User:Cara Chao/Notebook/Biology 210 at AU
February 25, 2016
Zebrafish Experiment: Testing the Effects of Metoprolol on Embryonic and Larval Development in Zebrafish
Introduction -
In this experiment, the effects of Metoprolol, a beta-blocker, on embryonic development in zebrafish were observed over the course of two weeks. Metoprolol is a drug used to treat high blood pressure, and is often prescribed to those who have suffered from heart attacks. It functions by slowing heart rates and relaxing blood vessels to lower blood pressure and improve blood flow in the body. The purpose of this experiment was to assess the effects of metoprolol on the development of zebrafish, a freshwater fish, in order to gauge the overall effects of metoprolol waste in freshwater marine life. Since metoprolol is a drug that is used to treat high blood pressure, the experimental fish that were exposed to the metoprolol treatment were expected to be smaller in size, due to reduced blood flow to developing body parts, and have slower average heart rates.
Link to fish data-sheet: https://docs.google.com/spreadsheets/d/1kMbv2FQr0OZ31zZ27pzXvfFnVU-tRuz8eJ3qNoz2stQ/edit?usp=sharing
Images of Control and Experimental fish can be found: https://drive.google.com/folderview?id=0B302eamWQF5nVFM0VVpYRDhBVWM&usp=sharing
February 25, 2016
Lab 6: Analyzing the 16S Sequences from the Bacterial Colonies
Introduction -
In this experiment, the results from the the PCR DNA sequencing of the bacterial colonies from Lab 3 were analyzed. With the results sequence results, the identities of the colonies were confirmed. From previous analysis of these colonies, both colonies were found on nutrient agar plates without tetracycline -- which indicated that they did not have tetracycline resistance genes -- and they both came out to be gram negative from the gram stain -- which showed that they did not have a thick layer of peptidoglycan on their cell walls. The 16S sequence is found in all prokaryotes and slight differences within this sequence helped to identify the tan and green bacterial colonies from lab 3.
Materials and Methods -
The returned bacteria DNA sequences were analyzed using the nucleotide BLAST program from the National Center for Biotechnology Information. The program used the sequences and found their closest matches in their database, which allowed for the identification of the bacteria from the list of the closest DNA matches.
Data and Observations -

This is the gel that came out from the PCR of the bacterial colonies. The 16S sequence was successfully cut and amplified, which can be seen boxed in the image.

This image shows the sequence results of the tan colony that was found on nutrient agar plates -tet with 10^-3 dilution. This colony was identified as Pseudomonas putida -- which is a rod-shaped, flagellated, bacterium that stains gram negative.

This image shows the sequence results of the green colony that was found on nutrient agar plates -tet with 10^-7 dilution. This colony was identified as Pseudomonas -- which is a rod-shaped, motile, bacteria that stains gram negative.
Discussion -
The sequence analysis results revealed that the green and tan bacterial colonies were Pseudomonas bacteria -- common bacterium that typically dwell in moist soil or water environments -- which is a very reasonable result for transect 4. They are both strictly aerobic and therefore can only be found in places with oxygen. They were both identified as bacillus, or rod-shaped, motile, and gram negative bacterium. These results support the previous observations made of these colonies (refer back to Table 2 in Lab 3).
February 11, 2016
Lab 5: Characterizing and Identifying Invertebrates and Vertebrates Collected from Transect 4
Introduction: In this experiment, vertebrates from Transect 4 were collected through a Berlese Funnel and were examined and identified under a microscope using a dichotomous key. A Berlese Funnel is a funnel with a net/filter on the bottom of the funnel tube that is filled with dead leaves and plant matter. Attached to the bottom of the funnel tube is a sterile 50mL conical tube that is filled with an ethanol/water solution. The funnel setup is placed beneath a light, which will force any critter that is dwelling in the leaf litter to migrate down, away from the light and heat, and into the ethanol solution, killing and preserving them for analysis. The filter serves to catch any large particles from falling into the conical tube. It was expected that all of of the organisms found, if any, would be vertebrates with hard exoskeletons because of the weather conditions as well as the general consensus that no large organisms could fit within a small sample of detritus.
Materials and Methods -
- After a week of waiting, the Berlese funnel was taken apart and the contents of the conical tube were removed -- the top half of the solution was poured into one dish while the remainder of the contents were poured into another dish.
- The dishes were observed under a dissection scope, and organisms that fell into the ethanol solution were analyzed and characterized using a dichotomous key.
Data and Observations -


Of the organisms that were found in the leave litter sample (figure 1), all of them were vertebrates with exoskeletons and were very small with sizes ranging from 1mm to 9mm. The largest specimen that was identified was an arthropod, specimen 4.1, that was approximately 9mm. These results show only a small portion of the organisms that reside in this transect, and it does not take into account the more complex organisms that also live there. In addition, all of the specimens found had bilateral symmetry. The two samples from the Berlese Funnel were different. All of the organisms were found in the bottom half of the solution, settled with soil particulates.
Discussion -
Invertebrates from Transect 4 were collected through a Berlese Funnel and were characterized with a dichotomous key as a means to get a general idea of what the invertebrate population on the floor surface of the transect. The results from this experiment support the original hypothesis of finding small organisms with hard exoskeletons because all of the identified organisms had exoskeletons.
Other vertebrates that could inhabit this transect include sparrows, pigeons, koi fish, squirrels, rats, as well as chipmunks. The classification of the vertebrates are as follows:
Sparrow - Phylum: Chordata, Class: Aves, Order: Passeriformes, Family: Passeridae, Species: Saxaul sparrow
Pigeon - Phylum: Chordata, Class: Aves, Order: Columbiformes, Family: Columbidae, Genus: Ectopistes, Species: Ectopistes migratorius
Koi Fish - Phylum: Chordata, Class: Actinopterygii, Order: Cypriniformores, Family: Cyprinidae, Genus: Cyprinus, Species: C. carpio
Squirrel - Phylum: Chordata, Class: Mammalia, Order: Rodentia, Family: Sciuridae, Genus: Sciurus, Species: Sciurus carolinensis
Rat - Phylum: Chordata, Class: Mammalia, Order: Rodentia, Family: Muridae, Genus: Rattus, Species: Rattus norvegicus
Chipmunk - Phylum: Chordata, Class: Mammalia: Order: Rodetia, Family: Sciuridae, Genus: Tamias, Species: Tamias striatus
Biotic factors that may benefit these organisms include plenty of foliage that serve as protection and shelter as well as the presence of humans, which could provide them with nutrients (ie. food) and protection from other predators that shy away from human contact. Abiotic factors that my benefit these organisms include the presence of a pond, which allows the survival of the koi fish, and the American University campus. The campus provides them with protection from natural and some urban disruptions, unnatural as it may be.
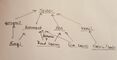
This food web depicts the relationships between as well as the trophic levels of the analyzed organisms from transect 4. It is apparent that almost all of the analyzed organisms, eukaryotic and prokaryotic, are at the lower trophic levels. Organisms at the higher trophic levels were not observed throughout this experiment, but it is safe to assume that these observed microorganisms serve as valuable resources to these organisms at the higher levels of the food web, therefore they play an important role in the carrying capacity of organisms that are higher up in the food web.
February 4, 2016
Lab 4: Identifying and Studying Plants and Fungi Collected from Transect 4
Abstract: In this section of the Hay Infusion experiment, plants and fungi samples from transect 4 are collected and analyzed. In addition, leaf litter from the transect are gathered for the setup of a Berlese funnel -- which will be used to isolate prospective invertebrates that dwell in the transect.
Plant Characterization Chart:

Images of Characterized Plants:
January 28, 2016
Lab 3: Identifying and Studying Bacteria
Abstract: In this experiment, the Hay infusion culture is observed a final time and the results from the previous bacterial culture on nutrient and nutrient plus tetracycline agar plates are analyzed. From the colonies found on the plates, 4 are chosen and are gram stained. A gram stain is a method in which microorganisms can be differentiated based on their staining characteristics. Bacteria that come out as gram (+) have a a thick layer of peptidoglycan in their cell walls, which allows them to retain the crystal violet dye (purple color) and appear purple under the microscope. On the other hand, bacteria that come out as gram (-) lack the thick layer of peptidoglycan in their cell walls, and therefore do not retain the purple color and appear pink under the microscope. This gram stain is used to help characterize the colonies grown on the tet- and tet+ plates. After the gram stain slides are observed and the chosen colonies are characterized, two colonies are picked out for PCR. This PCR is for the amplification of the 16S gene, which is a section of prokaryotic DNA that is found in all bacteria and archaea and is commonly used to characterize bacteria. The results of this experiment show that fewer colonies grew on tet+ agar plates; however, this indicates that the presence of a tetracycline resistance gene may be present among the colonies, if they are bacteria, on the tet+ plate.
Observations of Hay Infusion Culture: Transect 4, after another week of incubation, has retained its foul moldy odor. There is a thin film of mold covering the surface the water, and the layer of detritus at the base has gotten thicker, which could indicate signs of decomposition. The water has become more clear and there is a white and cloudy layer that has accumulated along the base above the brown detritus. In addition, there is a dry rim that is less than an inch thick above the surface of the water indicating that water has evaporated. As expected, Archaea organisms were not present in this hay infusion; it would have been an anomaly if archaea were found because they generally dwell in extreme environments, and transect 4 cannot be considered as one that is extreme.

Table 1: 100-fold Serial Dilutions Results
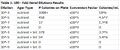
The results show that there was a drastic decrease in the overall number of colonies that grew on tet+ plates compared to those on tet- plates. Plates without tetracycline contained colonies that were not able to grow on tet+ plates. Small light green colonies were identified on tet- plates that could not be found on tet+ plates. However, there were colonies that grew on both the tet+ and tet- plates. Both agar types contained tan, white, and orange colonies. Tet+ plates also had fungal growth in the form of fuzzy white colonies, which indicates that there was either fungal contamination or tet+ plates reduced competition and provided room for any fungi in the sample to grow. There are currently twenty-nine different tetracycline resistance genes, tet (1). If the colonies on tet+ plates, other than those that are fungi, are confirmed to be bacterial colonies, then there is a chance that they have the tet gene which would explain why they were able to grow on tet+ agar.
Mechanisms of Action for tetracycline:
Tetracycline is an antibiotic that functions by inhibiting bacterial protein synthesis. It does so by preventing the association of aminoacyl-tRNA with bacterial ribosome (NCBI). Tetracycline affects bacteria that are gram-negative (NCBI), which indicates that they do not have a thick layer of peptidoglycan in their cell walls.
Colonies Analyzed:
- Tan colony from the 10^-7 -tetracycline agar plate
- Light Green colony from 10^-3 -tetracycline agar plate
- Orange colony from 10^-5 +tetracycline agar plate
- White & fuzzy colony from 10^-5 +tetracycline agar plate
Table 2: Bacteria Characterization


Colonies were observed under a dissection scope.
Gram Stain:
Materials and Methods
- A metal loop was sterilized over a flame and a small amount of growth from the chosen colonies was scraped off. A drop of distilled water was added onto a clean slide and the loop was mixed in the water and the slide was heat fixed with a flame.
- After it dried, the slide was covered with Crystal violet for a minute.
- The Crystal violet was rinsed off with distilled water and covered with Gram's Iodine mordant for another minute.
- The Iodine was rinsed off with distilled water and decolorized by covering it with 95% alcohol for 10-20 seconds, and rinsed again with water
- The smear was covered with safranin stain for 20-30 seconds and rinsed one last time with distilled water and then it was dried.
- The slide was then observed under a microscope to determine if the growth is gram (-) or gram (+).
Gram Stain Images:
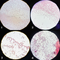
Magnification:
Colony 1: 1000x
Colony 2: 400x
Colony 3: 1000x
Colony 4: 1000x
PCR for 16S Gene:
Materials and Methods -
- 20µL of the primer/water mixture was added into each PCR tube and mixed to dissolve the PCR pellet
- A sterile toothpick was used to retrieve a small amount of the colony (1 for each tube) and was mixed into the PCR tube
- The tubes were then put into the PCR machine.
PCR Results - Gel Electrophoresis on agarose gel

Conclusion - The results of the agar plates show that there was bacterial growth on both tet+ and tet- plates, however, tet+ plates made room for fungal growth due to the presence of the antibiotic, tetracycline. To further characterize four select colonies from these plates, the PCR samples were sent to a lab to get their DNA sequenced. Plants and fungi that are present in the transect will be collected and observed to get a sense of some of the eukaryotes that reside in transect 4.
Citations:
1. Chopra, Ian and Marilyn Roberts. "Microbiology and Molecular Biology Reviews". Tetracycline Antibiotics: Mode of Action, Applications, Molecular Biology, and Epidemiology of Bacterial Resistance. Jun. 2001. National Center for Biotechnology Information. (9 Feb. 2016).
January 20, 2016
Lab 2: Identifying Algae and Protists
Abstract: From the previous lab, samples of transect 3 were obtained and cultured in a powdered milk and water medium, or a Hay Infusion Culture. The culture was set up using 500mLs of water, 0.1 grams of dried milk, and 10 to 12 grams of samples obtained from transect 3 mixed together in a plastic jar. After a week of incubation in room temperature, the culture was analyzed and samples from the top and bottom layers were extracted and observed. The sample obtained from the top was extracted from a layer of mold with no plant matter around, while the sample from the bottom was extracted from an area around plant matter. Using a microscope, the extracted samples were analyzed and organisms were identified using a Dichotomous key. A culture with agar plates and agar plus tetracycline plates was set up through a serial dilution of 100µL of the Hay Infusion, after it was thoroughly mixed, in 10 mLs of sterile broth.
Since tetracycline is an antibiotic, bacterial cultures will not grow on the agar plates with tetracycline unless. If there is any growth on tetracycline plates, the growth would most likely be fungus or strains of bacteria that contain a tetracycline resistant gene.
Observations of the culture: - Smelled of decomposition; stagnant water from a creek or lake; earthy-dirt scent; smelled moldy.
- Appearance: fluid in the container was murky light-brown color that was still translucent. There was a layer of mold on the top layer of water with a white film-like layer below. On the bottom of the container was an accumulation of settled particles, most likely soil and rocks. The middle area of the medium contained growth in the form of green shoots from plant samples that were collected and roots.


Identified Organisms:
(1) Gonium (40x), Length: 10.0 µm Width: 1.88 µm Small, rice-shaped body that is composed of a chain of smaller green bodies. Motile and ranges in length. Photosynthesizing and motile.
(2) Eudorina (40x), 10.0 µm Small, round body that contains smaller colonies inside what looks like a membrane. Motile and photosynthesizing algae.
(3) Pandorina (40x), 37.5 µm Large, round body that contains multiple colonies inside. Motile and photosynthesizing algae.
Image of Identified Protists:

Serial Dilution:
Materials -
- Four tubes of 10mL sterile broth labeled 10^-2, 10^-4, 10^-6, & 10^-8
- P1000 micropippetor with sterile tips
- 4 nutrient agar plates & 4 nutrient agar plus tetracycline plates
- 100µL of Hay Infusion Culture
- sterile rod
Procedure -
4 tubes of sterile broth were labeled according to their dilution factor; the 8 agar plates (both -tet and +tet) were also labeled according to their dilution (10^-3,1 10^-5, 10^-7, & 10^-9). The Hay Infusion Culture (transect 3) from the previous lab was mixed; 100µL of the mixed culture was extracted using a P1000 and added to the sterile broth tube labeled 10^-2. From this tube, a serial dilution was carried out to 10^-8 dilution. 100µL from each diluted broth tube was removed and added to the corresponding agar plates. The sterile rod was then used to spread the mixture around the agar plates; the rod was sterilized with ethanol and fire.

The Eudorina meets all the needs of life, as described in Biological Science, by Freeman, Quillin, and Allison by having cells, as it is a colony of green algae, acquiring and using energy through photosynthesis and respiration, containing genetic material, replicating through asexual reproduction, and having the ability to evolve as a species.
If the Hay Infusion Culture grew for another two months, one possible outcome would be an increase in the population of the microorganisms. Any physical changes to the culture would most likely be an increase in mold. Selective pressures that may influence populations in the culture may be a decrease in resources (ie. food), which would likely lead to an overall increase in the populations of organisms that can photosynthesize and a drastic decrease in the populations of heterotrophs.
January 13, 2016
Lab 1: Examining a Transect at American University (AU)
Transect: 3
Location: Transect 3 is a 20x20 meter plot located in the garden behind the AU amphitheater and next to Hughes Hall. It is a quiet area with an abundance of foliage and shade. There is no body of water around the transect. On the day that this transect was analyzed the weather was sunny with clear skies with temperatures around 30 degrees Fahrenheit.
Topography: The transect is surrounded by cement side-walks and stone, and contains a light post and a wooden bench. It also contains bushes, trees, detritus, wood chips and dormant plants.
Biotic Factors: bacteria and other potential living organisms in the soil, leaves, stems, and roots. Abiotic Factors: soil, metal, water, and rocks.

Hay Infusion Culture:
Materials:
- 10-12 grams of transect sample
- 500mLs of deerpark water
- 0.1g of dried milk
- clean plastic jar with lid
A Hay Infusion Culture was set up using the samples collected of the transect. 10 to 12 grams of the collected samples were put into a plastic jar with 500mLs of water, along with 0.1 grams of dried milk. The jar was closed and mix for 10 seconds, then was opened up and left to incubate for a week.