User:DavidRamos/Lab/2006-6-14
From OpenWetWare
Jump to navigationJump to search
June 14th
Morning
Plasmid Miniprep
- lac operon promoter
- R0010
- promoter and GFP
- E0241
- GFP
- E7104
- DNA Miniprep of transformant colonies
- Out of 5mL of liquid culture, reserved 1mL for gylcerol+freeze and 4mL other
- followed QIAprep Miniprep Kit for Microcentrifuge directions
- Eluted with warm dH20
- Put at 40C for ~2 min to evaporate ethanol before elution
- Forgot to label after elution --> don't know what is what
- Sol'n: During digest will have to run PCR, can tell R0010 from rest, but E7104/E0241 is only different by 30bp; if doesn't work can flip and try two experiments.
- Nanodrop demonstration
Nanodrop results -1: 31.2ng/uL 260/280:1.75 260/230:1.92 -2: 38.5ng/uL 260/280:1.83 260/230:1.87 -3: 33.8ng/uL 260/280:1.70 260/230:1.44
PROBLEM: Messed up labeling of the plasmids! To diagnose, ran a 15min e-Gel to find out which is R0010.
- Digestion of vector/insert
- Digested R0010 (200bp cutout) as vector at S and P site.
- .5uL Spe1, .5uL Pst1
- 11uL h20
- 2.5uL 10X BSA
- 2.5uL #2 NebBuffer
- 8uL DNA
- Digested other 2 (~900bp cutout) as insert at X and P site.
- .5uL Xba1, .5uL Pst1
- 11uL h20
- 2.5uL 10X BSA
- 2.5uL #3 NebBuffer
- 8uL DNA
- Incubate @ 37C for 1h
- Digested R0010 (200bp cutout) as vector at S and P site.
- Phosphatase
- 80C@15min to kill enzyme activity
- Used CIP (1 unit) into the R0010, 1h@37C
- Run on 1% agarose gel
- Image, Cutout, and Purify
- Can isolate the three from the gel
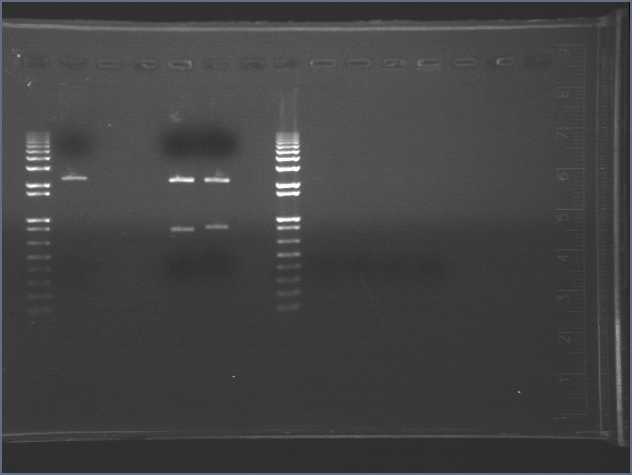
Result
Ladder=1kb+ Lane 1=R0010 (#1) Lane 2=E0241 (#2) Lane 3=E7104 (#3)