User:Cassandra V. Sharp/Notebook/Biology 210 at AU
4.7.15 Missing 16S entry. SK
Embryology and Zebrafish Development 2/25
Purpose: The purpose of this second unit of study is multifaceted. In it, we will observe and understand the various stages of embryonic development, particularly those of the zebrafish. We will also observe the effects of environmental factors on embryonic development by conducting a study in which we expose developing zebrafish to a variable of our choosing.
Materials and Methods:
Day 1: Thursday -we paired up with partners and chose a variable to study in our embryogenesis experiment. I am working with Lexi Schultz. -we decided to work with caffeine, so we read an experimental paper about the effects of caffeine on embryonic development in zebrafish. We used the information in this paper to develop a procedure for our own experiment -we obtained forty live zebrafish embryos, placing twenty in a petri dish with just water and twenty in a petri dish with a water-caffeine solution. -we set these petri plates aside and allowed them to develop
Day 2: Friday -we returned the next day to make sure that the majority of our zebrafish had survived the night -we counted the number of living and dead embryos for each of our petri dishes and recorded it in our notes -we identified what stage of development our embryos were in using a chart provided to us
Day 3: Monday -we returned to our developing zebrafish after the weekend and counted the number of living and dead embryos in each plate as well as the number of zebrafish that had hatched -we identified what stage of development the remaining embryos were in and recorded this in our notes -we recorded the heartbeat rates of the control and experimental zebrafish and averaged these numbers -we measured the length of the control and experimental zebrafish and recorded this information -we fed the hatched zebrafish 2F from a small dropper -we added fresh water to these petri dishes
Day 4: Thursday -we returned to our zebra fish and counted the number of living and dead fish in each plate -we identified what stage of development these hatched fish were now in -we attempted to record the rate of heartbeat for each kind of zebrafish, but found that tracking their pulses and observing their blood was no longer possible -we measured the length of the control and experimental zebrafish and recorded this information -we added fresh water to the two petri dishes
Day 5: Monday -we returned to our zebrafish, counted number of living and dead -we measured the length of the two different types of zebrafish and recorded this information -we made observations about the movement displayed by the two types of zebrafish -we added fresh water to the two petri plates
Data and Observations
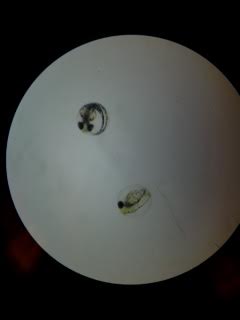
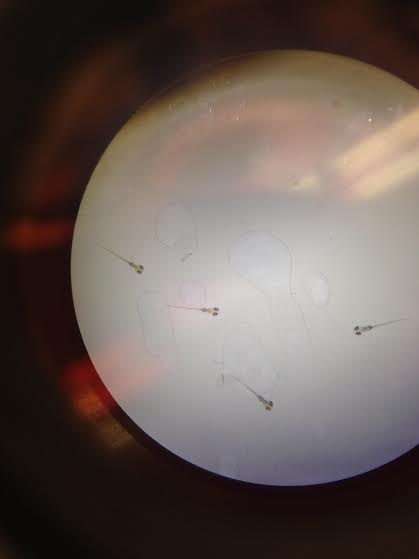
Day 2: Friday Img 1. Zebrafish embryos without caffeine exposure
Img 2. Zebrafish embryos with caffeine exposure
Control: 0 dead embryos, 20 live embryos. Appear to be at the 20-somite stage of development. Test: 3 dead embryos, 17 live embryos. Appear to be at the 20-somite stage of development.
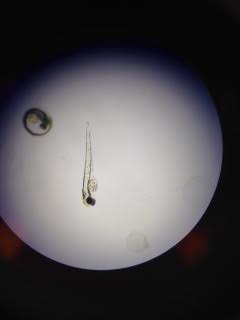
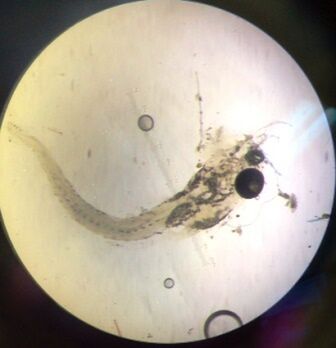
Day 3: Monday Img 3. Zebrafish embryo and hatchling without caffeine exposure
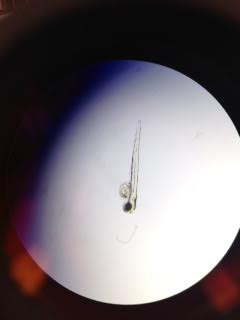
Img 4. Zebrafish embryo with caffeine exposure
Img 5. Zebrafish hatchling with caffeine exposure
Control: 8 dead, 12 alive. 10 embryos in stage H-1, 2 hatchlings in 48 hour. Avg Heartbeat: 150 BPM Avg Length: 3mm
Test: 7 dead, 10 alive. 7 embryos in stage H-1, 3 hatchlings in 28 hour. Avg Heartbeat: 122 BPM Avg Length: 3mm
Day 4: Thursday
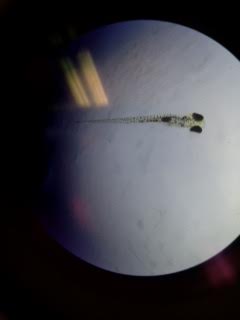
Img 6. Zebrafish without caffeine exposure
Img 7. Zebrafish with caffeine exposure
Img 8. Group of zebrafish with caffeine exposure
Control: 12 alive, 0 dead. Heartbeat: unknown Length: 5 mm
Experimental: 8 alive, 2 dead. Heartbeat: unknown Length: 4 mm
Notes: We noticed that the control zebrafish- who had not been exposed to caffeine- were zipping around their petri dish quickly with fast, tail-swishing movements. The experimental zebrafish- who had been exposed to caffeine- were at almost a complete standstill. We further observed that the control zebrafish measured an average of 5mm, while the caffeinated zebrafish measured an average of 4 mm- 1 full mm shorter than the control.
Day 5: Monday
Img 9. Zebrafish control, no caffeine
Img 10. Zebrafish with caffeine exposure
Control: 12 alive, 0 dead. Length: 5.5 mm Notes: our control zebrafish are now exhibiting a much slower speed of movement than the last time we observed them. They have not yet reached a complete standstill as the zebrafish exposed to caffeine have, but they are definitely moving less. They also seem to have grown, on average, 0.5mm.
Experimental: 7 alive, 1 dead Length: 4 mm Notes: The zebrafish exposed to caffeine are almost entirely motionless and remain at an average length of 4mm.
4.7.15 Missing conclusions SK
SS
2.23.15 Good entry. Good organization and detailed description of invertebrates. Missing a food web. SK
Lab 5: Invertebrates 2/16
Purpose This lab marked the final section of our unit studying the various ecosystems around AU. In it, we took a second look at the Berlese Funnel we had previously constructed (Lab 4) and examined the invertebrates that we had collected from our transect. This lab helped us to familiarize ourselves with another sect of life in an ecosystem and helped us to practice identifying organisms under a microscope.
Materials and Methods 1. Acoelomates, Pseudocoelomates, and Coelomates -We examined three different kinds of worms- planaria, nematodes, and earthworms- in various locations around our lab room
2. Arthropods -We walked around the room and observed various organisms from the arachnida, diplopoda, chilopoda, insect, and crustacea classes
3. Analyzing Invertebrates Collected with a Berlese Funnel -we returned to our burlese funnel that we had set up the previous lab -we took apart our funnel and poured out the liquid that had been in it into a petri dish -we obtained a microscope and examined this petri dish -using the chart in our lab books, we identified different invertebrates that had been in our funnel
Data and Observations
1. Acoelomates, Pseudocoelomates, and Coelomates: We observed the acoelomate Planaria using a dissecting microscope. It is incredibly small and looks somewhat like a leech does. It appears to move in a manner somewhat like swimming, propelling itself by wiggling its tail. We then observed the pseudocoelomate Nematode, which is also a minute organism we needed a dissecting microscope to observe. The last type of worm we examined was the common coelomate Annelida, better known as the earthworm. These worms were long, plump, and visible to the naked eye. We used a pair of tweezers to pull one out of a mound of earth and it came out twisting and wriggling its entire body.
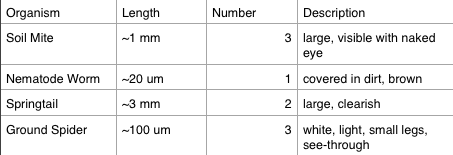
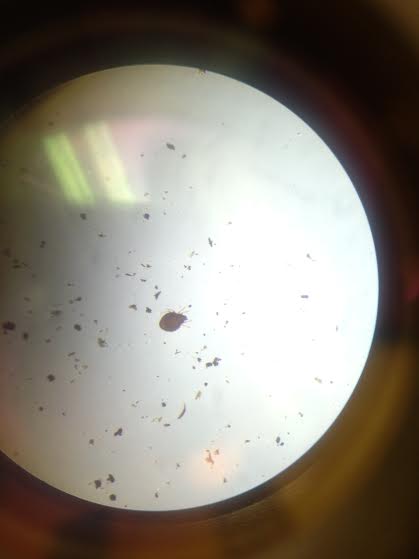
2. Analyzing Invertebrates Collected with a Berlese Funnel:
The smallest organism that we observed was the nematode, and the largest was the springtail.
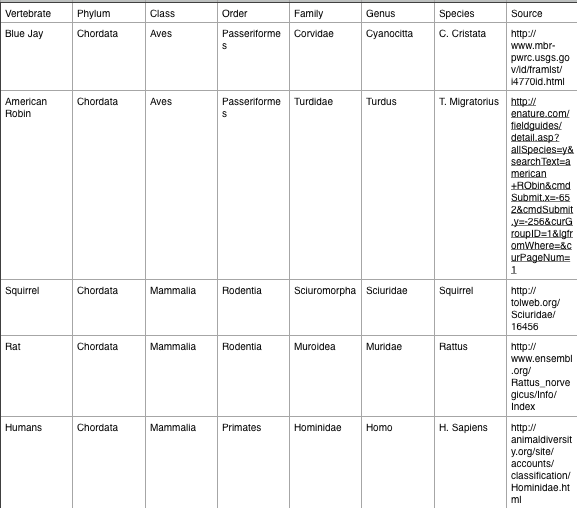
3. Vertebrates and Niches
Table 2: 5 vertebrates that might inhabit our transect
Conclusions
Biotic aspects of the transects probably with benefit and are harmed by the birds and rodents that inhabit it. These creatures eat many of the crops that grow in it, but also carry around the seeds of these plants and help them to grow in new locations. Humans have more of an effect on the abiotic features of the land, such as by constructing the fence and repairing it if it gets damaged, but also contribute to the biotic features of the land by gardening and watering the plants. Together, these various organisms and the abiotic and biotic features of the land form an ecological community of interaction. Because the garden is so small, it has a small carrying capacity and can not be home to a large population of vertebrates. The nutrients in the soil, the plants, the invertebrates, and the vertebrates that all interact together in this niche represent different trophic levels in the environment.
SS
Lab 4: Plantae and Fungi 2/15 (I was originally absent for this, so bear with me)
Purpose In this lab, we focused on collecting plant and fungi samples from the transect we had been assigned and identifying which plant orders they belonged to. We examined their internal structures and methods of reproduction, then set up a Berlese funnel to collect and study the invertebraes that might be found in our niche. Plants and fungi are an invaluable part of any ecosystem as they provide food, shelter, and important gasses. Understanding the families to which these lifeforms belong and the ways in which they operate are vital to understanding the ecosystem as a whole.
Materials and Methods 1. Collecting and observing samples -we obtained two plastic bags from Dr. Knight and returned to our transect, located behind Leonard Hall in a farming area -we gathered a sample of vegetation, including lettuce, dead leaves, kale, and top soil
2. Observing plant vascularization and specialization -we observed the patterns of vascularization in each of our plant samples, noting the shape of the plant veins and how this might relate to how they obtain water and nutrition -we recorded these observations in our notebooks and made guesses as to how these might affect the plants specialized function and reproduction -we dissected a lily plant to observe its structure and function
3. Setting up a Burlese Funnel -we obtained a conical beaker and put 25 mL of 50% ethanol and 50% water into it -we put a funnel onto the end of the tube and taped it closed -we clamped the funnel we constructed to a stand and placed it under a light -we let this funnel sit for a week so that the invertebraes in our sample would crawl away from the light
Data and Observations Lily Flower: has both the anther and the stigma. The anther acts as the male portion of reproduction and produces haploid pollen cells which then attach to the stigma. The pollen goes down a pollen tube into the ovaries of the plant where it creates a zygote. We concluded that the lily was a monocot. Our transect: we had a lot more trouble identifying the structure of the plants gathered from our transect. The only plant we were really able to observe was the Kale plant.
SS
2.10.15 Good start on notebook. Try using http://excel2wiki.net to make tables to show results. You make the table in excel and then copy and paste it into the website which converts it to OWW format. Pictures would be good too.SK
Lab 3: Identifying Bacteria with DNA Sequences 2/5/15
Purpose: This lab picked up where Lab 2 finished. One week after setting up our 100-fold serial dilution agar plates we observed the bacteria colony growths that appeared at the both the visible and microscopic levels. Through this we practiced identifying and characterizing microorganisms and observed the effects of antibiotics upon these organisms.
Materials and Methods: 1. Quantifying and Observing Microgansims -we retrieved our agar plates (set up last week) and observed the bacterial growth upon them -we counted the number of colonies per plate and converted them to colonies/mL -we compared the number of colonies on the plates containing the tetracycline to those of the agar-only plates
2. Bacteria Cell Morphology Observations -we obtained four glass slides and placed a drop of water on each -using a sterilized metal loop, we scraped samples of the bacteria growth on the 10^5 and 10^7 agar plates and put them onto two of the glass slides -using a sterilized metal loop, we scraped samples of the bacteria growth on the 10^5 and 10^7 agar + tetracycline plates and put them onto two of the glass slides -we observed our four wet mounds under the 10x and 40x objectives, taking care to notice the shapes of the cells and their possible motility -we obtained four new glass slides and labeled them 1-4 -using a sterilized metal loop, we scraped samples of the bacteria growth on the 10^5 and 10^7 agar plates and put them onto two of the glass slides -we passed these glass slides through a flame to fix the bacteria to them -using a sterilized metal loop, we scraped samples of the bacteria growth on the 10^5 and 10^7 agar + tetracycline plates and put them onto two of the glass slides -we passed these glass slides through a flame to fix the bacteria to them -we placed our four slides atop a staining tray -we covered the bacterial slides with crystal violet for one minute then washed it off with water -we covered the bacterial slides again, this time with Gram's iodine mordant -we rinsed this off using a 95% alcohol solution -we covered the slides once more with safranin stain, this time for 30 seconds -we rinsed the safranin stain off with water and dried them using a kimwipe -we put a large drop of oil on the bacteria smears on each side -we observed our oil immersed slides at the 100x objective, taking care to note whether they were gram positive or negative
3. Preparing PCR for Next Week's Lab -we took a sample of the bacteria on the 10^5 agar plate and put it into 100 ul of water -we took a sample of the bacteria on the 10^5 agar + tetracycline plate and put it into 100 ul of water -we put these into a 100*C water bath for 10 minutes -we removed our samples from the water bath and centrifuged them for 5 minutes at 13,400 rpm -we obtained test tubes containing PCR beads -we added 20 ul of a primer and water solution to the PCR bead test tubes -we took 5 ul of the bacteria supernatant out of each of the centrifuged tubes and added them to the PCR bead test tubes -we put these on an ice bath to observe next week
Observations Transect Observations -all the vegetation is now rotten -bottom of the jar is murky and muddy -theres a foul smell, very fungal and mouldy -a dirty ring runs around the jar at the surface of the water -fungal growth along water's surface
Table 1: 100-fold Serial Dilutions Results
Antibiotic Resistance: The plates that contained both the agar and the tetracycline showed significantly less growth than those lacking the tetracycline, proving that it well serves its purpose as an antibiotic. The small amount of bacteria that grew despite it most likely has a resistance to it. According to a report by Ian Chopra and Marilyn Roberts, resistance to tetracycline results from possessing one of 29 tetracycline-resistant genes, some of which code for ribosomal protection proteins and others that code for efflux pumps (Chopra and Roberts).
Table 2: Bacteria Characterization [[File:http://openwetware.org/images/4/47/Screen_Shot_2015-02-05_at_4.34.02_PM.png |thumb|left|alt=.]]
Conclusions and Future Directions This lab effectively demonstrated how certain strains of bacteria can be resistant to certain antibiotics. It was a little unnerving to see that there was still some bacterial growth on the agarose plates containing tetracycline, especially considering that we gathered our bacteria samples from farmland. It just goes to show how real and ubiquitous antibiotic resistance is and that it is a problem we need to address. This lab also gave us firsthand understanding of using gram staining techniques to determine cellular make up and characteristics. We finished up the lab by preparing a PCR and bacteria solution.
Citations
Chopra, I and Marilyn Roberts. (2001, June 1). Tetracycline Antibiotics: Mode of Action, Applications, Molecular Biology, and Epidemiology of Bacterial Resistance. Retrieved February 5, 2015, from http://mmbr.asm.org/content/65/2/232.full#ref-list-1.
SS
1.27.15
Good first entry but is for lab 2 not lab 1. Organization is a little tough to follow. Don't include every labeling event but write about the most important parts. I couldn't see the pictures. We will work on how to upload pictures on Thursday.
Need to write up lab 1 with transect description, 5 abiotic and biotic features and describe Hay infusion set up.
SK
Lab 1: Observing Life in the Niches of AU 1/27/15
Purpose: This lab held two purposes. The first was to observe how natural selection acts upon organisms through the examination of several different bacterial cells under a microscope. The second was to construct a Hay Infusion Culture out of a niche found at AU.
Materials and Methods: 1. Observing Evolutionary Specialization -obtain three pre-made slides containing Chlamydonas, Gonium, and Volvox -observe each slide beneath the microscope and record each organism's cell number, colony size, motility, any specialization, and reproduction characteristics
2. Creating a Hay Infusion from a Niche at AU' -go to the location of group's assigned niche -observe the area and draw a picture of what it contains -take a sample of the soil and vegetation in a 50 mL tube -return to lab -obtain a jar with 500 mL of water -add 10g of the sample into the jar -put 0.1gm of dried milk into the jar, mix -label jar and leave it on the back counter of the lab
Date and Observations Table 1. Evolutionary Specialization of Members of Volvocine Line File:Http://openwetware.org/images/6/6b/Screen Shot 2015-01-29 at 12.56.22 PM.png
Transect Observations: 20x20 ft area of farmland on AU Campus' north side. It appears to have rained recently, as everything is kind of dewey and wet. There are four raised garden planters containing soil and a variety of vegetation. Around these four structures the ground is more firm and strewn with woodchips. The area of our transect is enclosed by a chain link fence Abiotic features: vegetable beds, metal fence, woodchips, wire, soil Biotic features: worms, brussel sprouts, broccoli, ants, lettuce, moss
Conclusions and Future Directions Observing the Chlamydomonas, Gonium, and Volvox samples was a good way of illustrating how evolution creates diversity amongst organisms. While the three organisms we examined share a common ancestor, evolution is responsible for their differences in reproduction, size, and motility. Now that we have made our Hay Infusion Culture, we can examine it in next week's lab and observe what bacteria and life grows in it.
SS
2.4.15 Good entry. Each new lab book entry should be posted at the top of the page, pushing down the older entries. Try not to just list things from the manual but to include what you actually observed. eg. don't tell me you observed your Hay Infusion, tell me what the Hay Infusion looked like. The picture links do not work properly. SK
Lab 2: Identifying Algae and Protists 1/25/15
Purpose The intended purpose of this lab was to familiarize ourselves with observing eukaryotes under a microscope and to identify key features of such microbes, then to match these key features to those listed by a dichotomous key. The eukaryotes we examined were sampled from the transects we collected last week in lab.
Material and Methods
This lab had three different components
1. Using a Dichtomous Key
-we made a wet mound of the organism gonium then place it under the microscope
-we observed the gonium slide at the 4x and 10x objectives
-we noted the organism's ocular size and defining characteristics in our notebooks
-using a dichotomous key, we compared our observed characteristics to those listed by the key
-we successfully identified the gonium sample
2. Observing our Hay Infusion Culture
-we obtained the hay infusion culture that we made last week and examined its appearance and smell
-we took samples from two different niches in our culture- one from the floating gel-like blob, one from the water surrounding the culture's vegetation
-we made two distinct wet mounds using these samples
-we observed our prepared slides, noting key features of the organisms we observed
-we compared our observed organisms to those listed on the dichotomous key and identified them
3. Preparing a serial dilution
(this part of the lab was mainly a set up for next week's lab)
-we labeled three test tubes containing 10 mL of broth each: 10^4, 10^6, 10^8
-we labeled three petri dishes containing agar: 10^5, 10^7, 10^9
-we labeled three petri dishes containing agar and tetracycline: 10^5, 10^7, 10^9
-using a micropipette we took 10 mL of the hay infusion culture and put it into the test tube labeled 10^4
-we took 100 mL of this new culture-broth mix and put it into the 10^6 test tube
-we took 100 mL of the 10^6 culture-broth and put it into the 10^8 test tube
-we pipetted 100 mL of the mixture in the 10^4 test tube onto the agar plate labeled 10^5 and onto the agar + tetracycline plate labeled 10^5
-we pipetted 100 mL of the mixture in the 10^6 test tube onto the agar plate labeled 10^7 and onto the agar + tetracyclne plate labeled 10^7
-we pipetted 100 mL of the mixture in the 10^8 test tube onto the agar plate labeled 10^9 and onto the agar + tetracycline plate labeled 10^9
-we left these agar and agar + tetracycline plates to incubate at room temperature for one week
Data and Observations
1. Gonium Wet Mound sample
4 ocular spaces large//observed at 40x, so 10uM
File:Http://openwetware.org/wiki/Image:Unnamed-1.jpg File:Http://openwetware.org/wiki/Image:Unnamed-2.jpg
2. Hay Infusion Observations
-smell: fungal, salt-watery
- gel-like growth floating in surface of the water
-apparent bacterial colonies floating at surface
-murky scum on sides of jar
File:Http://openwetware.org/wiki/Image:Unnamed-3.jpg File:Http://openwetware.org/wiki/Image:Unnamed-4.jpg
Niche 1: collected from Gel
File:Http://openwetware.org/wiki/Image:Unnamed-5.jpg
We observed two different specimen in this sample. Using the dichotomous key, we identified the long, stringy, amber colored organisms as being Spirostomum. Spirostomum is a ciliophora and a eukaryote. This was due in part to the organism's texture, color, shape, and size- 1-3 um in length. The second type of organism, the colorless, bubble-like circles, we identified as being Chlamydomonas.
Niche 2: collected from vegetation File:Http://openwetware.org/wiki/Image:Unnamed-6.jpg
The two organisms we observed in this wet mound were Colpidium, which we measured to be from 50-70 um, and didinium cysts. File:Http://openwetware.org/wiki/Image:Unnamed-7.jpg File:Http://openwetware.org/wiki/Image:Unnamed-8.jpg
3. Serial Dilution Procedure File:Http://openwetware.org/wiki/Image:Unnamed9.jpg
Conclusions and Future Directions
This lab showed us just how useful dichotomous keys can be when identifying an unknown eukaryote. The simple descriptions and detailed images of each organism we might have found made it very easy to identify the organims we found in our hay infusion culture. If I were to repeat this experiment again, I would probably observe organisms from more niches of our culture to see how the organisms differed from niche to niche.
SS