User:Federico Castro M/Adaptive Landscape
Last updated February/26/2008
Project status: In stasis
Objective: To design and assemble a devise that would allow us to map the adaptive landscape resulting from artificial genes that confer antibiotic resistance.
The theory
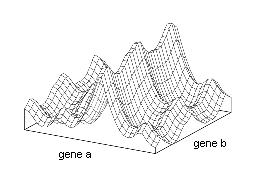
In the adaptive landscape, of S.Wright, the fitness associated with different genotypes as well as the relation between two genotypes is visualized in a map. Different evolutionary forces such as endogamy, genetic drift, horizontal transfer and natural selection move the population across the adaptive landscape.[1] An analogy to the movement of populations across adaptive landscape could be a ball falling into a cone by the action of gravity (although natural selection usually moves populations uphill).
The antibiotic resistance acquired by bacteria and drug resistance developed by virus can be seen as a movement of the population to peak of fitness in the adaptive landscape. The speed and direction of a population that moves across the adaptive landscape, are given by the topology of the landscape and the evolutionary forces that act upon the population. Interestingly, this is supported by the observation of twins infected with the same strain of VIH; having the same genotype they had the same adaptive landscape and developed similar immunological responses to the virus[2]. The forthcoming of personal DNA sequencing stresses the need to understand how the genotype of one individual shapes the adaptive landscape for a pathogen.
Given the medical importance of the process that lead to drug resistance, some strategies have been devise to prevent it:
- High doses of drugs are administered (Ex. antibiotics) to reduce a population and hence reduce the number of new individuals with a mutation that are produced.
- Many different drugs are administered at the same time as cocktails (Such as the HAART that is used for AIDS treatment)so that the population has to move a very large distance in the adaptive landscape in order to reach a fitness peak.
- Drugs are designed in such a way that their action is not inhibit by a single mutation, they have a less specific active area thereby widening the valley the populations have to cross.
The aforementioned approaches are evolutionary approaches that deal with drug resistance yet they are very crude and to often their benefits are partial; after some time with the treatment the viruses and bacteria develop drug resistance. In order to deal with drug resistance in a parsimonious way it would be useful to study how does the drug affects the virus o bacteria.
The design
I aim to construct a devise in which the relation between the different genotypes and some of the different evolutionary forces are known and they may be easily measured and modified. By analyzing the data derived from the devise, an adaptive map can be represented, and subsequently the movement of the population across the adaptive landscape could be predicted.
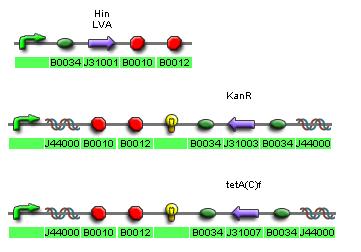
In order to achieve such an ambitious goal I intend to construct a devise with two different regions susceptible to the action of invertases. An inversion in the system would allow the expression of a reporter and will confer the bacteria with resistance to certain antibiotic.
I intend to culture all the bacteria with nonlethal concentration of Kanamycin and Tetracycline and to estimate the fitness by means of colony counting. Having only two region susceptible to inversions the system will represent two different genes each one with two alleles. The topography resulting from the mutations will be fairly simple being that there are no intermediate mutations between the two alleles.
The Promoters are unspecified and I just require them to be constitutive. The reporters are unspecified since most reporters will do the job, I will choose the ones that will allow to assemble the construction easy with few ligation steps.
Some proteins are inverted to highlight that they will only be expresed with the inversion.