User:James Chappell/Weak Promoter
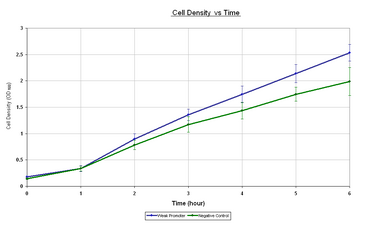 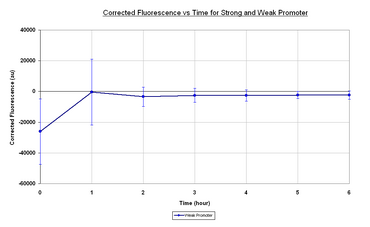 |
| Figure 1.1 it can be seen that the growth rate of the E.coli containing the I20260 is lower than that of E.coli with no plasmid. Both samples appear to have an initial lag phase for the first hour followed by the exponential phase for the remainder of the experiment. The reason for the slow growth rate of I20260 could be put down to the expression burden that I20260 is placing on the E.coli. Figure 1.2 shows the corrected fluorescence vs time. Corrected fluorescence was calculated by normalising by the OD600 and taking away the negative control to remove background fluorescence. As it can be seen by taking away the negative control we get an negative initial value that increases over the first over and reaches a steady value for the remainder of the sampling. The reason why the negative control fluorescence is higher is not known, however it is clear that throughout the sampling there is minimal GFPmut3b expression from the I20260. The steady-state reached at 1 hour could be due to inaccuracies in the initial measurement or due to a true steady-state of GFPmut3b per cell.
|