User:Jen17

Hi, my name is Jennifer Auchtung.
I was a grad student/post-doc in Alan Grossman's lab.
Back to the Grossman Lab Webpage
I'm now a post-doc in Jim Tiedje's lab at Michigan State University. Tiedje Lab
This is how to contact me:
E-mail: j.auchtung@gmail.com
Phone: (517)355-0271 ext 1+286
Address:
Jennifer Auchtung
Center for Microbial Ecology
540b Plant & Soil Sciences Building
Michigan State University
East Lansing, MI 48824
Education
Ph.D. in Biology, January 2006
Massachusetts Institute of Technology
B.S. in Microbiology, May 2000, cum laude
Michigan State University
Research
Horizontal gene transfer plays an important role in bacterial evolution. Acquisition of foreign DNA can be beneficial to a cell, such as when the DNA encodes proteins that allow the cell to adapt to a new niche. However, acquisition of foreign DNA can also be detrimental, such as when the DNA encodes a toxic product. Therefore, mechanisms that regulate horizontal gene transfer can balance the potential risks of acquiring foreign DNA with the potential rewards. My work has focused on mechanisms that regulate two types of horizontal gene transfer, competence for genetic transformation and transfer of the mobile genetic element ICEBs1, in the Gram-positive soil bacterium Bacillus subtilis.
Regulation of competence development
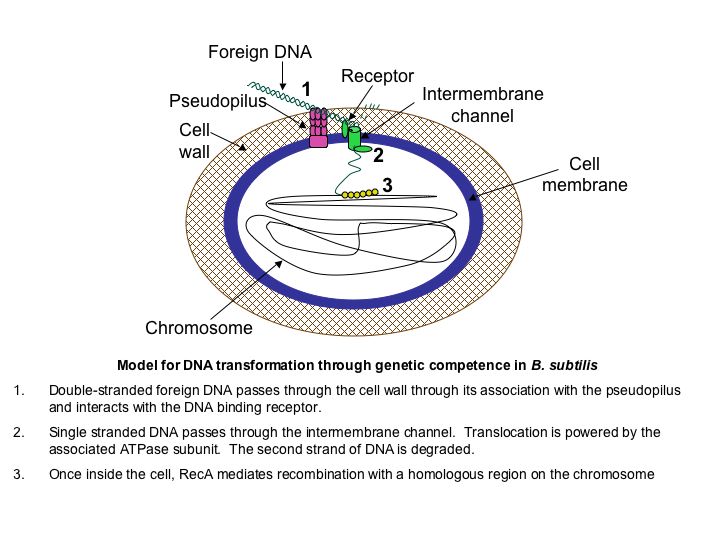
Competence for genetic transformation is the ability of cells to take up naked DNA from the environment and incorporate this DNA into the chromosome through homologous recombination. Here is a simplified diagram of competence in B. subtilis:
In addition to expressing the proteins required for DNA uptake and processing, B. subtilis competent cells are in a transiently differentiated state - cell division arrests and the cells’ metabolism likely changes. Because these changes may be disadvantageous for the cells under certain conditions, competence development is highly regulated.
One mechanism that regulates competence development in B. subtilis is quorum sensing. Quorum sensing is a mechanism by which cells use small, intercellular signaling molecules to monitor the concentration of other related cells. B. subtilis cells become competent when they sense they are surrounded by a threshold concentration of related cells. This regulation is likely beneficial to the cells because it limits competence development to conditions when DNA from closely related cells is more likely to be present. B. subtilis cells evaluate the concentration of related cells present by producing and responding to small signaling peptides. Previous work had shown that two signaling peptides regulate competence development. My work identified two additional signaling peptides that regulate competence. All four signaling peptides interact with different cellular targets and the expression of the signaling peptides as well as their target proteins are regulated by a variety of different transcription factors which respond to different cellular signals. The involvement of multiple signaling peptides allows for the integration of many regulatory signals in the decision to become competent.
Regulation of ICEBs1 transfer
ICEBs1 is an integrative and conjugative element (also known as a conjugative transposon), a type of mobile genetic element that is normally found integrated into the chromosome of a host cell that can excise from the chromosome and transfer to a recipient cell through conjugation. Here is a simplified diagram of the life cycle of an integrative and conjugative element:
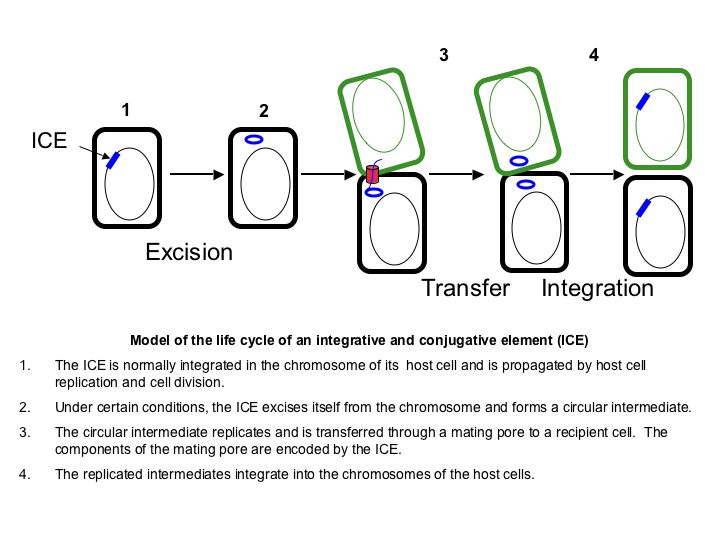
Under most conditions, the genes required for ICEBs1 excision and transfer are not expressed. This is likely due to the fact that unrepressed expression of the ICEBs1 excision and conjugation genes is detrimental to the element - it is toxic to the host cell and causes the element to be lost at a high frequency. However, under certain conditions it is beneficial for the element to express its excision and conjugation genes.
One condition that induces expression of the ICEBs1 excision and conjugation genes is the global DNA damage response. Similar to many bacteriophage, ICEBs1 uses this regulation to sense host cell genomic stress and initiate its escape from distressed cells.
Intercellular peptide signaling also regulates excision and transfer of ICEBs1. Chromosomally-encoded signaling peptides function along with starvation signals to stimulate expression of the ICEBs1 excision and conjugation genes, thereby promoting excision and mating when the concentration of potential mating partners is high. A second peptide signaling molecule encoded within ICEBs1 inhibits expression of the ICEBs1 excision and conjugation genes. This peptide functions in self-recognition, inhibiting mating when the concentration of cells that already contain ICEBs1 is high. The combined effect of regulation by intercellular signaling peptides is to limit expression of the ICEBs1 excision and conjugation genes to conditions where mating is most likely to be efficient - when there are high concentrations of potential recipient cells that lack ICEBs1.
If you interested in reading more about horizontal gene transfer in bacteria, Nature Reviews Microbiology had several good reviews on the subject in the September 2005 issue.
A more thorough description of competence is available in this review written by Ines Chen and Dave Dubnau:
Chen, I. and Dubnau, D. 2004. "DNA uptake during bacterial transformation." Nat. Rev. Microbiol. 2:241-9.
A more thorough description of integrative and conjugative elements is available in this review written by Vincent Burrus and Matt Waldor:
Burrus, V. and Waldor, M.K. 2004. "Shaping bacterial genomes with integrative and conjugative elements." Res. Microbiol. 155:376-86.
Publications
Auchtung, J.M. and Grossman, A.D. “Cell-cell signaling and the recognition of self in the control of horizontal gene transfer in Bacillus subtilis.” In Cell-cell signaling in Bacteria, Second edition, B.L. Bassler and S.C. Winans, eds. (Washington D.C., ASM Press). In Press.
Auchtung, J.M., Lee, C.A., Garrison, K.L., and Grossman, A.D. “Identification and characterization of the immunity repressor (ImmR) that controls the mobile genetic element ICEBs1 of Bacillus subtilis.” In Press.
Teaching Experience
CO-INSTRUCTOR, MIT FIELD TRIP FOR HIGH SCHOOL SCIENCE CLASSES (Annual, 2005 – 2007)
As an outreach to the community, the MIT biology department invites students and teachers from biology classes at local area high schools to spend the day at MIT and experience biological research. In March of 2005, Melanie Berkmen and I designed a short lesson that allowed the students to investigate how the bacterium Bacillus subtilis responds to starvation by forming dormant spores. The students had the chance to use light microscopy to distinguish sporulating cells from non-sporulating cells. They also learned that spore formation in Bacillus subtilis is an interesting process to study because it is an example of bacterial development and because of the similarities to spore formation in pathogenic bacteria, such as the anthrax-causing bacterium, Bacillus anthracis. I enjoyed this first experience so much, that I volunteered to teach the same lesson again in 2006 with Melanie and in 2007 with Kasia Gora, a rotation student in Alan's lab.
TEACHING ASSISTANT, MOLECULAR BIOLOGY, MIT (Spring 2004)
This senior-level undergraduate/graduate course was taught by Tania Baker and Steve Bell. During the course of the semester we covered a wide range of topics, including DNA replication and recombination, transposition, transcription, translation, and gene regulation. My responsibilities for this course involved preparing and leading weekly discussion sections with students, preparing problem sets for students in collaboration with another teaching assistant, and grading students’ exams. It was a great opportunity to watch intelligent students learn to engage challenging concepts that I find very interesting.
TEACHING ASSISTANT, EXPERIMENTAL BIOLOGY LAB, MIT (Fall 2001)
This introductory biology was taught by Angelika Amon, Alex Rich, Mary Lou Pardue, and Andrew Chess. During the course of the semester, the students had the opportunity to try a wide variety of experimental techniques to investigate questions in microbial genetics, protein biochemistry, recombinant DNA methods, and zebrafish development. My responsibilities for this course involved preparing biweekly discussion sections with students during the microbial genetics section of the course, preparing section notes for online publishing, supervising students' lab activities, and grading students’ exams and lab reports. This lab course was unlike any I had ever experienced as an undergrad because the students had the opportunity to use so many techniques. It was also interesting to watch some students learning how to do lab work for the first time and to meet others who were already quite experienced due to previous independent research experiences.
Personal
When I'm not hard at work, I enjoy reading, running, playing tennis, and spending time with my son, Julian.

My husband, Tommy Auchtung, is also a microbiologist and a graduate student in Colleen Cavanaugh's lab at Harvard University. He's studying Korarchaeota, a division of Archaea that is thought to be deeply rooted on the evolutionary tree. One perk about studying Korarchaeota is the interesting places he goes to collect samples (Yellowstone National Park, Kamchatka, Russia, the bottom of the ocean.) Unfortunately, I don't get to go along.