen that, as these cells progressed, they became more complex. This can be seen by the increase in complexity from the Chlamydomonas to the Gonium; the same can be observed from Gonium to Volvox. Judging off how the cells increased in complexity as a result of evolution, it could be inferred that evolution moves to increased complexity. The reason for evolution is to put an organism at an advantage in its environment. As a result it only makes sense that evolution leads to more complexity. This can be seen in the example of the Volvocine Line where it can be seen that evolution lead from isogamy and asexual reproduction to oogamy and asexual, as well as, sexual reproduction.
Procedure 2: Defining a Niche at AU
1) Transect 5 was assigned to my group (Located in the main Quad)
2) It was analyzed for both biotic and abiotic matter
3) A sample of half soil and the other, plant matter was collected for further analysis
4) The collected matter was used to make a hay infusion back in the laboratory.
5) 12 grams of the soil and plant matter was added to a plastic jar along with 500 mLs of deerpark water. Additionally, 0.1 g of dried milk was placed in and it was all mixed up then placed in a safe place.

Biotic Matter: Rosebushes, grass, leaves, wood chips, moss
Abiotic Matter: Rocks, Cement bench, Slate tiles
The Transect Matter in the jar was stored away in order to be able to observe the various changes which occurred in it physically in the following labs. Additionally, wet mounds will be placed on slides to observe the microscopic organisms in future labs. -Kirk Blackmoore 01/31/14
Excellent start. Clear entry describing tasks undertaken and answering questions. Could include a little more detail. SK

