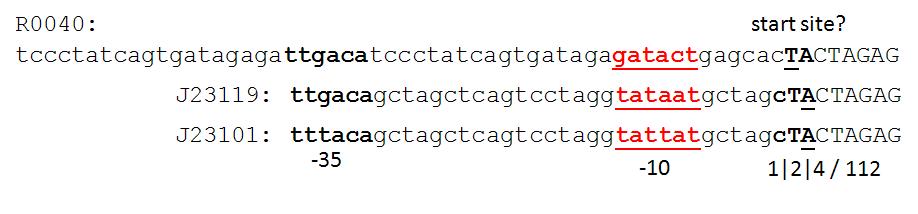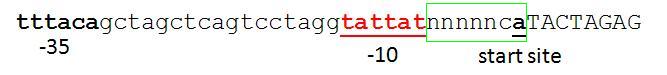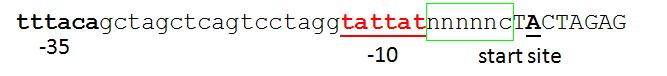User:Bill Flanagan/Sandbox
From OpenWetWare
Jump to navigationJump to search
My Images
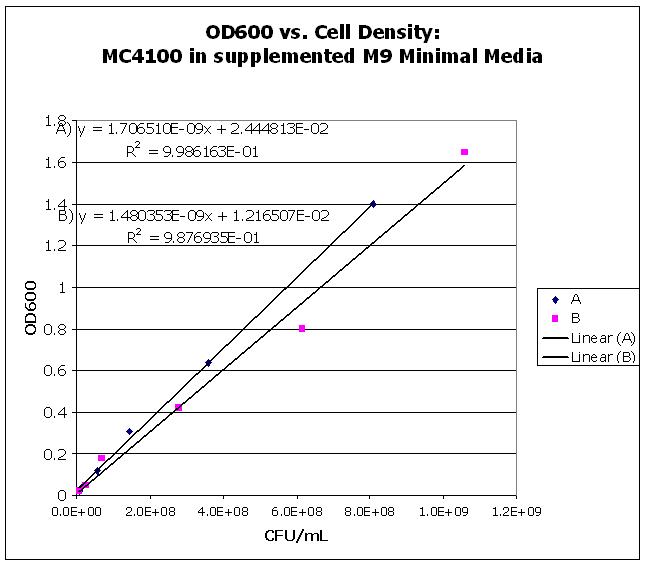
| OD600vsCellDensity.JPG: | OD600 vs. CFU density | |
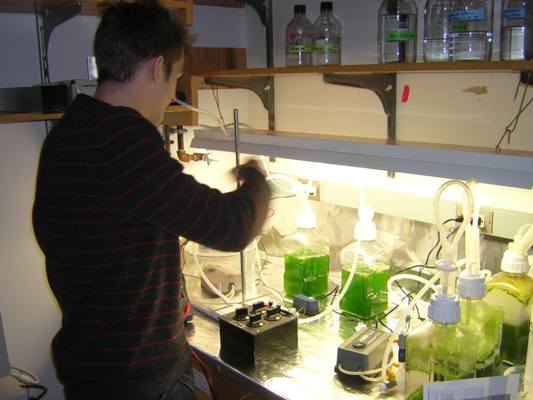
| 2005.4.5-2.jpg: | Closeup of pump with cells | |
| Headbg.jpg: | banner | |
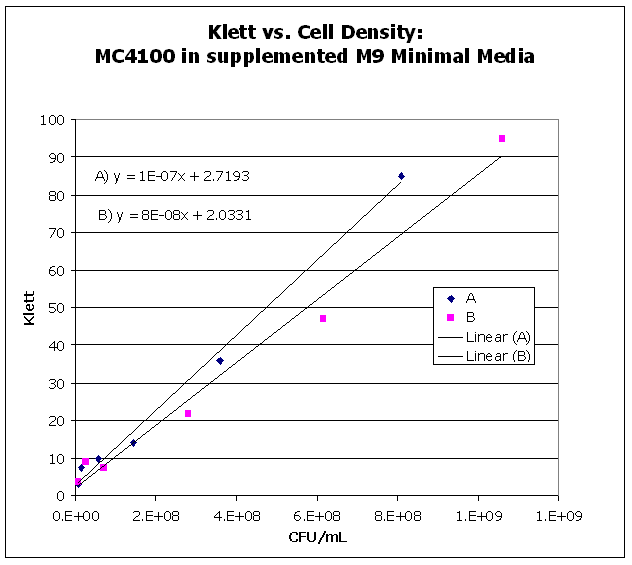
| KlettvsCellDensity.png: | Calibration Curve for Klett vs. Cell Density | |
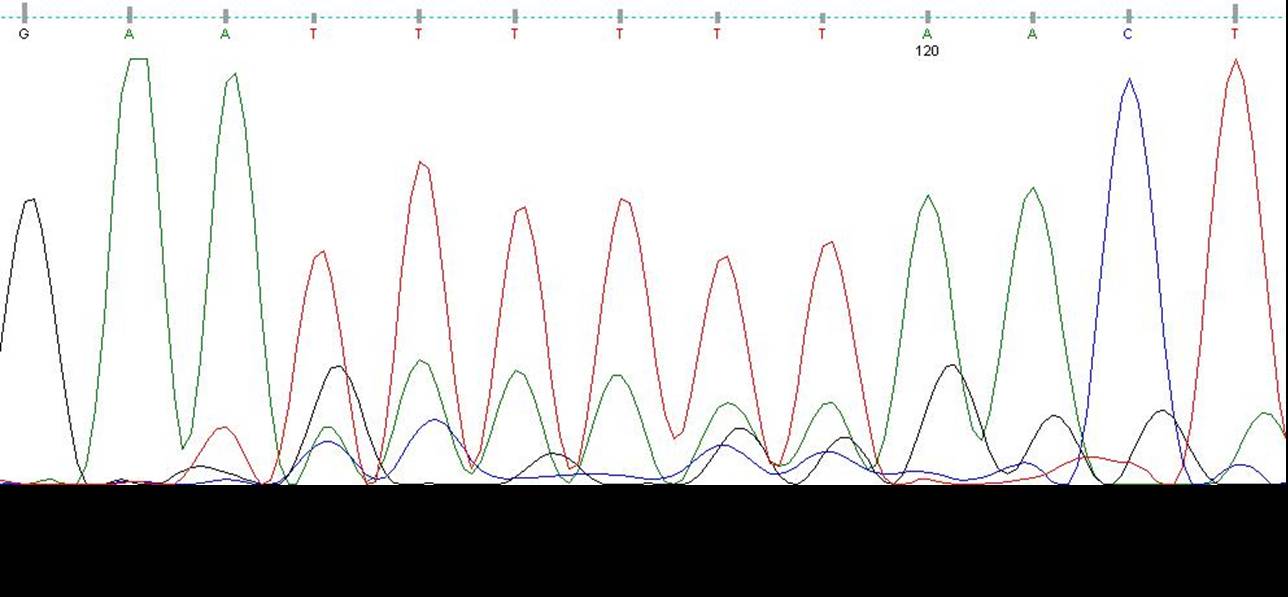
| Seq1.jpg: | seq of library | |
| Spec calibration.xls: | test of excel upload | |
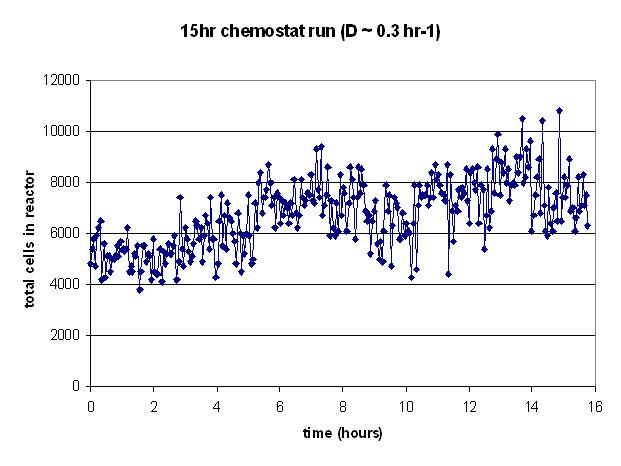
| SSchemo.PNG: | early chemostat results | |
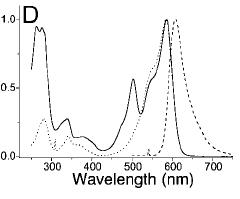
| MRFP1.JPG: | mRFP1 spectra | |
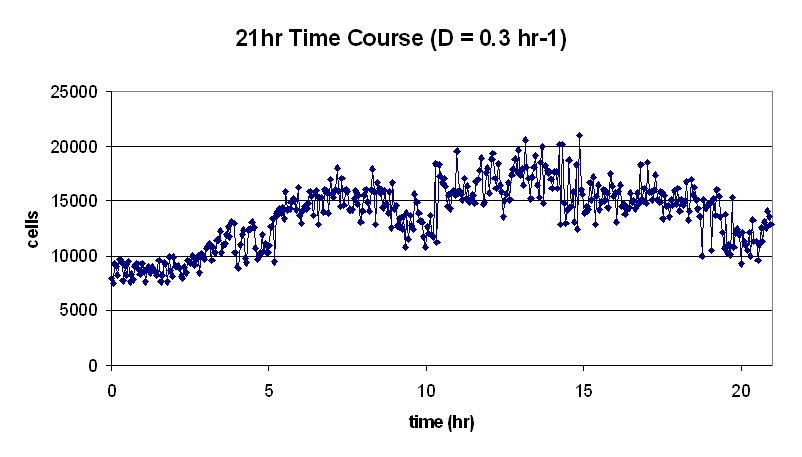
| 2005.06.07.ChemostatRun.JPG: | 06.07.2005 Chemostat Run | |
| 050607Chemo12hr.PNG: | 06.07.2005 Chemostat Run | |
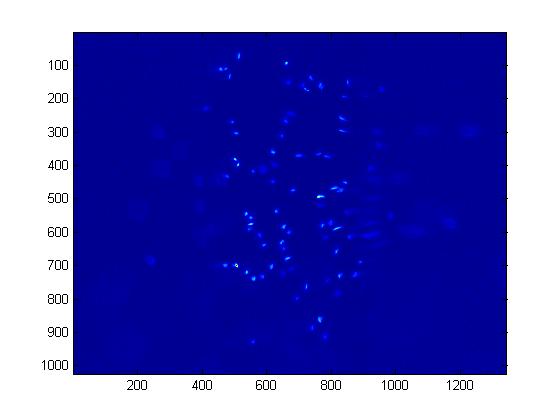
| ChemoIP1 orig.jpg: | chemostat image process original | |
| ChemoIP1 orig bright.jpg: | Chemostat IP orignal brightened | |
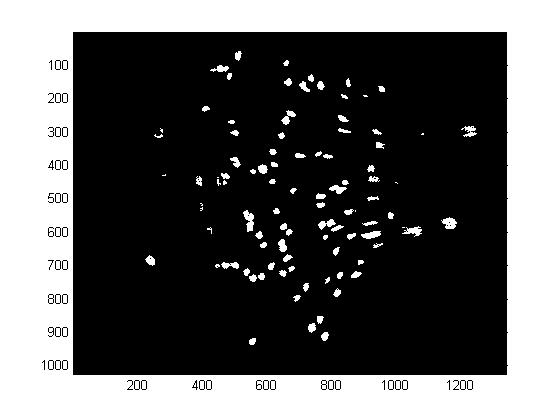
| ChemoIP1 process.jpg: | chemostat image process processed | |
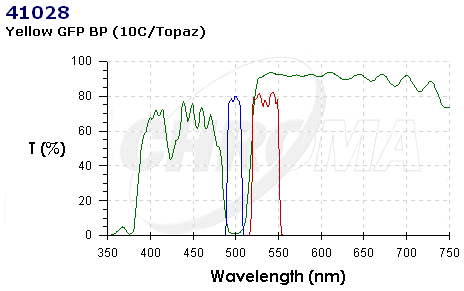
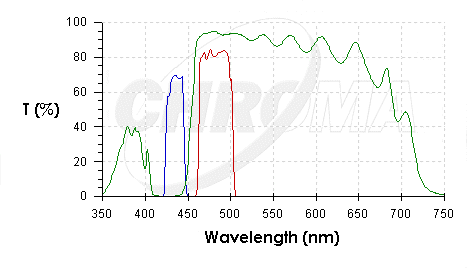
| CHR41028.PNG: | 41028 Chroma Spectra | |
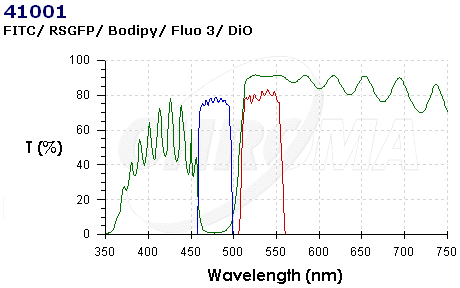
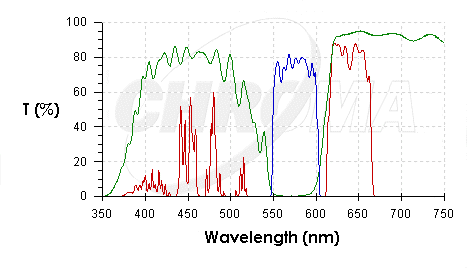
| CHR41001.PNG: | Chroma spectra 41001 | |
| CHR31044v2.gif: | Chroma spectra 31044v2 | |
| CHR41043.gif: | Chroma spectra 41043 | |
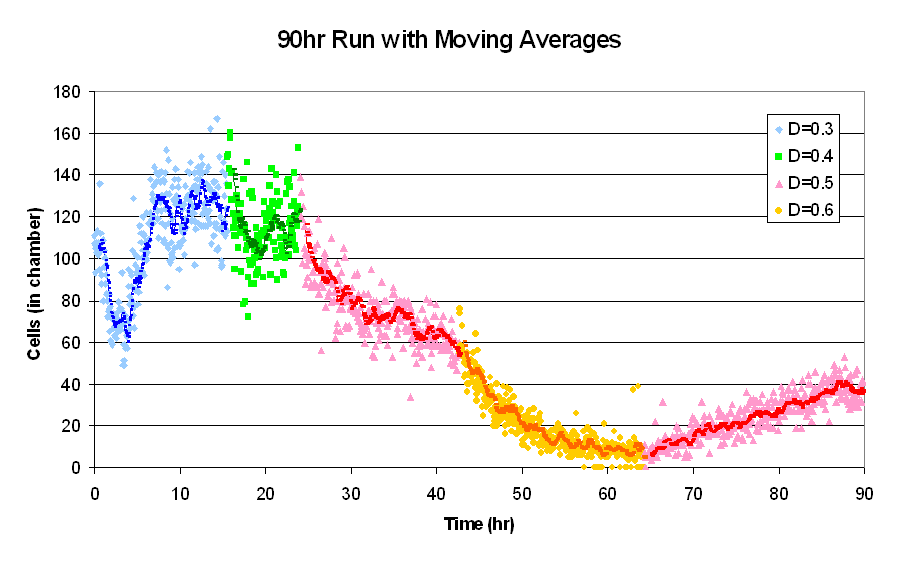
| 90hrRunf.PNG: | 90hr chemostat run | |
| BEING-leaf-logo.jpg: | Logo for BEInG front page | |
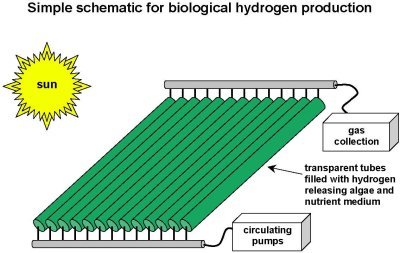
| Bioreactor-schematic.jpg: | Schematic of a hydrogen photobioreactore (BEInG) | |
| Luciferyeast.JPG: | luciferyeast | |
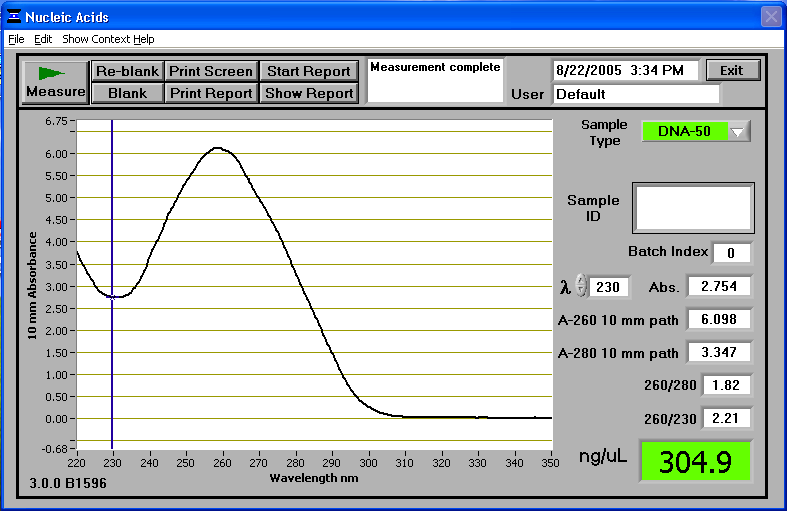
| DNANanodropExample1.PNG: | Example of wavelength vs. absorbance curve for a miniprep sample on the nanodrop. | |
| Anthropology.PNG: | anthropolgy home page pic | |
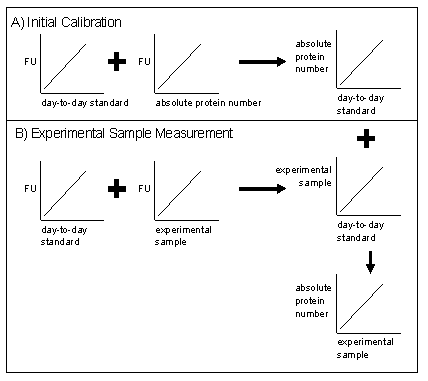
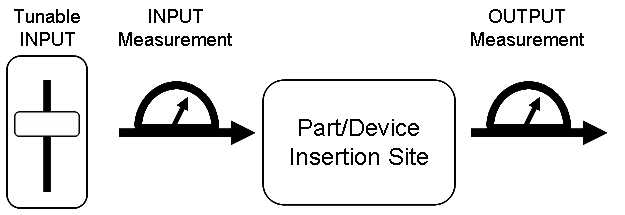
| GFPstandardProcess.PNG: | process for GFP measurement standardization work | |
| Streak plates.png: | Agar plate being streaked with bacteria. Drawn by Theresa Knott. thanks wikipedia! | |
| BEBoard.JPG: | BE Board Logo | |
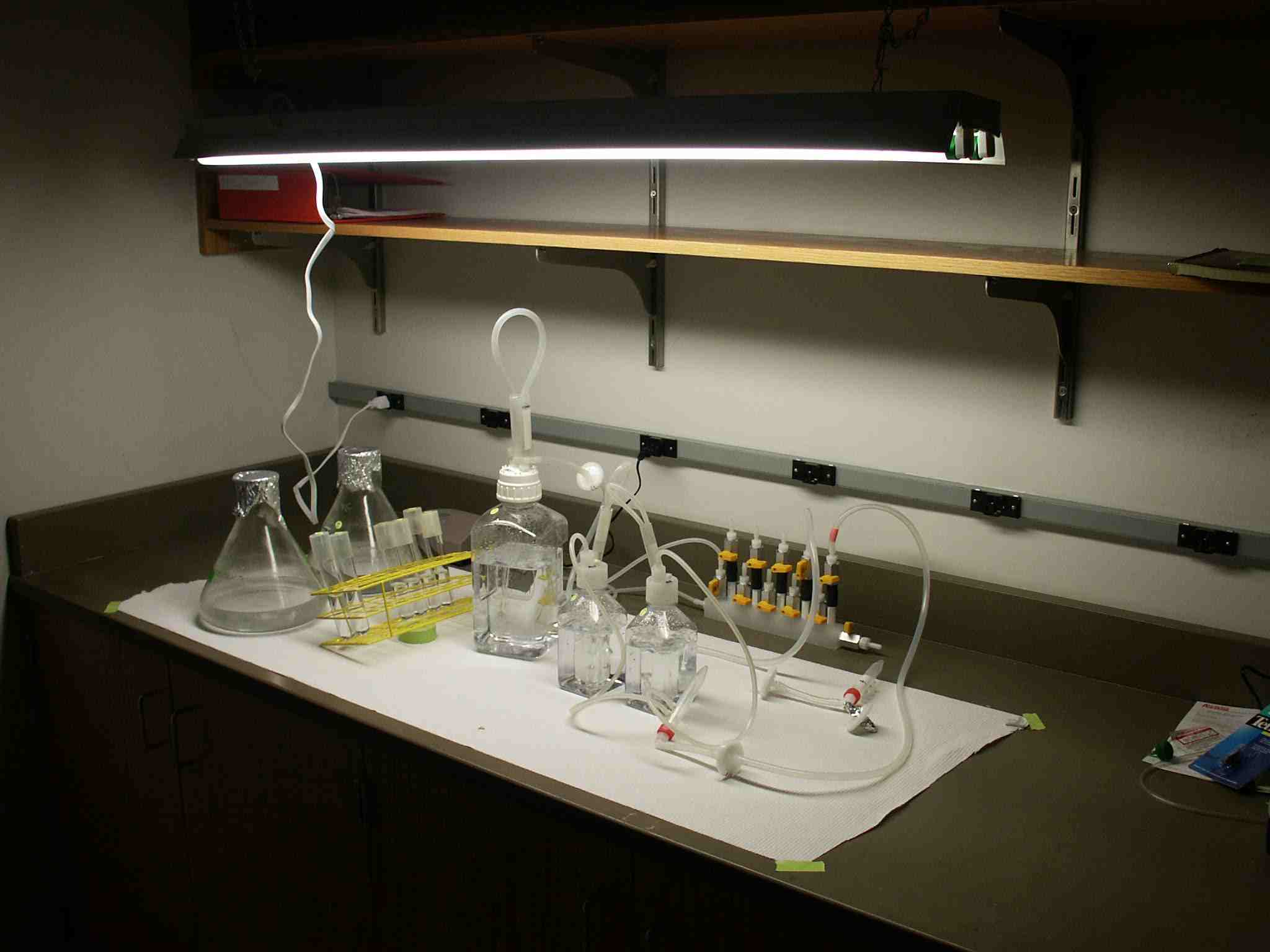
| BEInG lab.jpg: | Setup for BEInG Hydrogen Bioreactor 9/26/2005 | |
| Changing Sensor Zero Values.pdf: | document for changing sensor zero values on solent incubator. | |
| JK ICSB 2005.pdf: | Jason Kelly ICSB 2005 Poster | |
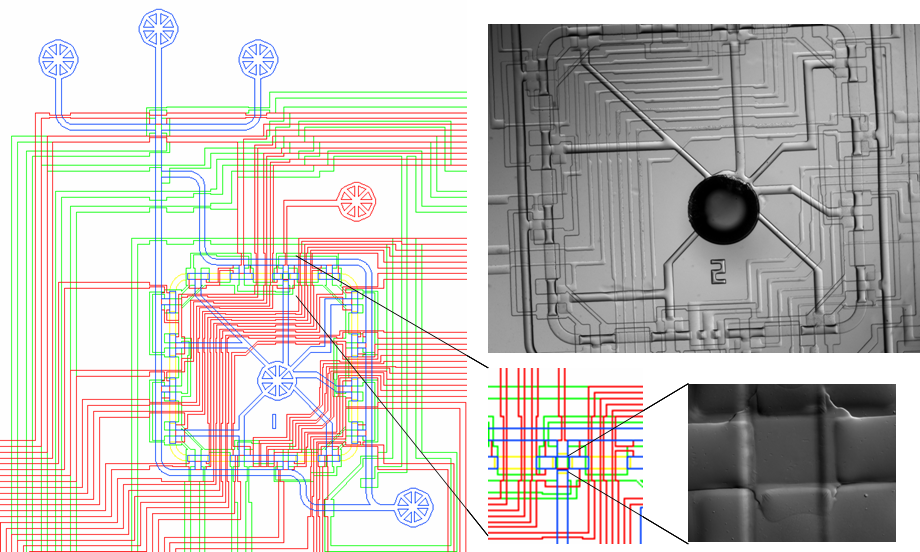
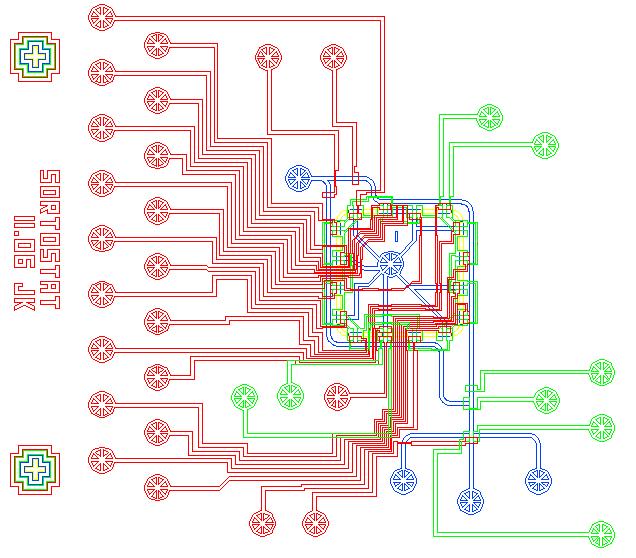
| SortoStatDesign.PNG: | Sortostat design image | |
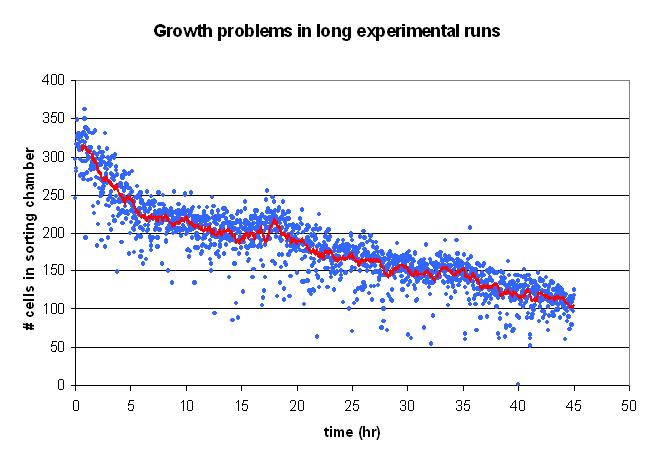
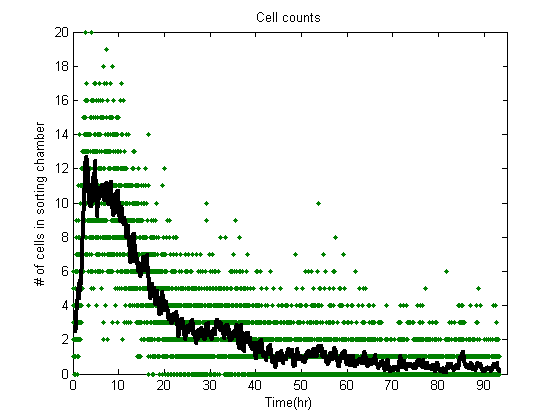
| GrowthIssues.JPG: | Graph of cells dying out insortostat in long runs. | |
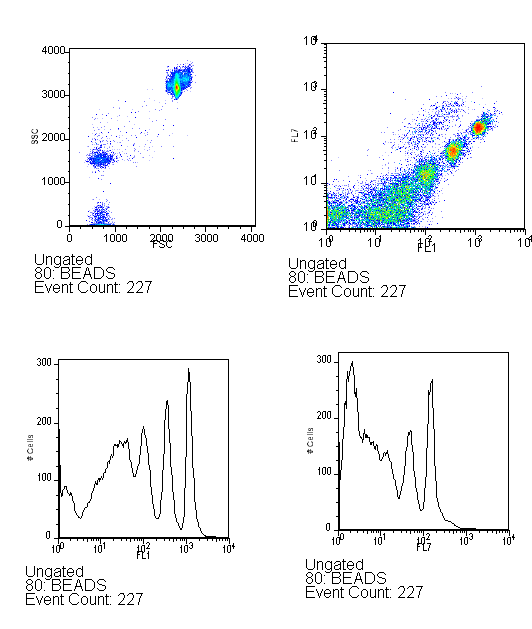
| MOFLObeads.PNG: | Example of what Spherotech RCP-60-5 Rainbow Calibration Particles (6.0-6.4um) should look like on the MOFLO under good alignment. | |
| TestImage.PNG: | test Image | |
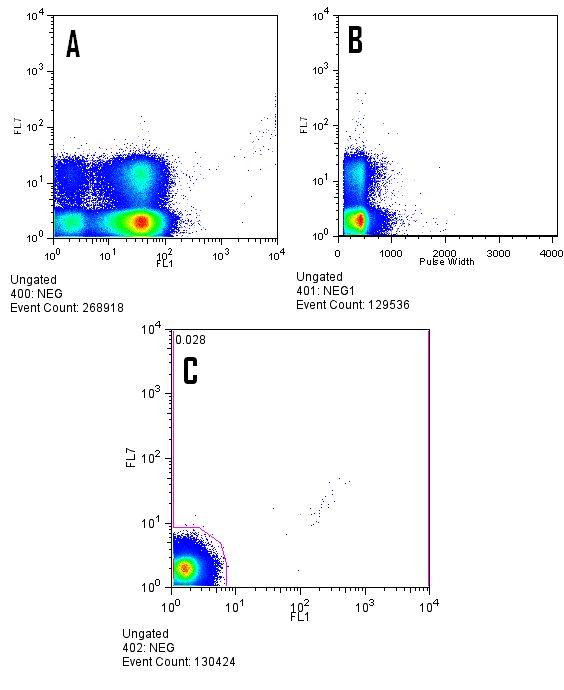
| MOFLODarkNoise.jpg: | Example of "dark noise" on non-flourescent cells on the MOFLO as a result of high PMT settings. (A) FL1=max / FL7=max (B) FL1 = 525 / FL7=max (C) FL1=525 / FL7 = 650. | |
| UT HelloWorld.jpg: | UT austin biofilm from wikipedia | |
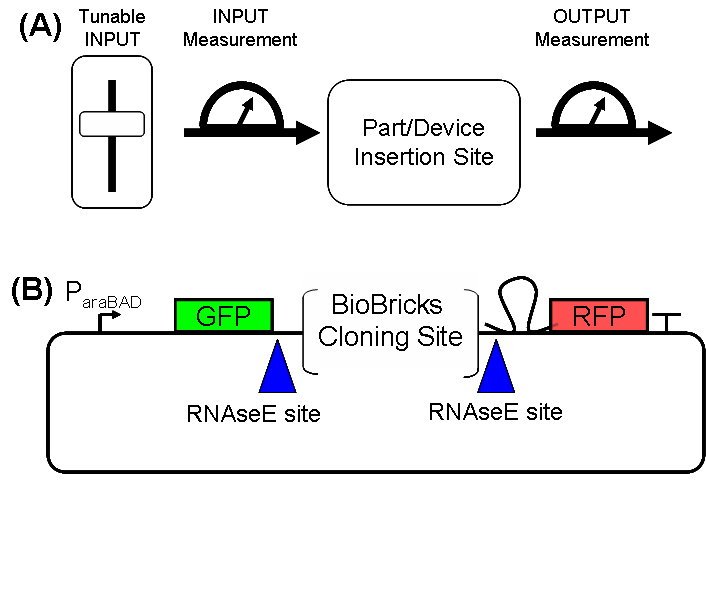
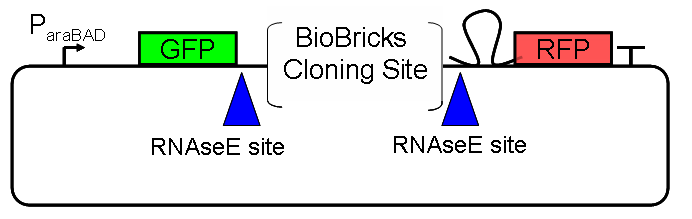
| ScreeningPlasmidDesign.PNG: | Schematic and actual design for the screening plasmid | |
| ScreeningPlasmidSchematic.PNG: | Schematic of Screening Plasmid | |
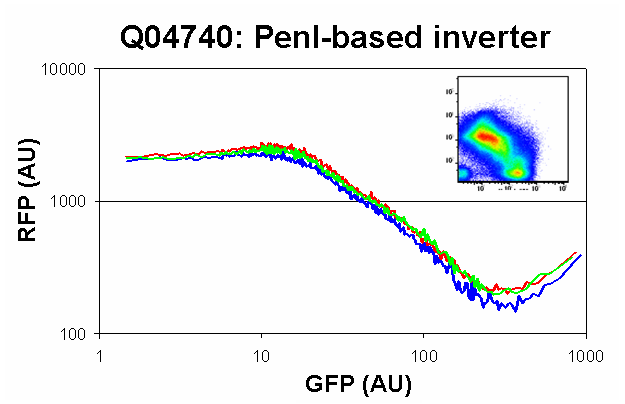
| Q04740sp.PNG: | q04740 characterization with SP1.0 | |
| ScreeningPlasmid1.0.PNG: | Screening Plasmid 1.0 | |
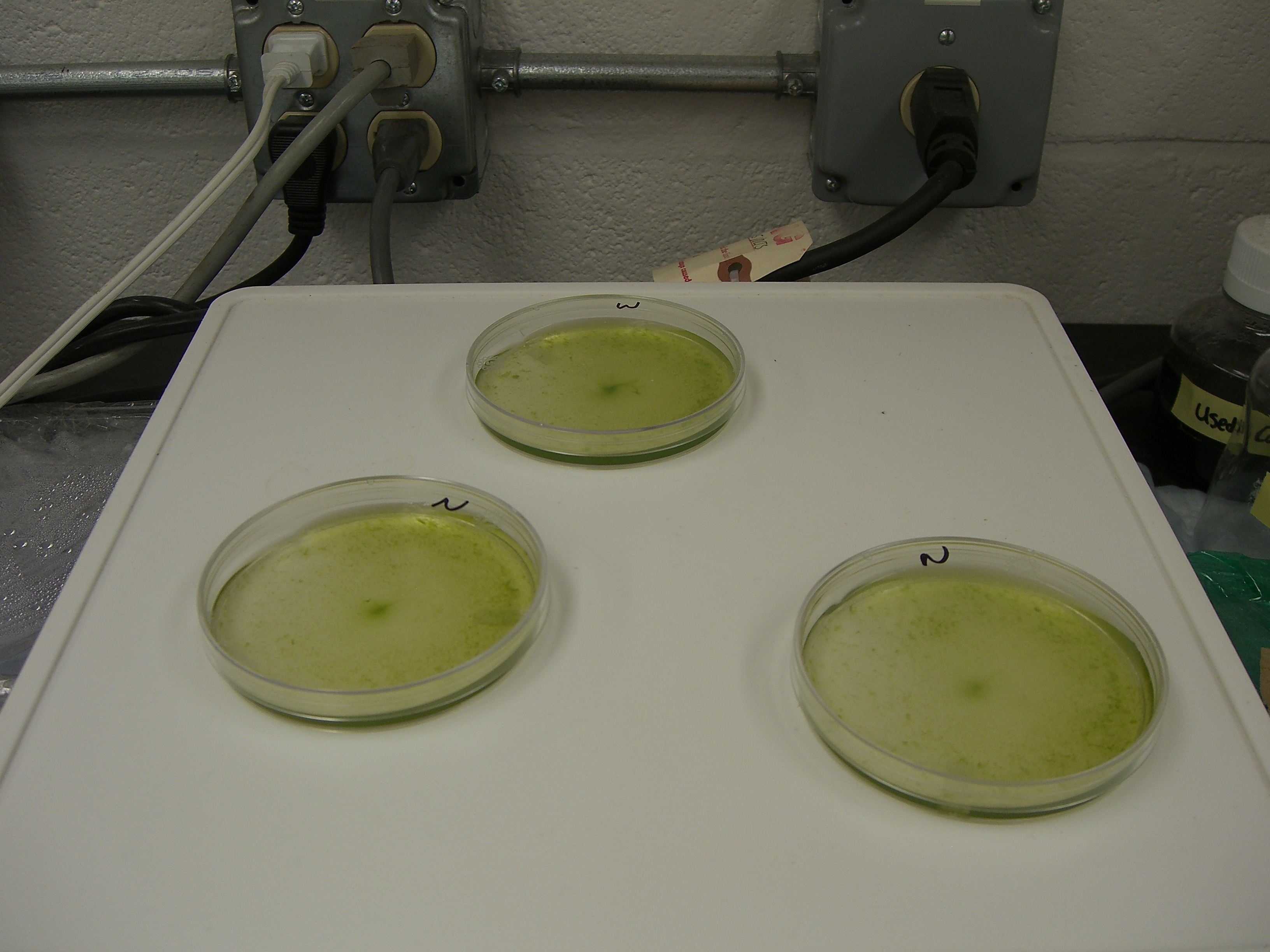
| BeingalgaeSetup.jpg: | BEInG's Setup for growing algae to make H2. | |
| JK BE Retreat 2006.pdf: | BE Retreat 2006 poster | |
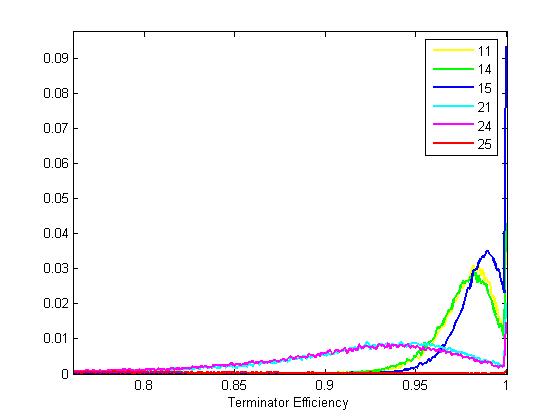
| TermsCorrected.jpg: | Terminator characterization with Screening Plasmid 1.0 | |
| TermDotPLot.JPG: | Dot plot of terminators characterizaed by screening plasmid 1.0 | |
| JK.SyntheticBiology2.0.Poster.pdf: | Jason Kelly's Synthetic Biology 2.0 Poster | |
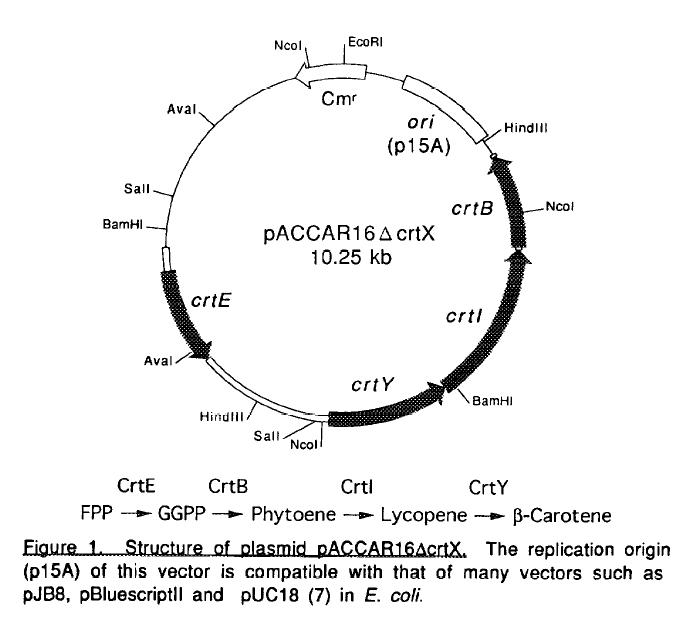
| PACCAR16.JPG: | pACCAR16 structure | |
| OpenWetWare iCampus Spring Review.pdf: | OpenWetWare Spring Review for iCampus Grant | |
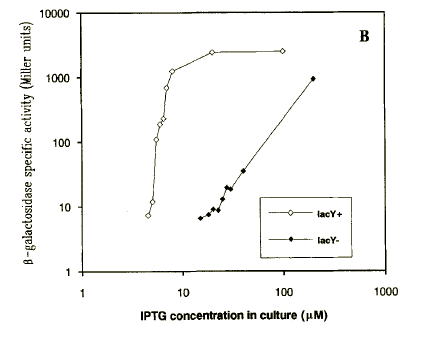
| DelLacYInduction.PNG: | Induction of lac-regulated promoter in lacY- and lacY+ strains, from Jensen et al. Eur.J.Biochem, 211, 181-191 (1993) | |
| MFruit Curves.bmp: | this is a test | |
| I13534vsI13457.JPG: | another test | |
| EndyCirculatorManual.pdf: | Manual for the Polyscienec Circulator. | |
| ICampus-SpringReview-FINAL.doc: | Word version of the OWW iCampus spring review for backup. | |
| July8 2006 gel.jpg: | ||
| Icampus.PNG: | MIT iCampus Logo | |
| Csbi.PNG: | MIT CSBI logo | |
| Jkpic.JPG: | be directory photo | |
| OWW tshirt.jpg: | OWW tshirt option | |
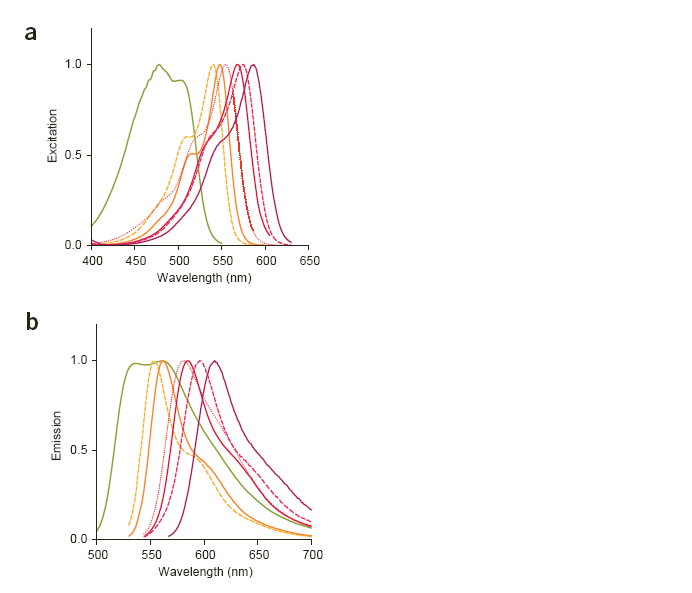
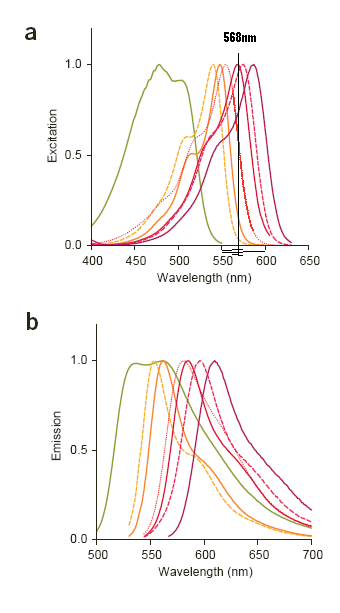
| MFruit Curves.PNG: | mFRuit curves with 568nm laster line approximately positioned on the graph. Image comes from Shaner et al, Nature Biotech, Nov 2004. | |
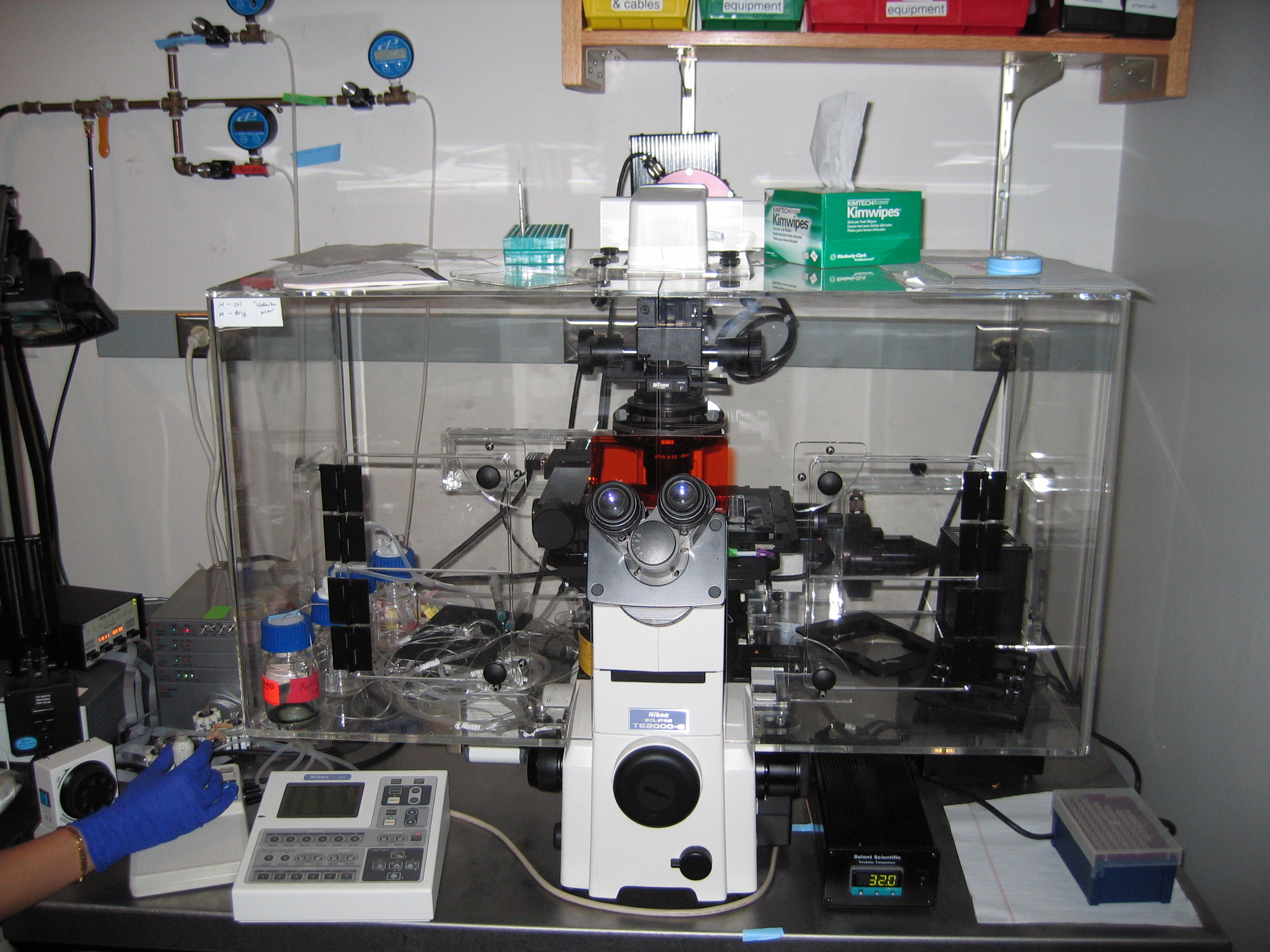
| Endy NikonTE2000.jpg: | Picture of Endy Lab Microscope | |
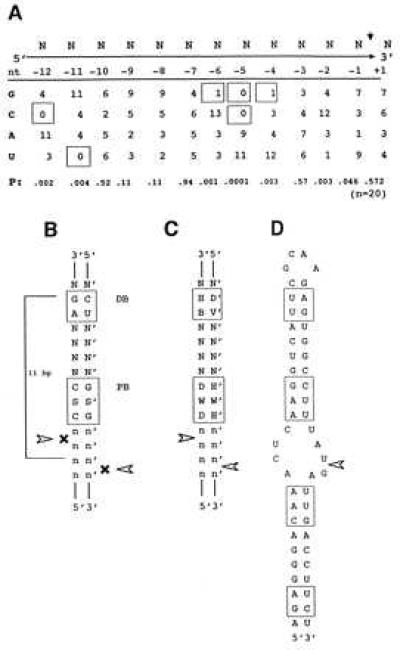
| Nicholson1997Fig1.PNG: | Figure 1 from Zhang and Nicholson, PNAS 1997. (A) Substrate alignment analysis. Ten substrates were aligned whose cleavage sites were accurately determined, either by direct RNA sequence analysis or by primer extension of RNA cleaved in vitro by purified RNase III. (B) The “disfavored” bp, displayed in a dsRNA structure. The proximal box (PB) and distal box (DB) are included within an 11-bp helix. S = C or G, with S′ complementary to S. N, N′ indicate complementary nucleotides; while n, n′ indicate less strict complementarity. Arrowheads indicate the (blocked) cleavage sites. (C) Absence of conservation (degeneracy) of RNase III substrate sequence. H = A, G, U, with D′ (A, C, U) complementary to H; B = C, G, U, with V′ (G, C, U) complementary to B; W, W′ = A, U. (D) Secondary structure of the T7 R1.1 RNase III substrate, showing the proximal and distal boxes and the single cleavage site (arrowhead). | |
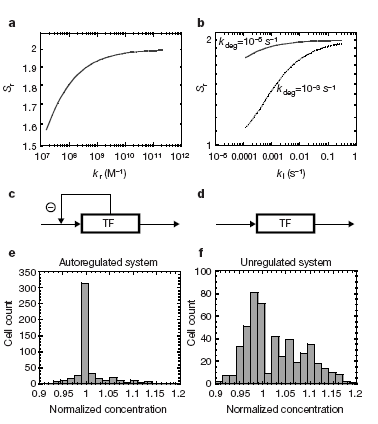
| Fig1-becskei2000.PNG: | figure 1 from becskei and serrano, nature 2000. | |
| Nature Presentation - Feb5-07.ppt: | Presentation to NPG NY office on Feb 5, 2007 by Sri, Jason, and Drew. | |
| OWW round.jpg: | Higher resolution copy of the sticker. | |
| 702px-I13600 - UV 254nm.jpg: | ||
| Valves-partialClosed.PNG: | partially closed valves | |
| PromoterChar Report Revised.doc: | doc on promoter char by Caitlin Conboy | |
| JK.BERetreat.Poster.2007.pdf: | Poster presented by Jason Kelly at MIT BE retreat 3/2007 | |
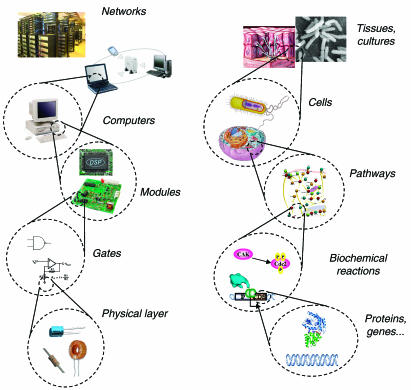
| Weiss-sbreview-f2.jpg: | Figure 2 from Andrianantoandro et al MSB 2006 (permission granted for re-use from NPG and EMBO). | |
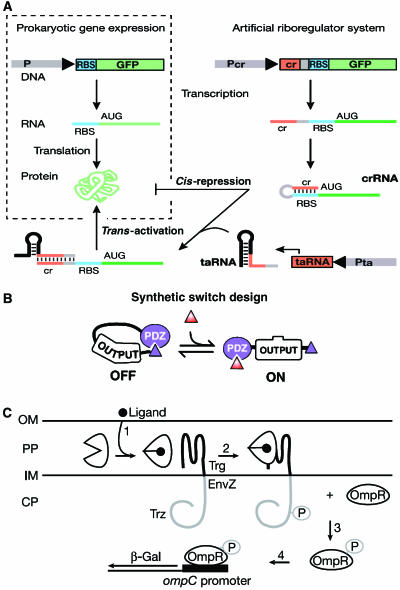
| Weiss-sbreview-f1.jpg: | Figure 1 from Andrianantoandro et al MSB 2006 (permission granted for re-use from NPG and EMBO). | |
| Example.jpg: | ||
| DNAonly vertical.png: | shirt with no circle around logo | |
| DNA only horizontal.png: | no circle around emblem horizontal | |
| MofloBEADS-070520.png: | Example of what Spherotech RCP-60-5 Rainbow Calibration Particles (6.0-6.4um) should look like on the MOFLO under good alignment. 05/20/07 | |
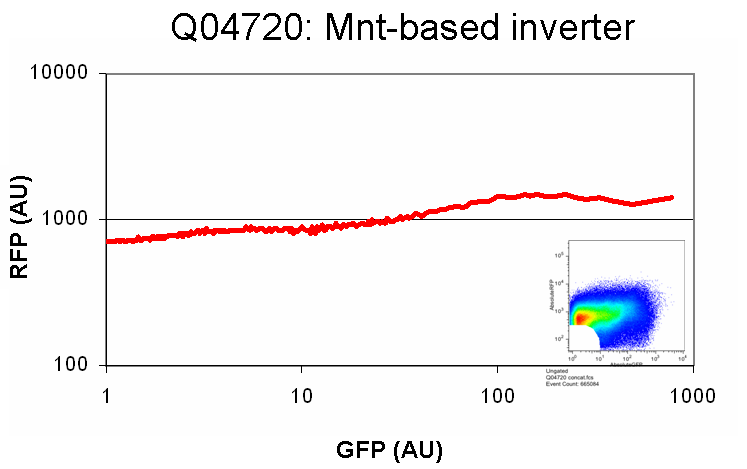
| MntInverter.png: | Mnt-based inverter in SP1.0 | |
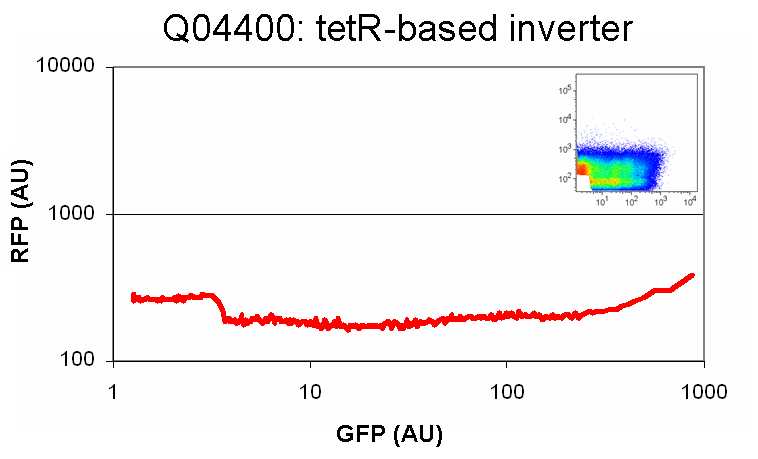
| Q04400InverterInSP1.0.PNG: | Q04400 in SP1.0 | |
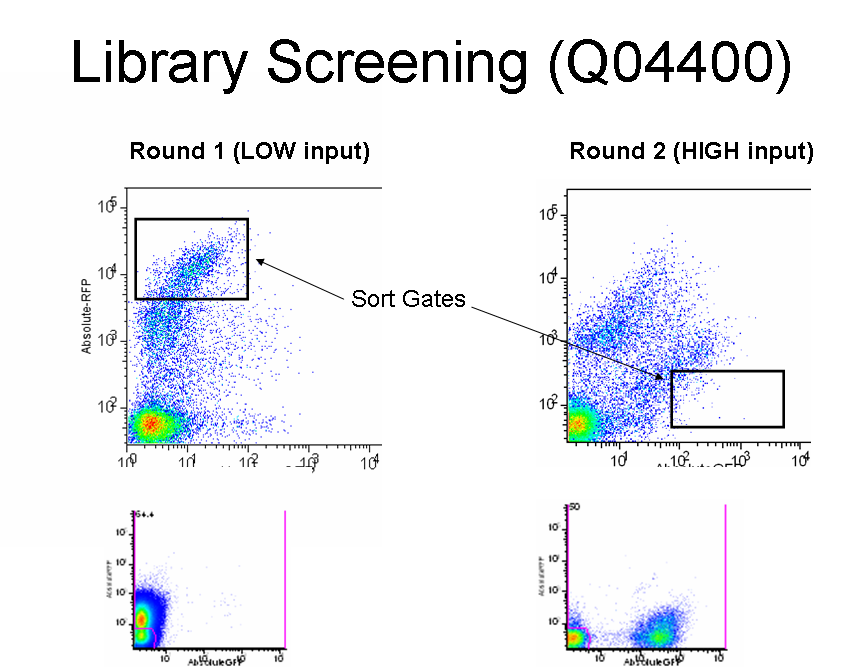
| Q04400LibInSP1.0.PNG: | Q04400 library in Sp1.0 | |
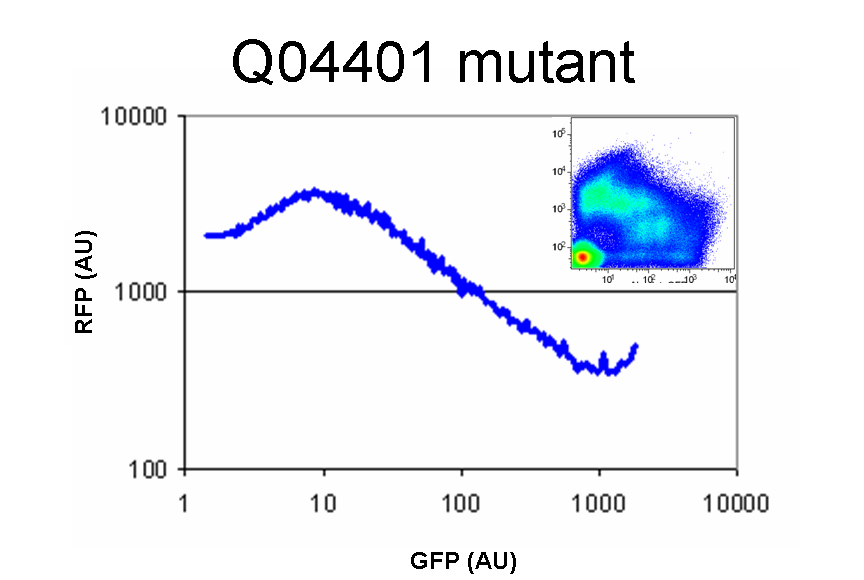
| Q04401InSp1.0.PNG: | Q04401 (mutant isolate from Q04400 library) in SP1.0 | |
| Sortostat v2.0.PNG: | Autocad image of Sortostat v2.0 | |
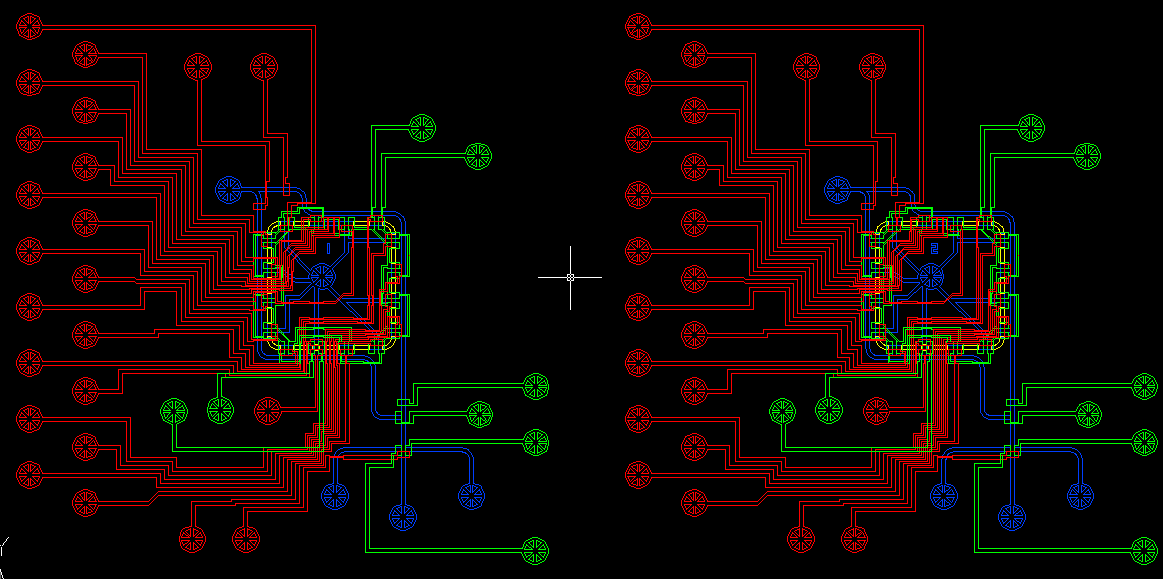
| Sortostat v2.0 - whiteBG.JPG: | AutoCAD image of the Sortostat v2.0 design on white background with layers colored (blue = fluid, red = push UP, green = push DOWN) | |
| Sortostat-valves-partialClosed.PNG: | Example of valves not sealing properly due to fabrication problems. | |
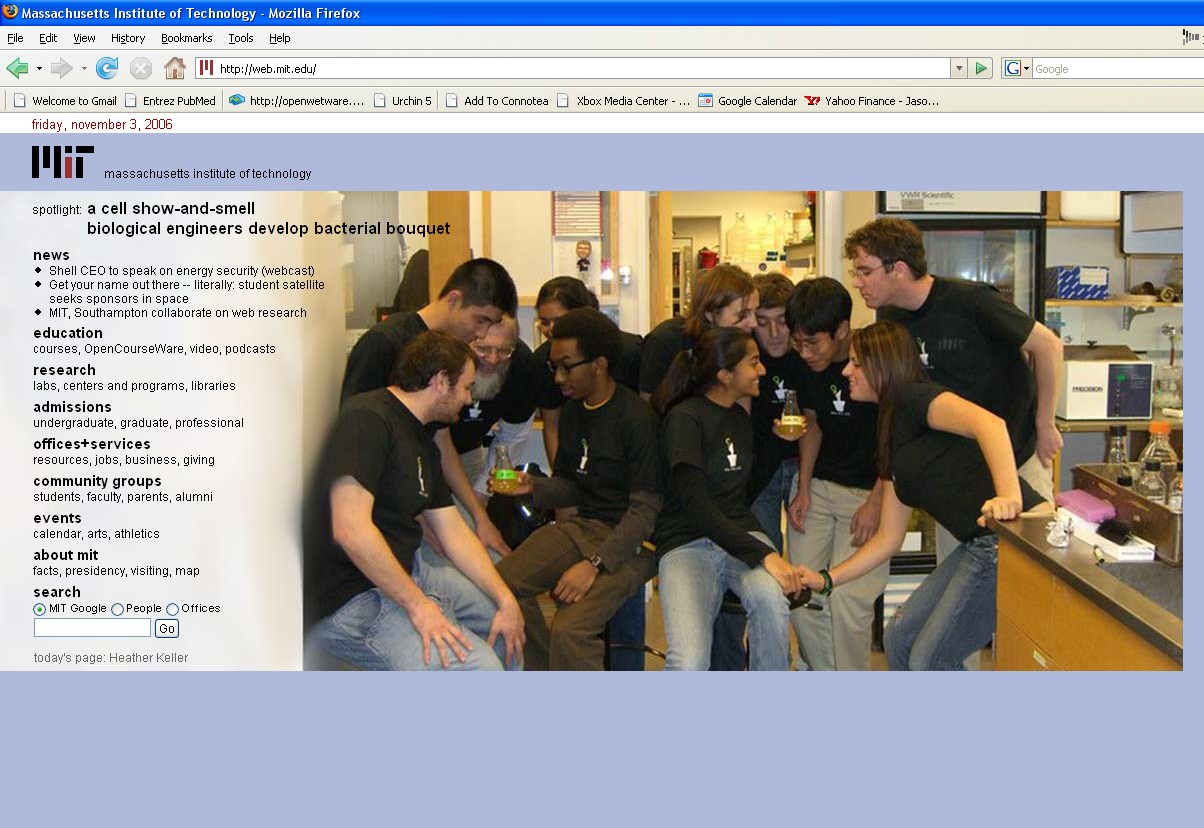
| IGEM-cropped-edited.PNG: | picture of MIT home page w 2006 iGEM team on it | |
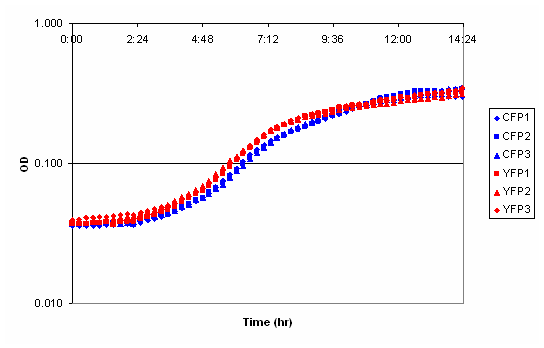
| CFP-YFP plate reader comparison.PNG: | Comparison of CFP and YFP expressing cells in the plate reader. | |
| 07.06.13.jpg: | Gel image | |
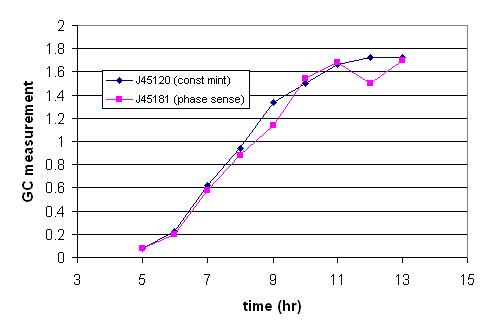
| IGEM06-Mint.JPG: | iGEM results for mint | |
| JK.SB3.0.Poster.2007.pdf: | Jason Kelly SB3.0 Poster | |
| 070909-pcr assembly.jpg: | PCR fusion test gel | |
| 070909-pcr assemblyB.jpg: | ||
| ESP-promoterTest.PNG: | Results of experiment by Jason Kelly to look at possible promoter in the empty screening plasmid | |
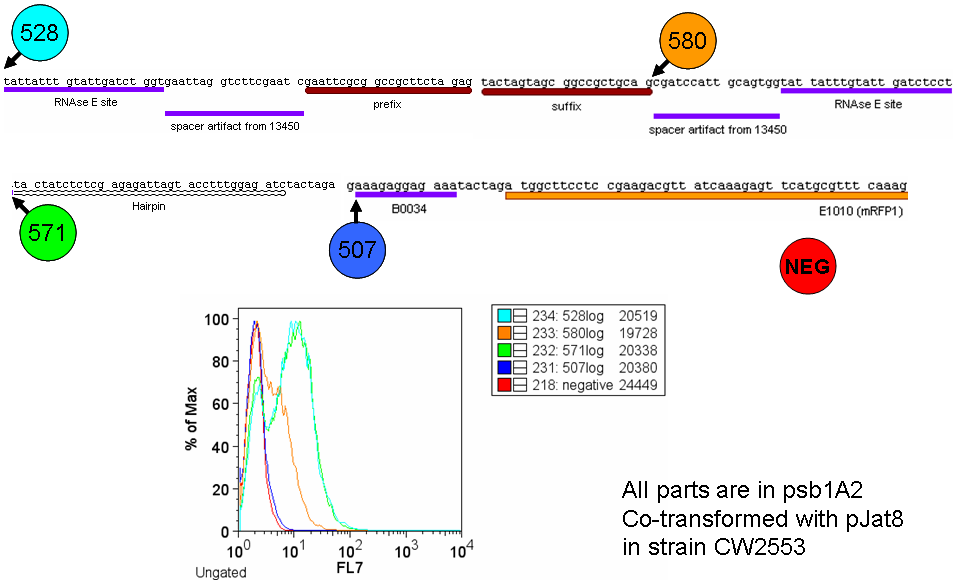
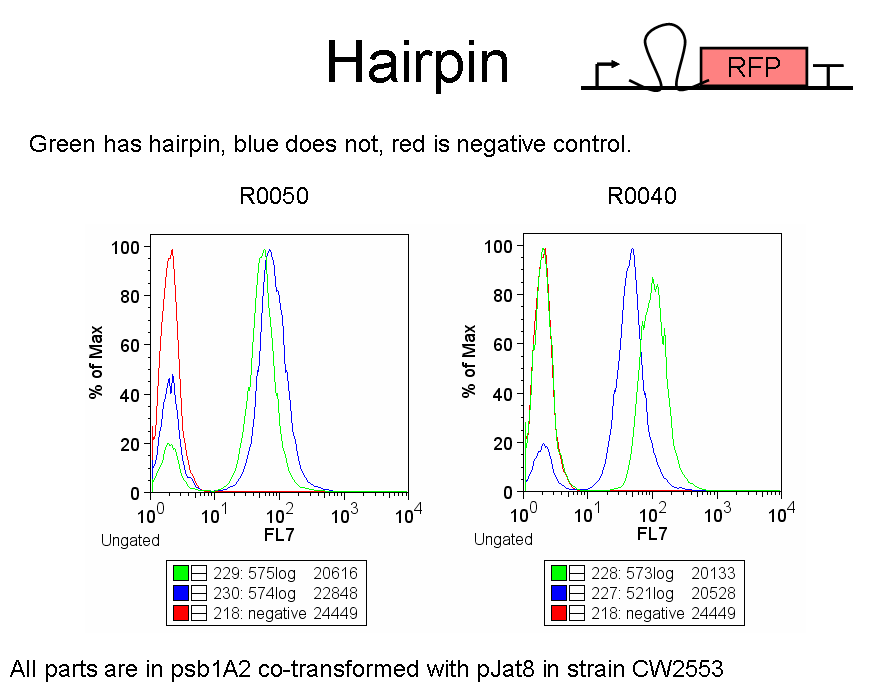
| ESP-hairpinTest.PNG: | Experiment to test hairpin effectiveness in the empty screening plasmid. | |
| Patent squad memo-Breetz-1.doc: | Memo by Hanna Breetz evaluating the viability of the patent goon squad | |
| Ecoli.jpg: | A plush microbe from Giant Microbes.com | |
| LacI-MullerHill94.JPG: | Figure 1 from Muller-Hill et al. EMBO J. 1994 July 15; 13(14): 3348–3355. | |
| OWW-mithomepage.PNG: | OpenWetWare on the MIT homepage | |
| BarberoECDimage.JPG: | Image from Robbie Barbero, Grad student at MIT, from virus-based electrochromic devices that can change from clear to opaque when a charge is applied. | |
| FreeGenes.JPG: | ||
| Psb3k3 Pcr prep.JPG: | preparatory PCR of psb3K3 on gel | |
| RBS tester colony PCR.JPG: | Colony PCR of 15 colonies following a construction using the RBS Measurement Kit | |
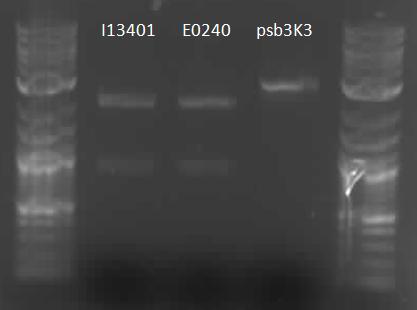
| MeasKit inputs.JPG: | inputs to the measurement kit run on a gel to verify length and digestion | |
| Sortostat f2620.PNG: | ||
| SynBERC talk Reisinger.ppt: | Sarah Reisinger Talk for SynBERC slides from Leonard Katz | |
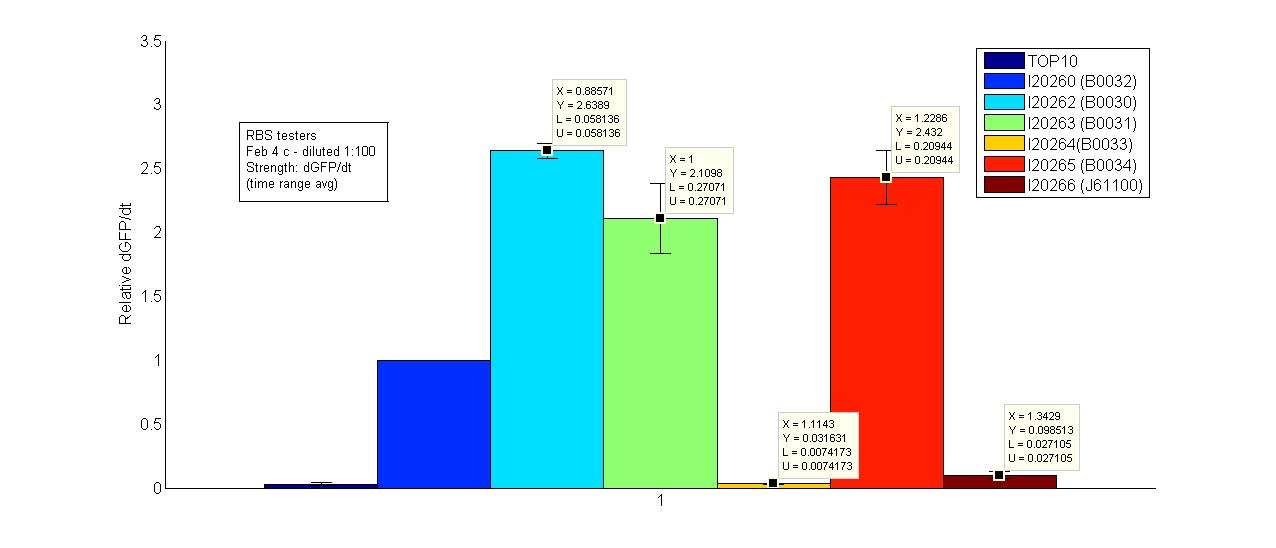
| GFP RBS relative dgfp 080204c.jpg: | RBS tester | |
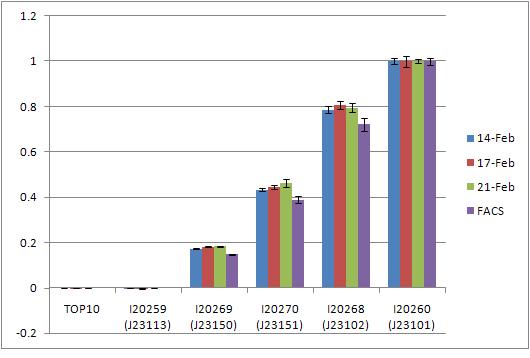
| Promoter tester demo.jpg: | Variety of constitutive promoters inserted into the promoter measurement kit. | |
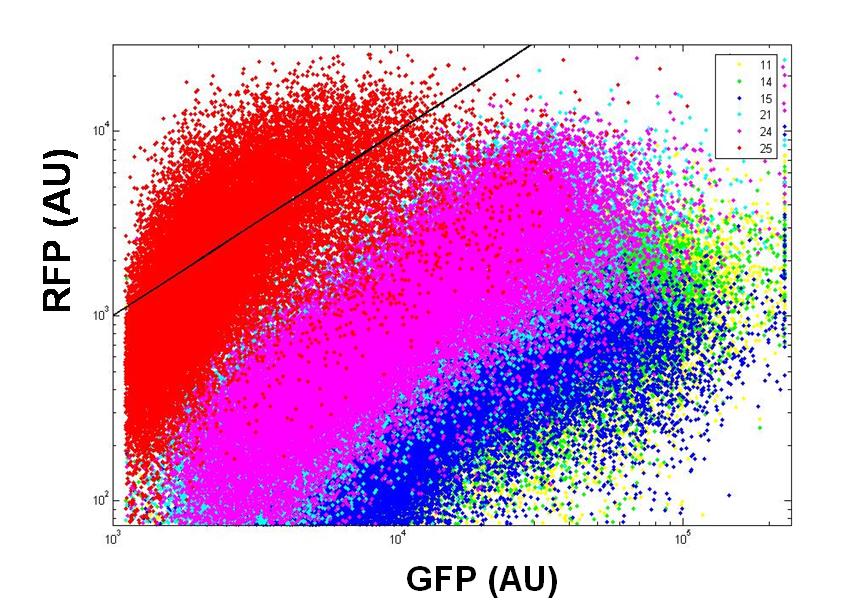
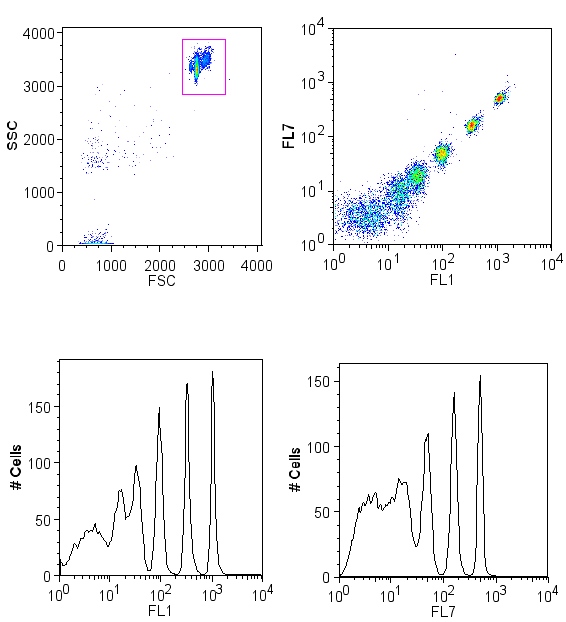
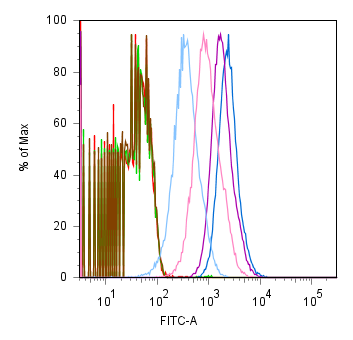
| Measkit promoter FACS.PNG: | ||
| Torch-01-june.gif: | torch | |
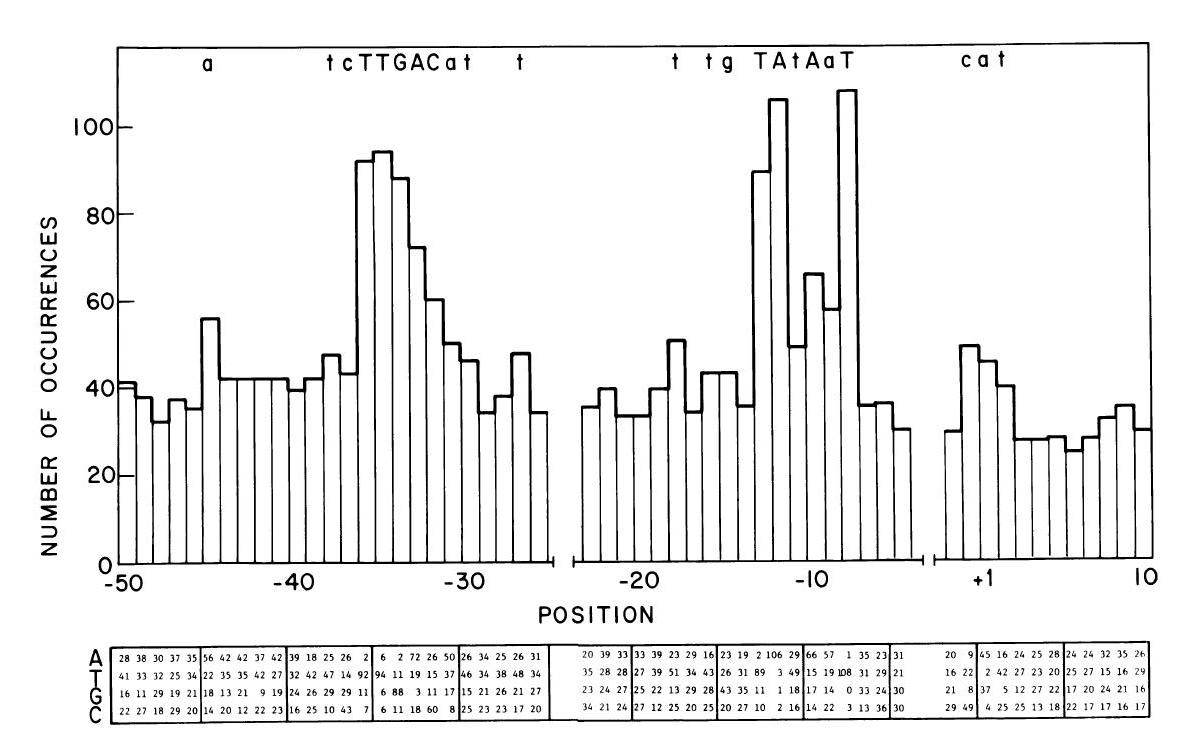
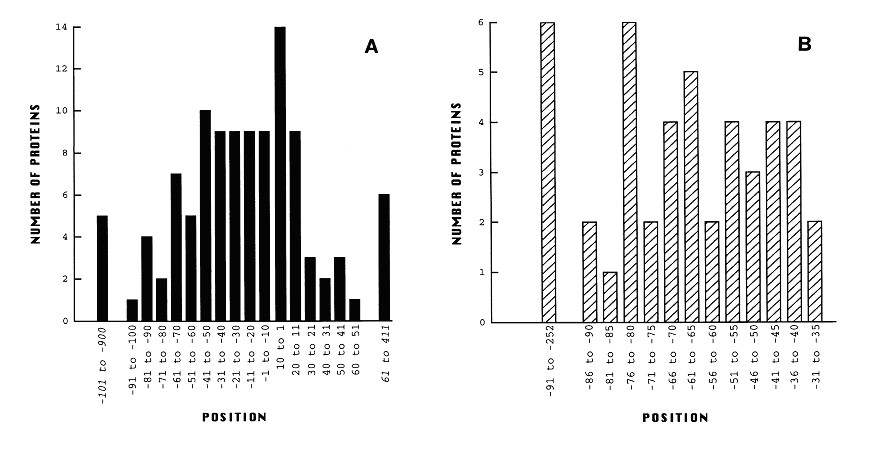
| Promoter table.JPG: | from Hawley & McClure, NAR Vol 11 (8) 1983 Table of nucleotide frequencies in the consensus e.coli sigma70 promoter | |
| PromotersInReg.JPG: | ||
| Proposed promoter standard.JPG: | proposed promoter standard with fixed +1 site | |
| Proposed promoter standard2.JPG: | another option for BB promoter standard. | |
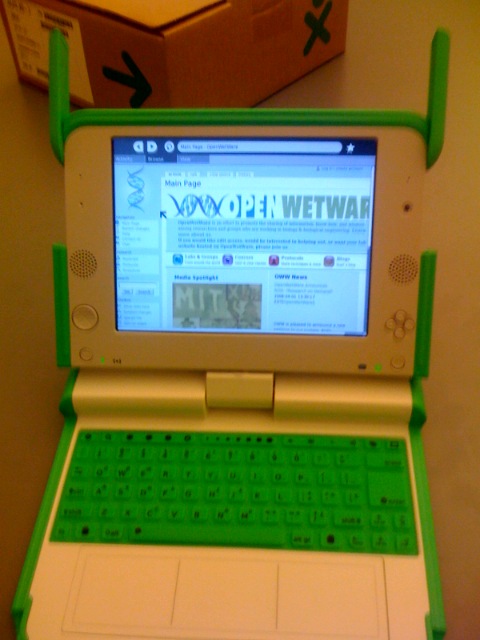
| Oww onOLPC.jpg: | OWW on the OLPC XO computer | |
| Harris-mage.PNG: | harris presents MAGE | |
| Repressor sites.gif: | ||
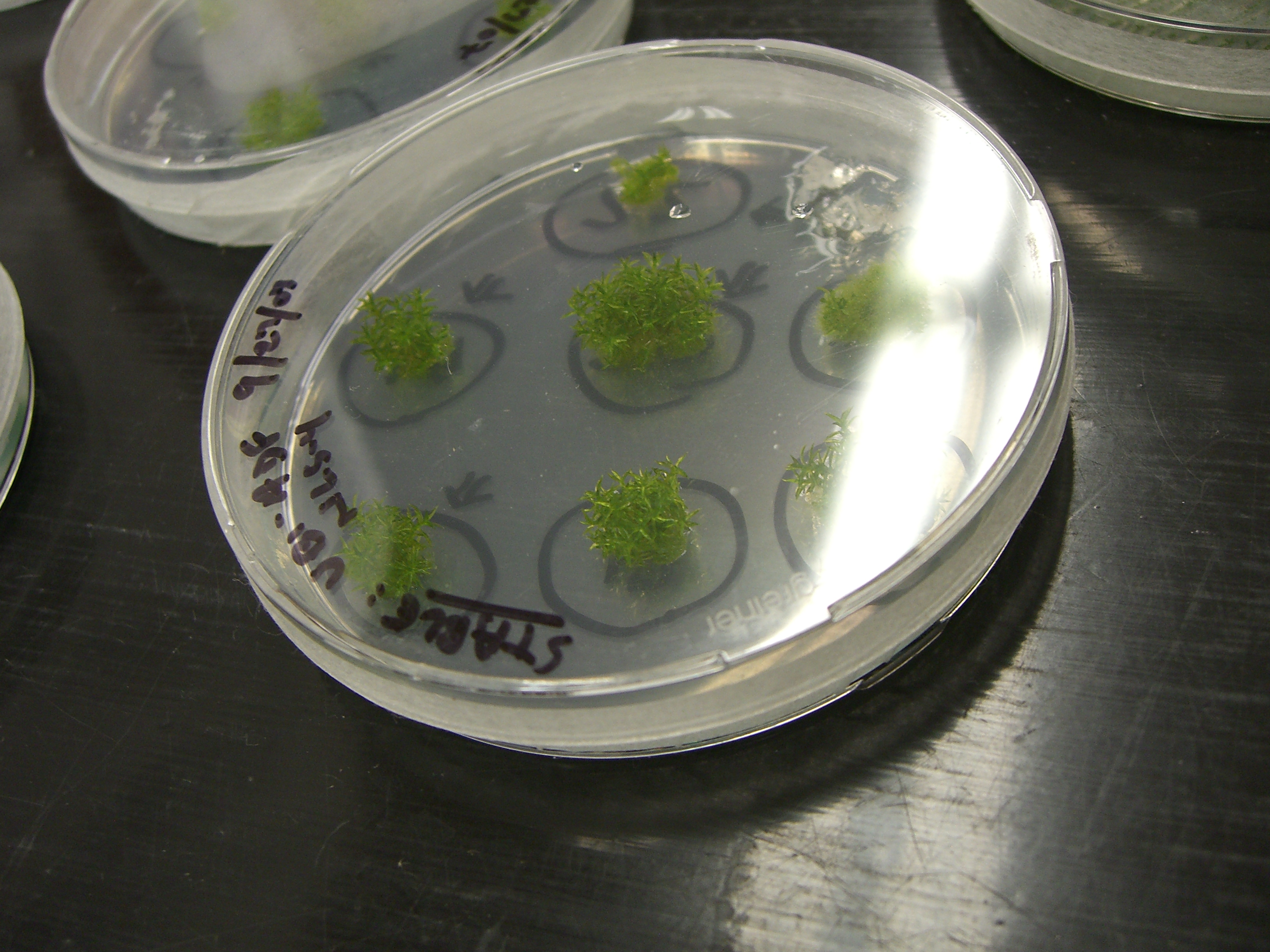
| Moss1.jpg: | ||
| Moss2.jpg: | ||
| Moss5.jpg: | ||
| Moss3.jpg: | ||
| Ginkgologo.PNG: | ||
| JKM Glycerols.xls: | JKM glycerols from endy lab | |
| KC Glycerols.xls: | KC glycerols |