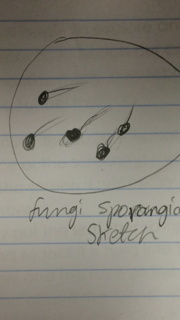User:Eyerusalem Abebe/Notebook/Biology 210 at AU
March 10, 2015 Zebrafish Development
Purpose:
The purpose of this lab is to learn the different stages of embryonic development of zebrafish. Be able to compare embryonic development in different organisms. Another purpose of this experiment is to set up an experiment in which the condition if retinoic acid, can affect embryonic development of zebrafish. If retinoic acid is added to the environment of the zebrafish embryos, then the development of these zebrafish will be stunted or negatively affected.
Materials and Methods:
Obtain 2 petri dishes; label one of these “control”, and the other “Retinoic Acid”. The retinoic acid is considered the experimental condition. Fill the “control” petri dish with 20 mL of Deerpark water use a pipeter to measure the amount of water used. For the experimental dish, pipette 20mLs of the diluted retinoic acid prepared by the lab instructor, make sure when changing the water or retinoic acid mix, to use the same water or mix that is necessary. Use 20 healthy and translucent embryos per dish. Transfer these embryos of zebrafish using a dropper, 20 embryos belong in each dish. Make sure to organize an observation schedule with the lab group, for the next two weeks. Make careful observations and record them over the next two weeks in Open Wetware. After 4-5 days remove 10mL of water and remove any dead embryos (removal is necessary or the dead embryos will mold causing more of the fish to die). Add 25mL of fresh water or retinoic acid, to the dishes. The dead embryos are saved in paraformaldehyde. After a week remove 5mL of water and any dead embryos, and add 5mL of fresh water or retinoic acid. Preserve 3 embryos from the control and test group, and preserve in the paraformaldehyde. Between days 7 and 14 remove 5mL of water or test solution and add 10mLs of water or test solution. After day 7 two drops of paramecium was added to each of the petri dishes in order to feed the zebrafish. On day 14, final observations were made, and the surviving embryos were collected and placed in an aquarium for safekeeping, and dead embryos were discarded.
Results:
Day 1: Number of dead control embryos: 0 Number of dead variable embryos:0 Number of control embryos: 20 Number of variable embryos: 20
Day 3:
Number of dead embryos control: 0
Number of dead embryos variable: 0
Number of living embryos still in egg cases control: 20
Number of living embryos still in egg cases variable: 20
Number of living hatchlings (larvae) control: 0
Number of living hatchlings (larvae) variable: 0
Number of dead hatchlines: 0
Day 5:
Number of dead embryos control: 6
Number of dead embryos variable: 4
Number of living hatchlings control: 14
Number of living hatchlings variable: 16
There were not any visible differences between the two groups. The zebrafish were mostly in 48-hour stage of development, while some were in the 24-hour stage of development. The zebrafish were moving really fast almost springing up after a slight tap. There was no heartbeat observed. The control zebrafish had a darker appearance. Also the water and retinoic acid were accidentally dumped out of the petri dish container until the instructor noticed and placed more within the dishes.
Day 7:
Number of living hatchlings (larvae) control: 11
Number of living hatchlings (larvae) variable: 14
Number of dead hatchlines control: 9
Number of dead hatchlines variable: 6
Degree of body and tail pigmentation (melanophores) control: darker definition lines
Degree of body and tail pigmentation (melanophores) variable: lighter pigmentation
Eyes and eye movement: eyes were developed on both the control and variable groups however
no movement was noted
Pectoral fin development: no pectoral fin development was noted
Yolk sac size (absorbed by day 5-7): developed by both the control and variable group
General movement control: fish glided across the petri dish, especially once disturbed
General Movement variable: fish glided across the petri dish at a faster more excited
In the variable petri dish the zebrafish were moving quickly and shaking, a couple of the fish do not move unless they are disturbed, the zebrafish in this petri dish have a yellow hue. The zebrafish in the control petri dish seemed to be slightly underdeveloped in comparison to the variable zebrafish, including with movement. The control zebrafish moved at a slower pace than the ones from the variable group.
Day 14:
Number of dead hatchlines control: 20
Number of dead hatchlines variable: 20
All of the embryos were found to be dead within the petri dishes.
Conclusion:
The purpose of this experiment was to learn the different stages of embryonic development and to set up an experiment to study how environmental conditions can affect embryonic development. The hypothesis of this lab states that the addition of retinoic acid to the environment of the zebrafish will stunt or negatively affect the embryonic development of the zebrafish. This hypothesis is negatively rejected because from the movement of the zebrafish, and the stage developments of the variable (retinoic acid), it seemed as though that variable allowed the fish to develop and move faster. The only difference in appearance between the two samples was the yellowish pigmentation of the zebrafish within the retinoic acid. The control (water) had a darker or blacker pigmentation of features, rather than the yellow of the retinoic acid. If this experiment was to be redone the observations should be taken daily, to see exactly how many fish die per day. Also the petri dishes should be stored in a safer manner, due to the fact that the liquid was spilled out of the petri dish at the beginning of the experiment.
EA
March 3, 2015 16s DNA Sequencing
Purpose: The purpose of this experiment is to identify the unknown bacterium that was grown in tetracycline and nutrient agar plates. This bacteria originated from the transect (the community garden) on American University’s campus. The identification is done through DNA 16s sequencing, which will allow for the practice using software relating to genetic identification.
Materials and Methods: Using a single colony from each of the two plates (One containing tetracycline, and one without), prepare a PCR tube for 16s sequencing. After one week, the PCR products of these bacteria will be ran in a gel, and sequenced. Using sequences provided by Genewiz.com (the lab) the copy and paste the sequences into NIH’s BLAST online identification system, and find the 16s DNA sequence this will describe what bacterium is present.
Results:
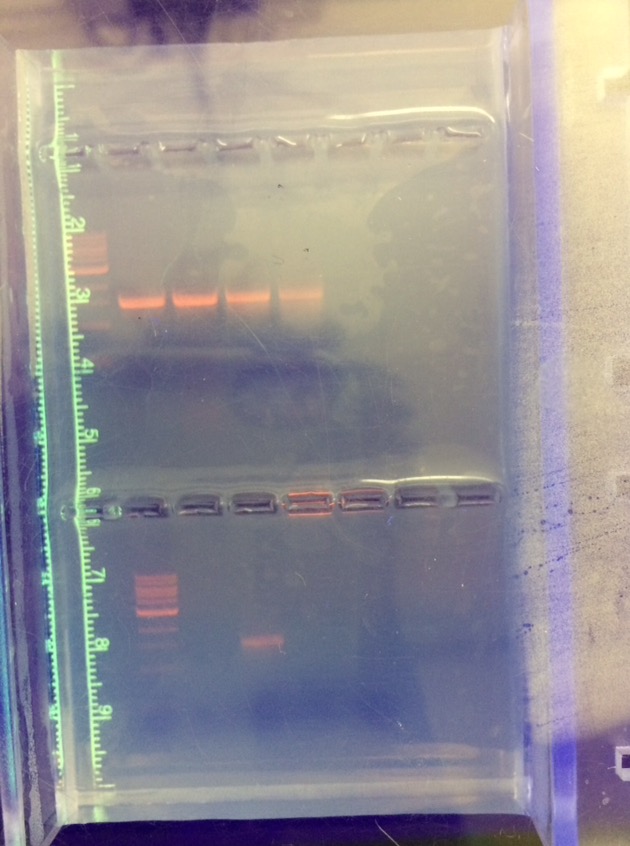
Figure 1: Gel with Samples of Each Bacteria Tested
Samples A and D are in lanes 2 and 3.
Raw Sequences: Sample A: NNNNNNNNNNNNTNNNNNNNGNNNCTCCTNNNACGNGTCACCGNCTTCAGGTACCCCATNNNNNCCATGGCTTGACGGGC GGTGTGTACAAGGCCCGGGAACGTATTCACCGCGCCATGGCTGATGCGCGATTACTAGCGATTCCAGCTTCATAGAGTCG AGTTGCAGACTCCAATCCGAACTGAGACCAGCTTTCGAGATTCGCATCCAGTCACCTGGTAGCTGCCCTCTGTACTGGCC ATTGTATTACGTGTGTGGCCCAAGGCGTAAGGGCCGTGATGATTTGACGTCATCCCCACCTTCCTCTCTACTTGCGTAGG CAGTCTCACTAGAGTCCCCAACTGAATGATGGCAACTAGTGACAGGGGTTGCGCTCGTTGCAGGACTTAACCTAACACCT CACGGCACGAGCTGACGACAACCATGCAGCACCTTGAAAAATGTCCGAAGAAAAGTCTATTTCTAAACCTGTCATTTCCC ATTTAAGCCTTGGTAAGGTTCCTCGCGTATCATCGAATTAAACCACATAATCCACCGCTTGTGCGGGCCCCCGTCAATTC CTTTGAGTTTCAGACTTGCGTCCGTACTCCCCAGGTGGCTAACTTATCACTTTCGCTTAGTCTCTGAATCCGAAAACCCA AAAACGAGTTAGCATCGTTTACGGCGTGGACTACCAGGGTATCTAATCCTGTTCGCTCCCCACGCTTTCGTCCATCAGCG TCAGTTGTTGCTTAGTAACCTGCCTTCGCAATTGGTGTTCTAAGTAATATCTATGCATTTCACCGCTACACTACTTATTC CAGCTACTTCAACAACACTCAAGACCTGCAGTATCAATGGCAGTTTCACAGTTAAGCTGTGAGATTTCACCACTGACTTA CAGATCCGCCTACGGACCCTTTAAACCCAATAAATCCGGATAACGCTTGCACCCTCCGTATTACCGCGGCTGCTGGCACG GAGTTAGCCGGTGCTTATTCGTATAGTACCTTCAGCTTNCCACACGTGGAAAGGTTNNTCCCTATACNAAAGAAGTTTAN NNCCATNNGNNNTCGNCTTCACGCGGGNTGNTGGNTCAGNTCTCANCCNTGNCNANNTCNTCANNGCTGCNTCCCGNAGN ANTCNGGNCCNNNNNNTCAGNACNNNNNGNGGGGNANNCN
Sample D: NNNNNNNNNNNNGNNNTTANNNNTGCAGTCGNNCGANNGAGTAGCNCNNNNTNNCGGACGCTGACGAGTGGCGAACGGGT
GAGTAATACTATCGGAACGTGCCCAGTCGTGGGGGATAACTACTCGAAAGAGTAGCTAATACCGCATACGATCTGAGGAT
GAAAGCGGGGGACCTTCGGGCCTCGCGCGATTGGAGCGGCCGATGGCAGATTAGGTAGTTGGTGGGATAAAAGCTTACCA
AGCCGACGATCTGTAGCTGGTCTGAGAGGACGACCAGCCACACTGGGACTGAGACACGGCCCAGACTCCTACGGGAGGCA
GCAGTGGGGAATTTTGGACAATGGGCGAAAGCCTGATCCAGCAATGCCGCGTGCAGGATGAAGGCCTTCGGGTTGTAAAC
TGCTTTTGTACGNAACGAAAAAGCTTCTCCTAATACNAGAGGCCCATGACGGTACCGTANNAATAAGCACCGGCTAACTA
CGTGCCANCAGCCGCGGTANTACGNANGGTGCGAGCGTTNATCGCGANTTTACTGNGCGTAAAGCGTGCGCANGCNGNTA
TGTNANGACANNATGTGAAATCCCCNNNNTCTNTCCTGNTNAACTGNCNTTTGTGNCTGCNNGGNTAAANTACGGNAGAG
GNGTGNTGGAAT
Results from BLAST (identification):
Sample A: “Chryseobacterium sp. C” 16s ribosomal RNA gene, partial sequence
Sample D: “Uncultured bacterium clone”16s RNA gene, partial sequence
Conclusion: Sample D is unidentified, because it is considered an uncultured bacteria, however sample A, is a Chryseobacterium. Chryseobacterium is a bacteria that is found in soil and water, it can grow despite the presence of chlorine (in water), it is also found to resist some antibiotics including tetracycline (Kirby et.al, 2003). This is a significant factor because the bacterium from the transect did come from soil, and was grown in tetracycline nutrient agar plates. The bacteria grew despite the presence of this antibiotic, on most plates, so the resistance described is accurate. The transect also has water because of the fact that it is outdoors, and whether (rain, snow, ice) is present, and the transect is watered by the keepers of the Community Garden, so it does make sense that the bacteria also appears in water.
Reference: Kirby, J., Sader, H., Walsh, T., Jones, R.. “Antimicrobial Susceptibility and Epidemiology of a WorldwideCollection of Chryseobacterium spp”. 2003. Journal of Clinical Microbiology. (4 Mar 2015). < http://jcm.asm.org/content/42/1/445.full.pdf>
EA
February 15, 2015 Vertebrates and Niches
Purpose: The purpose of this experiment is to identify the different vertebrates that are present within the niche of the explored transect. The identification of different vertebrates will allow for the better understanding of what is taking place within the transect. The identification of the different vertebrates will also allow for an understanding of whether the transect is a single niche specifically for those organisms, or if the transect is a part of their niche.
Materials and Methods: The observations were made during the daytime between the hours of 2:35-5:15pm, on days where the temperature was about 20°C. The observations were taken from near the transect location and on the transect location, through footprints or appearance of the vertebrates.
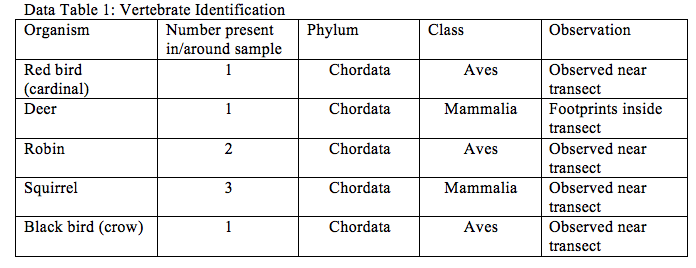
Data and Observations (Results):
Discussion:
The biotic factors of the community garden that would help the found organisms are possibly the live plants that they would be able to ingest. The abiotic factors of the community garden that would help benefit the vertebrates for example the squirrels is the structure and soil, in order to dig and bury items.
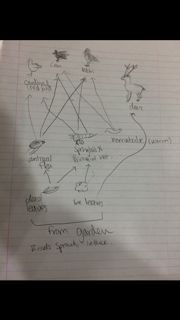
Food web based on community garden:
This demonstrates how the organisms work together in order to sustain the community. The birds are on the same topographic level, meaning the same feeding level, while leaves shared the same level, and the arthropods share the same level. The carrying capacity of this system, may be greater, however we observed the eight items within the food web. Carrying capacity is the maximum number of individuals that can exist within an area, without depleting the resources of the area (McConnel & Abel, 2013).
References:
Linzey, A.V., Koprowski, J. & Hammerson, G. “Sciurus carolinensis”. 2008. The IUCN Red List of Threatened Species. (15 Feb. 2015) < http://www.iucnredlist.org/details/42462/0> McConnell, R.I. & Abel, Daniel C. “Population Size”. 2013. Western Oregon University. (18 Feb. 2015). < https://www.wou.edu/las/physci/ch371/lecture/popgrowth/carrying.htm.> n.a. “American Crow- Corvus brachyrhynchos”. 2015. Wildlife Journal Junior. (15 Feb. 2015) < http://www.nhptv.org/wild/americancrow.asp> n.a. “American Robin - Turdus migratorius”. 2015. Nature Works. (15 Feb. 2015) <http://www.nhptv.org/natureworks/robin.htm> n.a. “Northern Cardinal - Cardinalis cardinalis”. 2015. Nature Works. (15 Feb. 2015) < http://www.nhptv.org/natureworks/cardinal.htm> n.a. “White Tail Deer”. 2015. Squam Lakes Natural Science Center. (15 Feb. 2015) < http://www.nhnature.org/visit/animal_info_sheets/white_tailed_deer.php>
EA
February 15, 2015 Invertebrates
Purpose: The purpose of this experiment is to understand the significance of invertebrates. Invertebrates are also described and identified by the structural and visual appearance. Another purpose of this experiment was to learn how simple systems evolved into more complex systems.
Materials and Methods:
Part 1: Acoelomates, Pseudocoelomates, and Coelomates Observe the acoelomate, Planaria, under the dissecting scope. The acoelomate has just been fed Planaria egg yolk so the process of digestion must be examined. Also examine the cross section of the whole mount of Planaria with a microscope, and the whole mount is stained in order demonstrate the appearance of the digestive tract of the organism.
Obtain the nematodes, and a cross sectional slide of their pseudocoelomate structure, and take note of their structure.
Observe the coelomate Annelida under the dissecting scope, and look for the position of their internal organs (the layers of muscle which are pink).
Part 2: Arthropods Observe the five major classes arachnida, diplopoda, chilopoda, insect, and crustacea. Look for the differences between each based upon their body parts, body segments, and number of appendages.
Part 3: Analyzing the Invertebrates Collected with the Berlese Funnel First carefully take down the Berlese Funnel, and pour the top 10-15mLs of the liquid and organisms into a petri dish. Pour the rest of the liquid and remnants into another petri dish. Use a dissecting microscope to observe to find the invertebrates. To identify which class of arthropoda invertebrates that are observed, use the dichotomous key, and figure 3, from the lab notebook. Identify at lease five invertebrates, and measure the length of each of the organisms.
Data and Observations:
Part1: Acoelomates, Pseudocoelomates, and Coelomates The movement of the acoelomate was smooth; it was sliding straight across the viewpoint of the microscope with ease and it appeared to be a small wormlike organism with what seemed to be almost an arrow like head, with two holes near the tip of it.
The movement of the nematode was squirmy and wormlike, which matches its structure. It was slowly moving in place.
The movement of the coelomate Annelida, was twisting and slowly moving, this makes sense for its structure because it was a bit thick and wormlike.
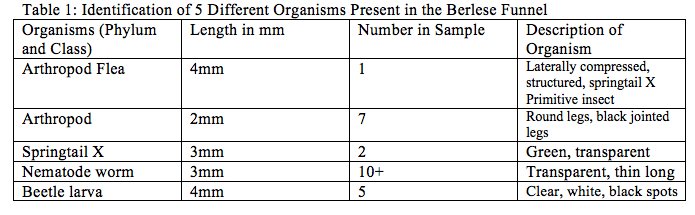
Part 3: Analyzing the Invertebrates collected with the Berlese Funnel.
The size ranges of the observed organisms is between 2-4mm. The smallest organism was 2mm it was the arthropod identified, and the largest organisms were both the arthropod flea and the beetle larva, both were 4mm. the most common organism within the leaf litter was the nematode worm, there were more than 10 present in the microscope.
Conclusions and Further Directions:
The purpose of this experiment was to understand the significance of invertebrates, and to learn how simple systems evolved into more complex systems. Using the dichotomous key and the images available in figure 3 this identification was done and recorded within Table 1, of this report. The five different organisms found were an arthropod flea, arthropods (without definite class/phylum organization), Springtail X, nematode worm, and beetle larva. If this experiment were to be redone a better understanding of what arthropods appearance should be established through more available images, because it was difficult to make sense of what the dichotomous key was describing at times. Also the dichotomous key should have more options available because for instance the second organism is listed in the table as “arthropod”, because it was clear that the organism present was an arthropod, however there were little to any descriptions that actually matched exactly what was being viewed under the dissection microscope.
EA
February 8, 2015 Plantae and Fungi
Purpose: The purpose of this lab is to understand the characteristics and diversity of plants. Fungus is also observed and studied within this lab. Another important factor of this lab is to appreciate the function and importance of fungi.
Materials and Methods:
Part 1: Collecting five plant samples from the transect First obtain three Ziploc bags and a pair of gloves, then go to your transect. Find a leaf litter sample from the transect and collect it within the bag. Place about 500 grams of leaf litter in one bag include dead leaves and plant and a little bit of the crumbly top layer of soil into another bag. This collection will be used to set up the Berlese funnel for collecting invertebrates. In the other bags, take a representative samples from five plants in a way that is not damaging, and make sure to choose a diversity of plants. Make sure to take a photo of any trees used in the sample, and collect any seeds, pinecones, and flowers available. Make sure to describe the location and identity of the plant, and use the information to describe and identify the plants.
Part 2: Plant Vascularization Observe the moss, Mnium make sure to write down the height of the plant, and compare it to the height of the lily plant stem. Look at the cross section slide of the lily stem and find the xylem and phloem layers. Make sure to characterize the vascularization in each of the plants from the transect, write down the height and observe the cross section from each plant.
Part 3: The Presence of Specialized Structures Look at the leaves of the moss using the dissection scope or under a low magnification of the compound scope. Observe the smooth surfaces that have the waxy cuticle. Describe the shape, size and cluster arrangement of the leaves collected from the transect. Look for attachment sites or evidence of leaves in the leaf litter collected.
Part 4: Mechanisms of Plant Reproduction Look at the moss, Polytrichum, and identify the male and female gametophytes and sporophyte, and identify the parts of the life cycle that are haploid and diploid. Using the seeds present from the transect, identify the parts of the seeds as listed in the hand out. (There were no seeds present at the transect)
Part 5: Observing Fungi • Understand why fungi sporangla are, and their significance. • Look at some samples of fungi with the dissecting microscope and decide if they are fungi and which of the three groups they belong to • Draw a picture of the fungus and explain why it is a fungus.
Data and Observations:
No seeds were collected from the transect.
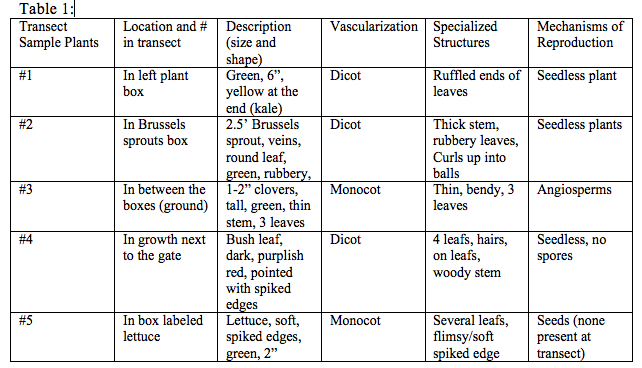
Part 5: Observing Fungi Sketch of fungi sporangia
Fungi sporangia are hyphae of fungi that grow upward and form small, black globelike structures. Inside of the sporangia there are cells that are called spores that release when the sporangia open.
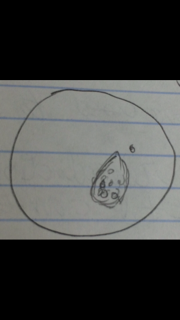
Example of Fungi:
This is a sketch of a mushroom. A mushroom is a fungus because it produces the cells called spores, which are released by its surface (Wilson, 2015). Since fungi release spores after the growth of the hyphae, the fact that mushrooms release spores is what allows them to be considered fungi.
Conclusions and Future Directions:
In this experiment several types of plant and soil matter were collected and observed from the transect (AU Community garden). Each plant sample was identified as either dicot or monocot using the appearance. We observed whether each item had a network of veins, or flowers. A description was provided of the appearance, mechanism of reproduction, and specialization was provided for each sample of plant. The hypothesis of this lab is accepted however no seeds were found, however it is important to observe that monocot sample #5, reproduces by seeds but no seeds were found. The other monocot #3 was an angiosperm (produces flowers) rather than a plant without seeds or flowers. If this experiment were to be redone, the transects should be more carefully observed, and pictures should be taken of each plant that the samples were drawn from, in order to better identify the specialized structures and mechanisms of reproduction.
References:
Wilson, Nathan. “Mushroom”. What is a Mushroom? 2015. Encyclopedia of Life. (11 Feb. 2015) < http://eol.org/info/453>
EA
February 4, 2015 Microbiology and Identifying Bacteria With DNA Sequences
Purpose: The purpose of this lab is to understand the characteristics of bacteria. Antibiotic resistance is also observed within bacteria. Also to learn how DNA sequences are used to identify species.
Materials and Methods:
Part 1: Quantifying and Observing Microorganisms Observe the growth from the Hay Infusion Culture from the agar plates from the previous week. Count the number of colonies on each plate, take note of the dilution and if there is too many colonies to count label it lawn, record this in table 1.
Part 2: Antibiotic Resistance Observe the results from table 1 and in the agar plates and observe the following: if there is differences within colony types between the plates with or without the tetracycline, how the tet affected the number of bacteria or fungi present, and how many species of bacteria present or unaffected by tet.
Part 3: Bacteria Cell Morphology Observe a wet mount and gram stain preparation of the bacteria grown in the plates. For the wet mount, sterilize a loop over a flame, and then scrape a small amount of grown from the surface of agar. Place a drop of water on a slide and mix the bacteria from the loop onto the slide, then place a coverslip over it. And observe the wet mount using 10X and then the 40X objective, it will be difficult to see the organisms, then determine the organism shapes and if the organisms are motile. With the gram stain process sterilize the loop over a flame and scrape a small amount of bacteria from plate. Mix it in a drop of water on a slide. Circle the area underneath with a sharpie or wax pencil. Label the slide. Heat the slide by passing it through the flame three times with the smear side up. While working on a staining tray, cover the smear with crystal violet for 1 minute. Then rinse the stain with water. Cover the smear with Gram’s iodine mordant for 1 minute, and then rinse off with water wash bottle. Decolorize the smear by flooding the bacterial smear with 95% alcohol for 10-20 seconds. Rinse it gently. It is finished decolorizing once the solvent flows colorlessly off the slide. Cover the smear with safranin stain for 20-30 seconds, and then rinse it off using a wash bottle of water. Dry the excess water by carefully blotting with a kimwipe and allow it to air dry. Focus the gram-stained sample on a low magnification, then under 40X and 100X oil immersion objectives. Record the results in Table 2. Repeat the gram stain and wet mount process for four of the colonies present.
Part 4: Set up PCR for 16S sequencing Take one from each of the two plates of nutrient agar and tetracycline that has the most complete characterization of bacteria. Transfer a colony of bacteria to 100 microliters of water in a sterile tube. Incubate the tube at 100°C for 10 minutes on a heat block and make sure the tubes are floating in the heat block. Centrifuge the samples for 5 minutes at 13,400rpm. During the centrifuging process take the PCR tube and add 20 microliters of the primer/water mixture, and mix it to dissolve the PCR bead. Take 5 microliters of the supernatant from the centrifuge samples into the 16S PCR reaction; place the tube in the PCR machine.
Data and Observations:
Observations of Hay Culture: • Half of the water evaporated • Smells like sewage • Brussels sprout shrunk • Clearer water • There still a growth that looks like skin on top • Soil on the bottom of the culture • Achaea species most likely did not grow in the agar plate, because it is no an extreme environment with a condition like high temperatures, it is a place that caters to the growth of bacteria.
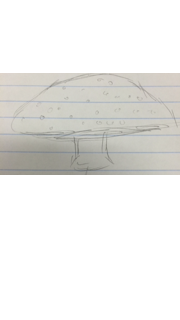
Table 1:100-fold serial dilutions results
Procedure 2 Observations:
• Different types of colonies are identified by appearance or shapes
• Tetracycline caused resistance
• There are yellowish mustard colonies, yellow colonies, and white colonies
• No fungus was present
• Two species of bacteria are unaffected by tetracycline
The mechanism of action that tetracycline takes is it inhibits the protein synthesis of bacteria by preventing the attachment of aminoacyl-tRNA to the ribosomal acceptor (A) site. Tetracycline inhibits bacteria that are gram-positive and gram-negative, such as mycoplasmas, chlamydiae, protozoan parasites and rickettsiae (Chopra and Roberts, 2001).
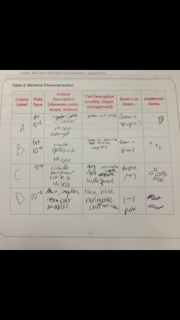
Table 2: Bacteria Characterization
Conclusions and Future Directions:
In this lab it was found that bacteria from the Hay Infusion Culture has various cultures of bacteria including some that resist the antibiotic tetracycline. It was also discovered that the nutrient plate with the least dilution produced the most colonies, while the higher dilutions tetracycline produced zero colonies. The bacteria characterization done after the gram stains presented that exactly half of the bacteria tested are gram positive, and the other half is gram negative. If this experiment were to be redone the procedures for the wet mount need to be done in a careful manner so that the bacteria can be viewed under the objectives, a stain or dye maybe necessary to view the bacteria as well because the samples were transparent. The gram staining procedure also needs to be observed and practiced carefully because the process of drying the wet mount over the flame caused the burning of bacterial cells.
References:
Chopra, Ian and Roberts, Marilyn. (2001). Tetracycline Antibiotics: Mode of Action, Applications, Molecular Biology, and Epidemiology of Bacterial Resistance. Microbiology Molecular Biology Review, 65(2). Retrieved from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC99026/
EA
January 26, 2015 Identifying Algae and Protists
Purpose: The purpose of this lab is to learn how to use a dichotomous key in order to identify unknowns. The characteristics of algae and protists are described and understood. Also algae and protists are examined from the assigned transects.
Materials and Methods:
Part 1: How to use a Dichotomous Key Using a slide and cover piece make a wet mount of the known organism. Observe the organism under a microscope at 4X and 10X. After finding an organism record information about it including its description and size using the ocular micrometer. With the 4X view each space is 25μm while for 10X each space is 10μm. Take a dichotomous key for the known organism and follow the key to find the organism, and confirm the image with the diagrams present in the key.
Part 2: Hay Infusion Culture Observations Carefully take the Hay infusion culture from the previous week to work area, and take note of its appearance. Describe in detail any growth, smell, and appearance. Make a wet mount from two different niches of the culture, observe them under a microscope and find what protists and algae are using the dichotomous key. Draw pictures of the different organisms observed, and describe and identify three different organisms each from the two niches (6 total). Make sure to identify if the organisms are motile or non-motile and identify if the organisms are algae or protozoa. Make sure to measure and record the sizes of each organism using the ocular micrometer.
Part 3: Preparing and Plating Serial Dilutions Take four tubes of 10mLs sterile broth and label with markings 10-2, 10-4, 10-6, and 10-8. Also retrieve a micropippetor set at 100μL. Take four nutrient agar plus tetracycline plates, and label with “tet” and the dilution values of 10-2, 10-4, 10-6, or 10-8. Place the lid on the Hay Infusion Culture, and swirl to mix up the organisms. Using the micropippetor pipette 100μL of the culture into the tube labeled 10-2, swirl retrieve another tip and take 100μL of the contents from the 10-2 tube and add it to the broth labeled 10-4. Continue this process for the next two tubes. To plate these cultures, take the first tube labeled 10-2, and pipette 100μL of the culture into a “tet” plate and spread the sample carefully on the plate, and repeat on the plate without “tet”. Label accordingly. Repeat the procedure with the following three dilutions. Store the agar plates agar side up on a rack and it will incubate for a week.
Data and Observations:
Part 1:Observations of known culture
Colpidium sp 50-70μm
Part 2: Observations of culture: Image of culture:
• Smells like spoiled vegetables • Appears to have almost a skin like collection on the top of the liquid, which includes mold, a fuzzy white clump, Brussels sprouts, and soil on the bottom of the liquid
• When viewing liquid near the fuzzy white clump, the culture was green in appearance and had very slight movement within the green clump, however it was non motile.
• When viewing the liquid near the Brussels sprout it had several structures present some that appeared to look like worms and others with nuclei. The structures were motile and had cilia.
• Using the dichotomous key, colpidum sp, paramecium bursaria, and paramecium multimicronucleatum were identified. Several other organisms were present that appeared to be various types of paramecium however they were not described on the dichotomous key so remain unidentified.
Image of microscope view sketches
Conclusions and Future Directions:
In this lab a dichotomous key was used in order to identified the different types of species present. The protists and algae were identified from the Hay infusion culture using a microscope and dichotomous key. It was very difficult to identify some of the organisms because they had several different types of characteristics present in each viewing so it was difficult to identify certain factors that the dichotomous key asked. Next time for this experiment it would be beneficial to get a more comprehensive dichotomous key where more types of algae or protists can be identified, because some of the ones from the hay infusion looked like a mixture of a few types of organisms.
EA
January 25, 2015 Biological Life at American University
Purpose:
The purpose of this experiment is to learn how natural selection allows for evolution of different species to exist. Another purpose of this experiment is to make careful observations about certain characteristics of a niche. A niche is the description of where organisms interact with the environment, and this area is described by both living and nonliving factors of the environment.
Materials and Methods:
Part 1: The Volvocine Line Chlamydomonas (a motile living culture) is obtained. A wet mount slide is prepared, this means pipetting a single drop onto a slide and covering it with a coverslip, it is then observed under a microscope. If the particles are moving too fast protoslo, is added at the side of the coverslip. After the microscope is adjusted the characteristics such as number of cells, colony size (in μm), specialization of cells, mechanisms of motility, whether or not the sample is isogamous or oogamous, and a picture of the specimen are recorded. These steps are then repeated for the live cultures of gonium, and volvox.
Part 2: Observing a Niche at AU With an assigned group observe the 20 by 20 meter dimensions of the community garden at American University (take note of the number of the transact). Use the number of the transact to label all of the samples collected from this part of the lab. Use a 50 mL conical tubes in order take a sample of the garden including soil and any ground vegetation present. Then once back in lab a Hay Infusion Culture is made. The culture is made by taking 10-12 grams of the collected soil vegetation sample in a plastic jar, with 500 mLs of deerpark water. 0.1 grams of dried milk, is then added in the jar and carefully mixed for about 10 seconds. The top of the jar is then removed, the jar and top is labeled with the group name and number, then kept in the lab for one week undisturbed in order for the bacteria present to grow.
Data and Observations:
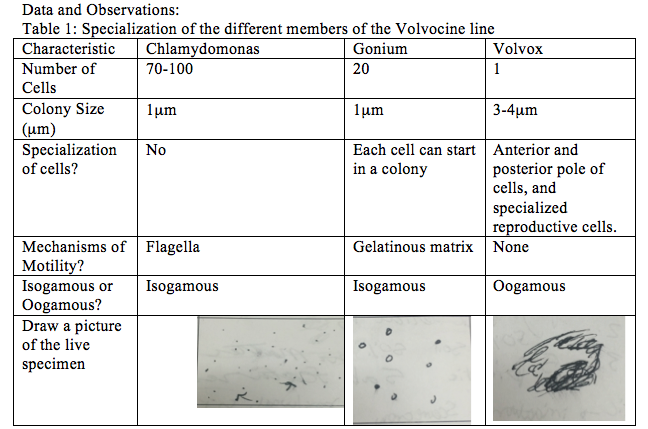
Figure 1: Topographic aerial map of Community Garden 
The assigned transect contained plants grown by the community garden members, including cucumbers, lettuce, brussel sprouts, and spinach, these were biotic (live) components. Pine leaves and dead leaves (which weren’t grown on purpose) were the other biotic components present within the dimensions. The abiotic (nonliving) factors that were present in the garden were the irrigation system, a scarecrow, snow, a box (made of wood), and soil. These biotic and abiotic factors were found within the boxes constructed for the plants to grow in.
Conclusions:
In this lab the colony size specialization, motility mechanism, and reproduction type was identified. The Chlamydomonas and Gonium were both found to be isogamous, and 1μm in size. The Volvox was found to by oogamous and 3-4μm in size. Gonium and Volvox both have reproductive specializations. In order to do this part of the experiment better next time, I would suggest using protoslo on most of the wet mounts, because the view of the specimens was too fast for some slides. The observation of a niche lab was a collection of certain biotic and abiotic factors of the transect, it allowed us to understand what to take note of when observing an environment. If this part of the experiment were to be redone I would suggest taking a picture of the 20 by 20 transect because it was difficult to understand the sketch from the site because it did not have enough details drawn.
EA
January 19, 2015
I would like to practice reading and writing Amharic so I can think, read, write and speak in both English and Amharic.
EA