User:Andy Maloney/Notebook/Lab Notebook of Andy Maloney/2010/03/17/Poly-L-lysine
Poly-L-lysine coated slides
So Emmalee and I attempted to coat glass slides with poly-L-lysine so that we can adhere microtubules to them. Our first attempt failed miserably and my second attempt was a wash as well. In the first attempt, we used 5 mg/mL poly-L-lysine in PEM and let this sit just like I would let it sit in a casein passivation study. I didn't wash it out before adding microtubules so we ended up with lots of clumps of microtubules.
My second attempt also failed. I again let the poly-L-lysine sit in a flow cell just like I would in a casein passivation, except I used only 1 mg/mL in PEM. After incubating, I washed out the flow cell with PEM and then introduced the microtubules. After another 10 minute incubation, I washed out the microtubules that didn't stick to the slide with more PEM. This sort of worked as I was able to get microtubules to stick to the slide with no clumping like we saw in the first attempt.
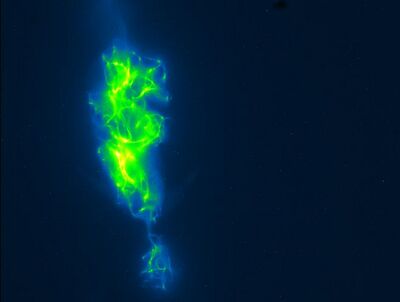
Attempt 1
Above, you can see the clumping that occurred in attempt one. The below picture also shows from attempt one logging effects going on with the microtubules.
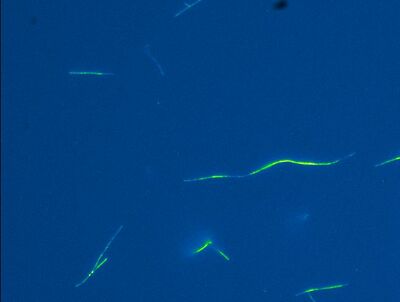
Attempt 2
In attempt 2, I used only 1 mg/mL poly-L-lysine in PEM and again I got the logging effects.
There is a lot less fluorescence in attempt 2 because I flushed out all the microtubules that were not stuck to the glass. This is definitely something I am going to do in the future when sticking microtubules to glass.
Attempt 3
For the third attempt, I am just going to use Sigma's protocol. They state that you should acid wash a slide or soak it in magnesium acetate for 2-3 hours. Then put a solution of 0.1 mg/mL poly-L-lysine in water on the slide. Let dry and you have your prepared surface.
An issue here is that I already put our poly-L-lysine in PEM. This means that when it dries, there will be salt residues left over. I'm guessing that I can just wash it away and I'm just going to try it. The first thing I'm going to do is just try and coat the slides with no acid wash.
Andy Maloney 14:22, 17 March 2010 (EDT): My first attempt at attempt 3 looks as if the slide has a bunch of residues on it. I don't like this at all. I have no idea how to prevent this.
- Steve Koch 14:32, 17 March 2010 (EDT): I guess you need to put it in water? Do we have dialysis available? Of course, just ordering more polylysine is good too.
- Andy Maloney: We have dialysis stuff here. No need since I dilute the poly-L-lysine solution from 5 mg/mL to 0.1 mg/mL.
Andy Maloney 16:55, 17 March 2010 (EDT): I have looked at other lab protocols and other websites that have poly-L-lysine coating protocols. Some say to put the poly-L-lysine in PBS so mine having it in PEM is okay. Predominantly, the prevailing protocol is:
- Wash the slide somehow. Either with an acid or base wash.
- Rinse the slide thoroughly with water.
- Incubate the slides in PLL from anywhere between 5-30 minutes. PLL can be in water or buffer but it is always around 50-100 µg/mL.
- You can rinse after incubation or not. It's up to you but I noticed residues when I didn't rinse so I think I will rinse after incubation.
- Drying is user specific. Lots of people dry in an oven, spin dry, or air dry. I think we will air dry since it is the easiest.
Attempt 4
Andy Maloney 11:09, 18 March 2010 (EDT): Today Emmalee and I will try to look at the fixed microtubules with some of her liposomes. I will redo my poly-L-lysine coating using the a version of the above protocol. I will write it again below so I remember it without myself looking at the above.
- Clean the slides. I'm going to do Ant's protocol with Alconox and the sonicator.
- Let the slides incubate in a bath of 0.1 mg/mL poly-L-lysine for at least 10 minutes which I previously put in PEM. My stock is 5 mg/mL poly-L-lysine in PEM.
- Wash the slides with water.
- Make flow cells from it.
New stock solutions
There are several things that need to be done today. They include:
- Polymerize 29% fluorescein labeled microtubules.
- PEM-T
- Poly-L-lysine slides
Results
Attempt 4 was also somewhat of a wash. But, the PLL coating did work great. Here's what we did.
- We incubated fluorescein labeled MTs on the 0.1 mg/mL PLL coated slides for 10 minutes. We did this uncovered where the MTs were susceptible to room lights.
- We then washed the slide with PEM-T, 3 volumes worth.
- We added a 1:2 dilution of Emmalee's liposomes that have TxRed on them and PG lipids.
- Sealed the slide and observed.
What we observed was that there was just way too much fluorescence coming from the liposomes in solution. We did find that there were some liposomes crowding around the fixed MTs.
Attempt 5
In order to get a better picture for liposomes crowding around the microtubules, we have opted to give it another try. In this attempt, we will do the following:
- Incubate 29% fluorescein labeled MT on the 0.1 mg/mL PLL coated slide for 10 minutes. In the dark.
- Wash the slide with 3 volumes of PEM-T+AF. We don't want to dilute the AF as previously done.
- Add a 1:2 dilution of liposomes labeled with TxRed and incubate for 10 minutes. I should note that we are illuminating the TxRed liposomes with a rhodamine cube.
- Wash the slide again with 3 volumes of PEM-T+AF to get rid of any liposomes not stuck to the MTs. It may turn out that we might have to incubate longer but this is just a first attempt.
- Seal and observe.
This turned out much better.
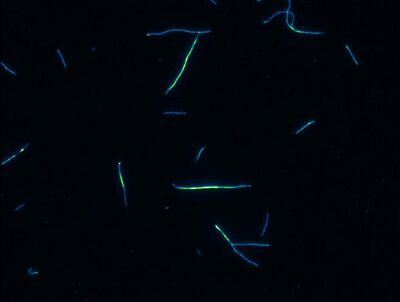
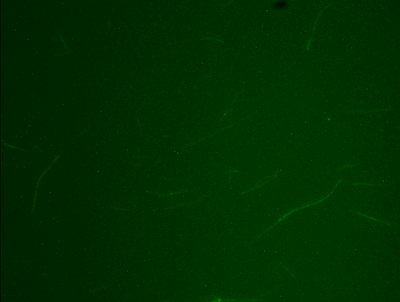
Liposomes under the rhodamine filter
Microtubules
As you can see, the fluorescein tagged MT can be seen using the fluorescein filter. It can also be seen using the rhodamine filter. I'd say that this worked pretty well.