User:James Chappell / Rolling Assembly
From OpenWetWare
Jump to navigationJump to search
Bacillus Genome Vector
Bottom-up genome assembly using the Bacillus subtilis genome vector
This paper describes a new technique that we could try to adopt for our iGEM project. The advantages and issues are summarized in the slides below:
Principle of Rolling Assembly
Our current strategy for insertion of multiple devices into the genome of B.subtilis is to integrate these devices as different defined positions within the genome. However an alternative is to develop a rolling assembly strategy that enables combination of devices at one loci within the genome.The possible advantages for this are as follows:
- Similar in situ position reduces variability in the transcriptional level.
- Only limited no. of biobricks needed for insertion.
- Only one insertion site is needed for initial insertion of the device.
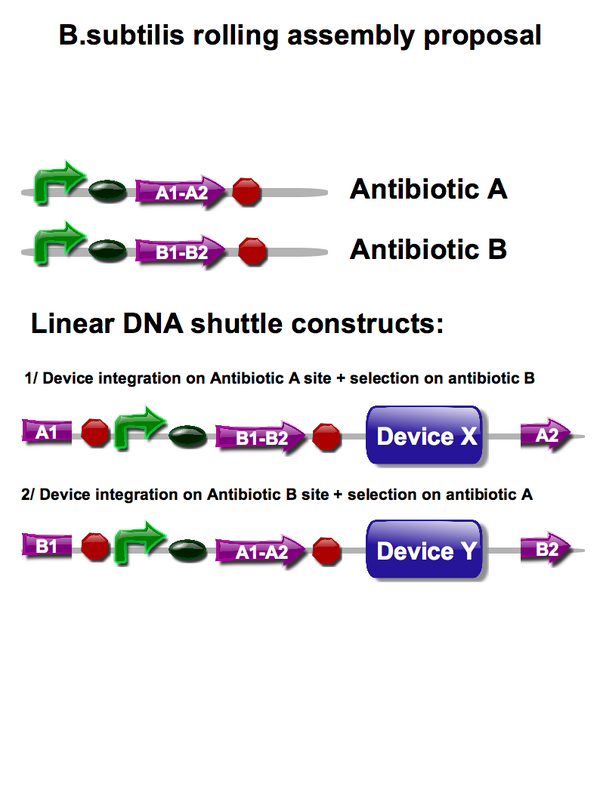
A possible solution for rolling assembly devised by adviser Vincent Rouilly is described below:
- Initial integrate into an integration site within the B.subtilis genome, with the device and a device encoding the constitutive expression of Antibiotic A. This antibiotic provides a means for selection.
- For the second integration the Antibiotic A gene is used as an integration site, so flanking regions of homology to antibiotic A are placed around the second device to allow insertion. In addition the device 2 contains a device encoding the constitutive expression of Antibiotic B. Again this antibiotic allows for selection and future integration sites.