User:Amie S. Krupnick/Notebook/Biology 210 at AU
16S Sequence March 5, 2015
Purpose The purpose of this experiment was to use PCR to sequence the 16S gene and identify the exact species of bacterium. DNA from the bacteria was isolated in the colonies and primers were used with PCR to selectively amplify the 16S gene. It was hypothesized that if PCR and gel agarose were run on bacterium samples, than sequencing the DNA of various samples would be possible, especially with the help of computer systems.
Materials and Methods A single colony of bacteria was transferred from the agar plate to a 100 micro liter of water in a sterile PCR tube. 4 different tubes were made using 4 different bacteria colonies. A primer/water mixture was in each of the PCR tubes. The tubes were incubated at 100 degrees Celcius for 10 minutes in a heat block. After heating, the samples were centrifuged for 5 minutes at 13,400 rpm. Once the centrifuging was complete, 5 micro liters of supernatant was transferred to the 16S PCR reaction and the tubes were place in the PCR machine. The following week, the 4 PCR products were run on an agarose gel. 2 of the 4 samples were determined clear enough to be used for DNA sequencing (Bentley et al 2015). This lab was then done using a computer program that translated gene sequences into species identification by giving the highest match to the MB number that was given after sequencing.
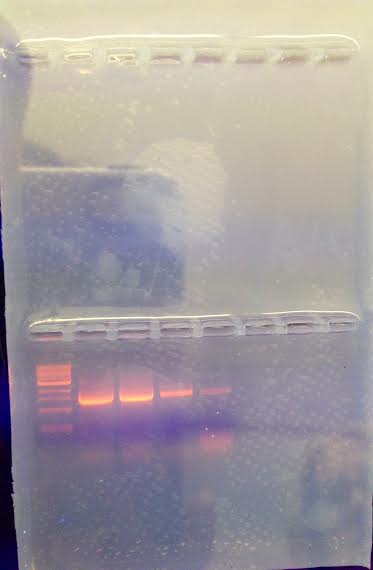
Results After running the gel, Figure 1 shows the resulting bands that appeared from the 4 samples. The 2 bands closest to the ladder were deemed to most successful, and were the ones sent for DNA sequencing. After using the computer system to translate the DNA sequence into a species, two types of bacteria were Chryseobacterium and Variovarax. Below, each bacteria's exact DNA sequence is shown.
Figure 1: Agarose Gel with 4 Sample Bacterium
MB 35: Chryseobacterium NNNNNNNNNNNNNGCTNNGCAGCCGAGCGGTAGAGATTCTTCGGAATCTTGAGAGCGGCGTACGGGTGCGGAACACGTGT GCAACCTGCCTTTATCAGGGGGATAGCCTTTCGAAAGGAAGATTAATACCCCATAATATATTAAGTGGCATCACTTGATA TTGAAAACTCCGGTGGATAAAGATGGGCACGCGCAAGATTAGATAGTTGGTAGGGTAACGGCCTACCAAGTCAGTGATCT TTAGGGGGCCTGAGAGGGTGATCCCCCACACTGGTACTGAGACACGGACCAGACTCCTACGGGAGGCAGCAGTGAGGAAT ATTGGACAATGGGTGAGAGCCTGATCCAGCCATCCCGCGTGAAGGATGACGGCCCTATGGGTTGTAAACTTCTTTTGTAT AGGGATAAACCTTTCCACGTGTGGAAAGCTGAAGGTACTATACGAATAAGCACCGGCTAACTCCGTGCCAGCAGCCGCGG TAATACGGAGGGTGCAAGCGTTATCCGGATTTATTGGGTTTAAAGGGTCCGTAGGCGGATCTGTAAGTCAGTGGTGAAAT CTCACAGCTTAACTGTGAAACTGCCATTGATACTGCAGGTCTTGAGTAAGGTAGAAGTAGCTGGAATAAGTANTGTAGCG GTGAAATGCATANATATTACNTANNNNACCAATTGCGAAGGCAGGTTACTATGTCTTAACTGACNCTGATGGACNAAAGC GTGGGGAGCGAACAGGATTNNATACCCTGGTAGTCCACNCCGTAAACGATGCTAACTCGTTTTTGGGNCTTCGGATTCNN ANACTAANCGAAAGTGATAAGTTAGCCNACCTGGGAGTACNGTTNNNNAGAATGAAACTNNNANNANNNACGGGGGCCNN NNNNACCGGNNNNTATNTGGTTTNAGTGCNATGATNCGCGAGNANCNNTACCACGCNTNAANTGGGGAAGTGAGGNNGGT TNGNNNGNNNNCGTNNNTNCNNCAANTTTCAANNNNNTCCNGGGTGNNGGNGGGNNGTNNNNNANNGNTANGTNTAGGTN CNNNGACNAGCNCNNCCCCCNNNNNCGGGNNNGGGCGGNTNNNTTNCNGGANNNCNNNNNGNGNNNCCCTTGNNNNN
MB 36: Variovarax
NNNNNNNNNNNNNNNNNTNNNNTGCAGTCGTANGGNGGTCAGCGCGTANCAATCCTGGCGGCGAGTGGCGAACGGGTGAG
TAATACATCGGAACGTGCCCAATCGTGGGGGATAACGCAGCGAAAGCTGTGCTAATACCGCATACGATCTACGGATGAAA
GCAGGGGATCGCAAGACCTTGCGCGAATGGAGCGGCCGATGGCAGATTAGGTAGTTGGTGAGGTAAAGGCTCACCAAGCC
TTCGATCTGTAGCTGGTCTGAGAGGACGACCAGCCACACTGGGACTGAGACACGGCCCAGACTCCTACGGGAGGCAGCAG
TGGGGAATTTTGGACAATGGGCGAAAGCCTGATCCAGCCATGCCGCGTGCAGGATGAAGGCCTTCGGGTTGTAAACTGCT
TTTGTACGGAACGAAACGGCCTTTTCTAATAAAGAGGGCTAATGACGGTACCGTAAGAATAAGCACCGGCTAACTACGTG
CCAGCAGCCGCGGTAATACGTAGGGTGCAAGCGTTAATCGGAATTACTGGGCGTAAAGCGTGCGCAGGCGGTTATGTAAG
ACAGTTGTGAAATCCCCGGGCTCAACCTGGGAACTGCATCTGTGACTGCATAGCTAGAGTACGGTAGAGGGGGATGGGAA
TTCCGCGTGTAGCAGTGAAATGCGTAGATATGCGGAGGAACACCGATGGCGAAGGCAATCCCCTGGACCTGTACTGACGC
TCATGCACGAAANNNGTGGGGAGCAAACAGGATTANATACCCTGGTAGTCCACGCCCTAAACGATGTCNACTGGTTGTTG
GGTCTTCACTGACTCANTAACNAANCTAACNCGTGAAGTTGACCGCCTGGGGANTACGGCCGCAAGGTTGNAAACTCANN
GAATTGANNGGGNNCCNCACAANCNGTGGATGATGTGGTTTATTTNNTNNCAACGCNAAAACCTTACCCACCTTTNGACA
TNTACGGAATTCGCCNGANATGGNTTANTGCTCGANAGNANAANCNNTACANCNNGTGCTGCATGGCTGTNNTCNNCTCG
TGTNNNTGNNANNNTGGNTNANNCCCNCAACNNNCGCAACCCNNNNTCATTNNNTGGCTACNTTCGNNTNGGCNNTCTNN
GGNNACTGCCNGNTGANNNAC
Conclusion The purpose of this experiment was to use PCR to sequence the 16S gene and ultimately identify the species of bacteria from the transect. The two species of bacteria that were identified from transect five were Chryseobacterium and Variovarax. Chryseobacterium is a gram-negative bacteria that is largely found in soil and water. This bacteria was responsible for the colony of bright yellow bacteria that was used as a sample in the previous labs leading up to this. Variovarax is a gram-negative, rod-shaped bacteria mostly found in soil (Kim 2006). IJSEM notes that Variovarax "was isolated from greenhouse soil cultivated with lettuce" which aligns with the findings of this experiment. Variovarax was found in the sample that was taken from the bottom area of the Hay Infusion Culture near some vegetation. The resulting bacterium species aligned very well with the experimental data.
Work Cited
Bentley, M., Walters-Conte, K., Zeller, N. (2015). Identifying Bacteria with DNA Sequences. General Biology II, 1(1), 25-32.
Kim, B., Weon, H., & Yoo, S. (2006). Variovorax soli sp. nov., isolated from greenhouse soil. International Journal of Systematic and Evolutionary Microbiology, 56, 2899-2901. Retrieved March 3, 2015
ASK
The Effects of Varying Saline Concentrations on the Development and Survival of Embryonic Zebrafish
Introduction This experiment tests the effects of salinity at varying concentrations on the development of embryonic zebrafish. The experiment will be done using a control, where zebrafish embryos live in distilled water and two variables- embryos living in 1% salinity and embryos living in 2% salinity. Over the course of two weeks, the development, maturation and survival of the zebrafish will be observed and recorded. The purpose of this experiment is to observe any and all effects that the concentration of saline has on the development and survival of embryonic zebrafish. It is hypothesized that if two sets of embryonic zebrafish are tested in two different salinity concentrations, then the higher saline concentration will have more detrimental effects on embryonic zebrafish development and survival.
Materials and Methods Day 1: Initial Set Up of Zebrafish Experiment During the initial set up, three petri dishes were prepared, each containing 20 living zebrafish embryos. The 60 embryos were selected from large tanks that contained hundreds of living and deceased embryos. A transfer pipette was used to individually pick up each living embryo. After transferring 20 embryos into each petri dish, a dissecting scope was used to ensure all selected embryos were, in fact, living. One petri dish was made the control- 20 mls of distilled water was added. The other two petri dishes were the variables- one dish had 20 mls of a 1% saline solution added and the other dish had 20 mls of a 2% saline solution added. Each dish was labeled accordingly and placed in the back of the room until day 2 observations. Day 2: First Day of Experimental Observation and Data Collection The first day of data collection occurred about 30 hours after initial set up. The dissecting scope was used to observe each dish and how many embryos were still living and how many were dead. Table 1 shows the number of zebrafish living and dead, as well as the developmental stage during each day of data collection. . All dead eggs were removed using a transfer pipette. Figure 4, the Zebrafish development table, was used to help determine in what stage each living embryo was in. Day 5: Second day of Data Collection Day 5 began just like day 2- a dissecting scope was used to examine each of the three petri dishes and all dead zebrafish were removed using a transfer pipette. The number of living and dead zebrafish were counted and added to Table 1.The living zebrafish were examined under the dissecting scope to observe developmental stage, movement, shape and other physical characteristics, all of which was also added into Table 1. 10 mls of solution was removed from each petri dish and 25 mls was subsequently added. Each petri dish also had 1 drop of paramecium added using a glass dropper, to feed the zebrafish. 3 depression slides were used to more closely observe the length, movement, hatching and development of a sample from each petri dish. Day 8: Third Day Data Collection Day 8 used the same methods as day 5. At this point, all zebrafish in the 2% S were dead. No observations on these were made, but one was preserved for comparison with a preserved 1% S living zebrafish. A dissecting scope was used to closely examine the zebrafish to observe developmental stage, movement, shape and other physical characteristics. As done in prior days, 10 ml of 1% S was added to the 1% petri dish by way of a pipette-aide. All phenotypic data, along with the numbers of living and dead zebrafish were recorded in Table 1. The zebrafish were not fed on this day.
Day 12: Fourth Day Data Collection This day of data collection used the same methods and materials as day 8.
Results
Over 12 days of data collection, all observable changes were recorded in an ongoing table, Table 1.
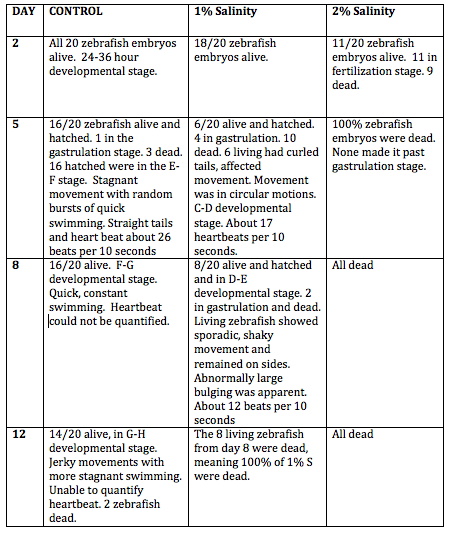 Table 1: Results of Zebrafish Embryo Observation
To discuss development in terms of A-G, Figure 1&2 below were used to determine embryonic development each day.
Table 1: Results of Zebrafish Embryo Observation
To discuss development in terms of A-G, Figure 1&2 below were used to determine embryonic development each day.
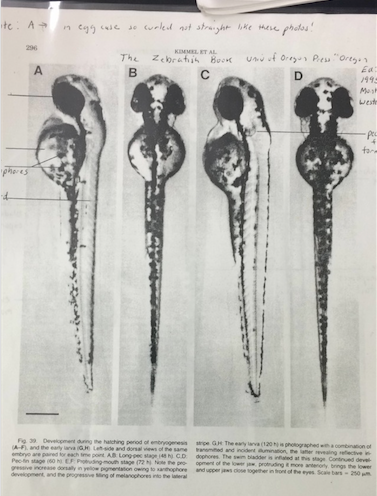
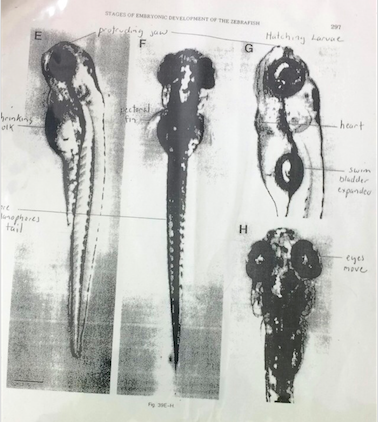 Figure 1&2: Photo Reference for Developmental Stage Determination
Figure 1&2: Photo Reference for Developmental Stage Determination
 Figure 3: Zebrafish embryo in 24 hour gastrulation phase.
These embryos were in the 24 hour gastrulation phase in 1% Salinity on day 5. All other observations were recored in Table 1.
Figure 3: Zebrafish embryo in 24 hour gastrulation phase.
These embryos were in the 24 hour gastrulation phase in 1% Salinity on day 5. All other observations were recored in Table 1.
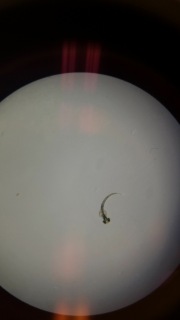 Figure 4: 1% Salinity Hatched Zebrafish Embryo
Figure 4 shows what the 1% Salinity hatched embryo looked like. All other observations were recored in Table 1.
Figure 4: 1% Salinity Hatched Zebrafish Embryo
Figure 4 shows what the 1% Salinity hatched embryo looked like. All other observations were recored in Table 1.
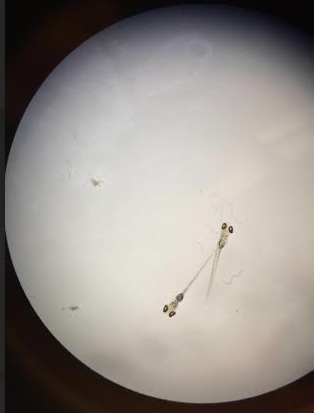 Figure 5: Day 8, Control Zebrafish
Figure 5 shows what the control looked like on day 8 of data collection. All other observations were recored in Table 1.
Figure 5: Day 8, Control Zebrafish
Figure 5 shows what the control looked like on day 8 of data collection. All other observations were recored in Table 1.
 Figure 6: Day 8, 1% Salinity Zebrafish
Figure 6 shows what the 1% Saline zebrafish looked like on day 8 of data collection. All other observations were recored in Table 1.
Figure 6: Day 8, 1% Salinity Zebrafish
Figure 6 shows what the 1% Saline zebrafish looked like on day 8 of data collection. All other observations were recored in Table 1.
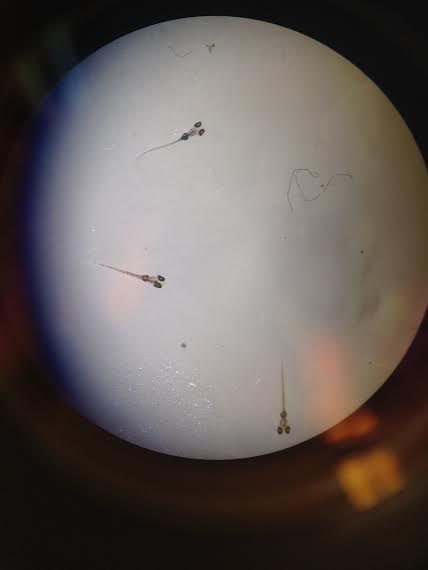 Figure 7: Day 12, Control Zebrafish
Figure 7 shows what the control zebrafish looked like on day 12 of data collection. All other observations were recored in Table 1.
Figure 7: Day 12, Control Zebrafish
Figure 7 shows what the control zebrafish looked like on day 12 of data collection. All other observations were recored in Table 1.
 Figure 8: Control Zebrafish Preserved After Day 12
Figure 8 shows what the preserved zebrafish looked like after 12 days of data collection.
Figure 8: Control Zebrafish Preserved After Day 12
Figure 8 shows what the preserved zebrafish looked like after 12 days of data collection.
Conclusion The purpose of this lab was to conduct an experiment in pairs and create different environments to then observe zebrafish development. In this lab, saline concentration was the variable of choice. A control group was made with distilled water and 2 variable groups were made- 1% S and 2% S. In each dish 20 zebrafish embryos were initially placed and observed for the course of the experiment. The results showed that 2% saline was much too high of a concentration for zebrafish embryos to develop and survive. As shown in Table 1, by day 5 100% of zebrafish in the 2% dish were dead. After 1 day, 9/20 zebrafish were dead in the 2% S solution environment. The majority of the experiment ended up comparing 1% S to the control, distilled water environment. Day 5 lead to vast differences between the control group and the 1% group. The control fish had 26 heartbeats per 10 seconds, whereas the 1% group had only 17 beats per 10 seconds. Similarly, the curled tails of the 1% group lead to affects on movement that the straight tails of the control group did not see. By Day 8, the development of the zebrafish was significantly different. The control zebrafish were in F-G stage, while the 1% were still in D-E stage. Similarly, the heart rate of the 1% fish was much slower, at 12 beats per 10 seconds, than the control group, which was too fast to be quantified. The further development of the control made it impossible to view the heart beating inside the fish. Figure 5 and 6 clearly show developmental differences between zebrafish on Day 8. 1% S fish show an abnormally large bulge that is not apparent in the control fish. This bulge lead to the zebrafish spending most of their time on their sides, instead of swimming upright as the control fish did. Figure 8 shows the preserved control fish for final comparisons. Zebrafish were preserved to be able to more closely examine their development. The 1% preserved fish was not alive upon preservation and was not as developmentally advanced as the fish in Figure 8. This lab supported the hypothesis that if zebrafish were placed in both 1% and 2% Salinity environments, the 1% S fish would develop at a more natural and normal rate when compared to the 2% S fish.
Work Cited
Bentley, M., Walters-Conte, K., Zeller, N. (2015). Embryology & Zebrafish Development. General Biology II, 1(1), 50-62.
ASK
2.24.15 Incomplete entry. No data for Invertebrate lab. No information on Vertebrates or food web. SK
Invertebrates February 19, 2015
Purpose This lab had two main objectives: "to understand the importance of invertebrates and to learn how simple systems evolved into more complex systems" (Bentley et al 2015). Acoelomates, pseudocoelomates and coelomates were observed with respect to their structure and function, as well as cross sections of slides under a microscope. The Burlese Funnel contents were examined after a week for the presence of invertebrates. Through this observational study, it was proposed that if a Burlese Funnel was created with vegetation and soil from the transect and left for one week, then during analysis various invertebrates would be identified.
Materials and Methods Part one of this lab was strictly observational. A dissecting scope was used to more closely analyze Planaria. Annelida, commonly known as earthworms, were observed in a white container full of soil. Nematodes were observed under a microscope as well. Part two of this lab dealt with Arthropods. Jars were set up all over the lab bench, each containing a different arthropod. Archnida, diplopoda, chilopoda, insects and crustacea were all viewed. Part three
2.19.15
Very good lab book entry. Included most relevant data and descriptions of procedures. You could have included location in transect where samples were collected from.
SK
Plantae and Fungi February 12, 2015
Purpose This lab had two main objectives: "to understand the characteristics and diversity of plants and to appreciate the function and importance of Fungi" (Bentley et al 2015). Five plant samples from the transect were obtained and identified based on various criteria. The collected samples were used to make a Berlese Funnel to collect invertebrates. Through this observational study, it was proposed that if five plants samples were collected from the transect for analysis, then the samples can be characterized based on structure and function.
Materials and Methods In this experiment, five plant samples from the transect were collected in one plastic bag. In a second plastic bag, about 500 g of leaf litter was collected from the transect, which was used to make the Berlese funnel. Back in the lab, the five samples were laid out and analyzed on a number of characteristics, as seen in Table 1. Vascularization, specialized structures and mechanisms of reproduction were the three categories of discussion. After these samples were identified, a lily flower was given to dissect and further understand the reproductive organs of a flower. Part V of the lab was observing fungi. In this section, various fungi, like mushrooms and black bread mold, were observed under dissecting microscopes. The respective structure and make up was noted. Lastly, the lab called for the Berlese funnel set up to collect invertebrates. 25 ml of a 50:50 ethanol/water solution was poured into a 50 mL conical tube. A piece of screening was taped into the mouth of a funnel to keep leaf litter from falling through. The leaf litter sample was placed into the mouth of the funnel and the funnel was inserted into the conical tube and taped together. The taped funnel and tube was attached to a ring stand with a 40 watt incandescent lamp above it. The lamp and funnel was covered with foil and sat for 1 week.
Data and Observations The 5 plant samples obtained from the transect were all leaves. The transect did not have much diversity, which limited options during sample collection. Table 1 outlines the location, size and shape of each sample along with the respective vascularization, specialized structures and mechanisms of reproduction. All 5 plant samples were vascularized and used alteration of generation as their mechanism of reproduction. In terms of specialized structures, all had the presence of a cuticle- a waxy layer on their outer surface- except for sample number 4. Sample number 4 was prickly to the touch and did not have the same thick cuticle the other 4 possessed. It was concluded that all 5 samples were dicots. This was concluded due to each sample having 2 cotyledon, being broad leaves with networks of veins and vascular bundles in rings. The 5 samples were all between 1.5-3 inches and were pulled from tall trees or low bushes. Sporangia are small, black, globelike structures that contain spores necessary for sexual reproduction when the sporangia open. The black bread mold was analyzed closely under the dissecting microscope and determined to be a fungi. The black bread mold falls under the Ascomycota category, which includes yeasts, mildews, molds and truffles. Many ascomycetes are pathogenic and reproduce sexually (Bentley et al 2015).
Table 1: Characterization of Plant Samples
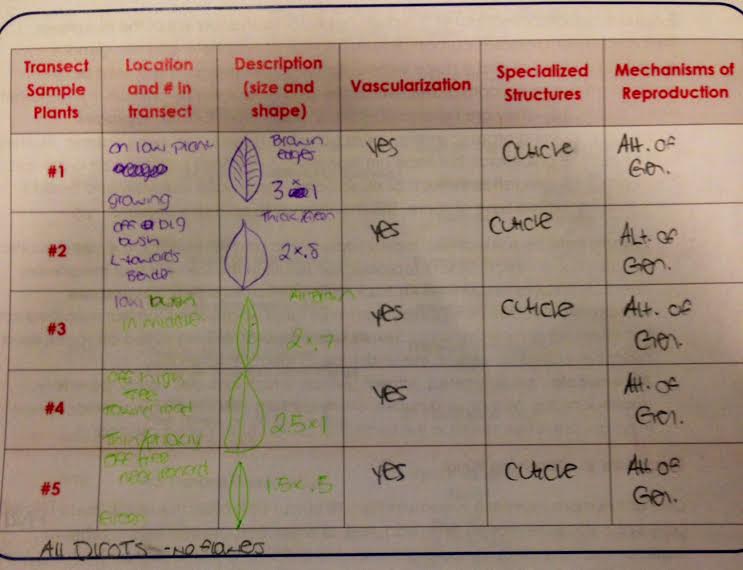
Conclusion
If plant samples were analyzed from the transect, then it is possible to characterize them based on various criteria. This observational lab allowed for further understanding of plant make up and characterizations. Table 1 clearly defined the similarities and differences between the 5 collected plant samples.
Work Cited Bentley, M., Walters-Conte, K., Zeller, N. (2015). Plantae and Fungi. General Biology II, 1(1), 33-43.
ASK
2.10.15 Very good entry. Only correction would be that the PCR tubes were placed in the PCR thermocycler and samples will be sent for sequencing the 16s gene. SK
Microbiology and Identifying Bacteria with DNA Sequences February 5, 2014
Purpose This lab had three main objectives: "to understand the characteristics of bacteria, to observe antibiotic resistance and to understand how DNA sequences are used to identify species" (Bentley et al 2015). In the week prior, agar plates were made to observe during this lab. I propose that mostly bacterium species will have grown on the agar plates. Before this experiment was started, the Hay Infusion Cultures were observed one last time. Procedure 1 allowed us to study the agar plates and quantify the colonies on each and procedure 3 allowed us to identify the bacteria morphologically. If six agar nutrient plates, three also containing the antibiotic tetracycline, then after a week of growth, the plates with tetracycline will show significantly less growth than those with strictly agar nutrient. This experiment also called for the preparation of PCR for sequencing in the coming week.
Materials and Methods In this experiment, the Hay Infusion culture was brought to the work bench for observation. In the first part of this lab, the agar nutrient plates inoculated from the hay infusion culture was obtained at the work bench and observed. The number, color, size and dimension of each colony on the plate was recorded. Table 1 was created to keep the six agar plates straight- regarding dilution, presence of tetracycline and growth. Prepared slides were obtained to practice viewing and locating bacteria through a microscope and the 100x magnification was used with oil. After practicing with prepared slides, wet mounts were made from the growth on the agar plates. A tiny loop sterilized over the bunsen burner was used to transfer agar growth onto the wet mounts. The wet mounts were observed using the 10x and 40x objectives. Table 2 depicts the description of the slides. Next, the gram stain procedure was done. A tiny loop was sterilized over a flame and used to scrape up a very small amount of growth from the agar plate surfaces. The sample was mixed into a drop of water on a glass slide and fixed on the plate by running the slide through a flame 3-4 times with the bacterial smear side up. With the help of a staining tray, the bacterial smear was covered with crystal violet for 1 minute and rinsed off with water. The slides were covered with Gram's iodine mordant for 1 minutes and rinsed off with water. The slides were decolorized by flooding the smear with 95% alcohol for 15 seconds and rinsed. Decolorization is complete when the solvent is completely colorless. Finally, the slides were covered with safranin stain for about 30 seconds and rinsed off with water. Kimwipes were used to dry off the slides before being put back under the microscope for observation. The last portion of the experiment was setting up the PCR for the coming week. 1 entire colony of bacteria from the agar plate was transferred into a 100 ul sterile tube of water. 4 tubes were created like this- 1 previously studied colony from each agar plate. The tubes were incubated at 100 degree Celsius in a heat block for 10 minutes. After 10 minutes, the tubes were placed into a centrifuge for 5 minutes at 13,400 rpm. While the centrifuge was going on, 4 PCR tubes were obtained with 20 ul of primer. Once the centrifuge was complete, 5 ul of supernatant from the samples were added to the PCR tubes and set on ice. The PRC tubes will be observed in the coming weeks lab.
Data and Observations The Hay Infusion Culture appeared thick and dark brown in color with all contents having completely settled. There was substantially less water from the previous observation and a film appeared around the inside of the jar, right above the water line. A similarly putrid oder emanated from the culture, especially when it was moved about. Table 1 showed the 10^-5 nutrient plate to have over 700 colonies, where as the 10^-7 nutrient and tetracycline plate only had 7 colonies. Two of the three plates with tetracycline also showed the presence of mold. Table 2 highlights the bacteria characterization that was seen from the slides, both pre and post gram stain. In this table, three plates did not have tetracycline. The one plate that did have tetracycline was the second 10^-7 dilution. The orange growth looked to be Sarcina and the clear growth, that we came across two times, appeared to be different strains of bacteria. The plate with clear growth containing tetracycline looked to be Bacillus, while the other clear plate without tetracycline looked to be Streptobucilli. All of the gram stains were negative, except for the first slide that lacked tetracycline and was 10^-7 dilution.
Table 1: 100-Fold Serial Dilutions Results
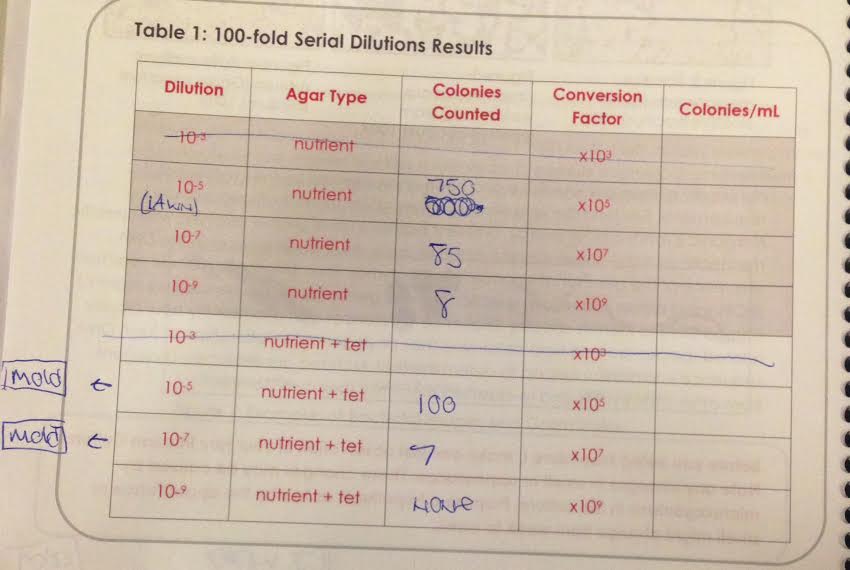
Table 2: Bacteria Characterization
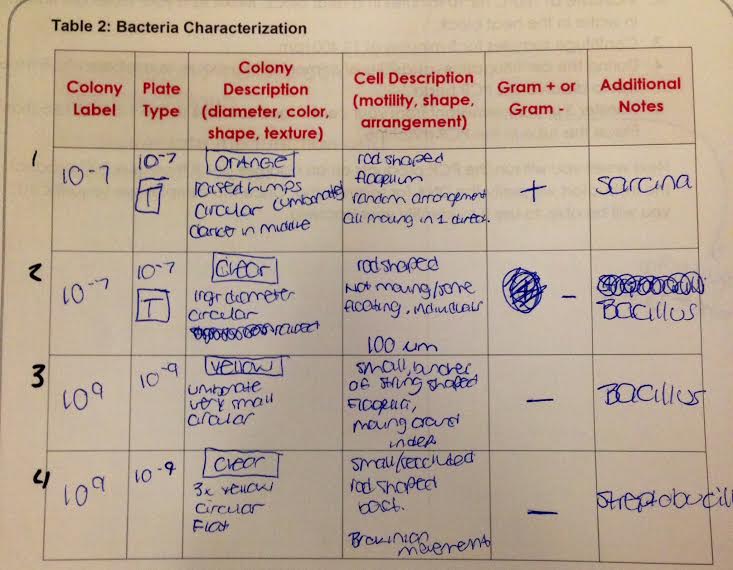
Table 3: Initial Agar Plate Colony Characterization

Conclusion
If a Hay Infusion culture is viewed two times, two weeks in a row, then during the second observation the water level will drop from natural evaporation and the degradation of vegetation will lead to different ratios of life forms and unpleasant appearances and odors. As Table 1 shows, dilution plate 10^-5 without tetracycline had over 700 colonies, almost a lawn of bacteria, whereas dilution 10^-7 with tetracycline only had 7 colonies.
Reviewing Table 1 strongly indicates the effectiveness of tetracycline with certain bacterium strains. The lower the dilution, the less effective tetracycline was. Table 1 shows that 10^-5 had 100 colonies and 10^-9 had no colonies. The less amount of solute and the higher amount of solvent seems to make it easier for the antibiotic tetracycline to be effective in preventing growth. That being said, certain strains may be resistant and others may not be. Based on these samples and results, all of the growth in the agar + tetracycline plates that was not also seen on the agar plates means affected by tetracycline. In the 10^-5 agar and tetracycline plates, both had yellow growth, which indicates whatever that bacteria is is unaffected by tetracycline. Similarly, in the 10^-7 agar and tetracycline plates, both experienced a yellow growth that looked very similar in colony, edge, surface and elevation. The six plates- three with tetracycline and three without- are labeled and described in Table 3.
Work Cited Bentley, M., Walters-Conte, K., Zeller, N. (2015). Microbiology and Identifying Bacteria with DNA Sequences. General Biology II, 1(1), 25-32.
ASK
2.4.15 Very good notebook entry. Well organized, nice pictures and detailed description of Hay infusion and unknown protists. SK
Identifying Algae and Protists January 28, 2015
Purpose The purpose of this experiment was to observe the transect sample in the hay infusion culture after one week. A week in the hay infusion culture allowed time for algae and protists to appear, reproduce and thrive. This experiment made use of a dichotomous key, which allowed accurate identification of different organisms. If a hay infusion culture is created and then observed after one week, multiple different life forms (algae and protists) will appear and be viewable under a microscope for analysis and observation. This experiment also called for the preparation and plating of serial dilutions, which will be used in the coming week.
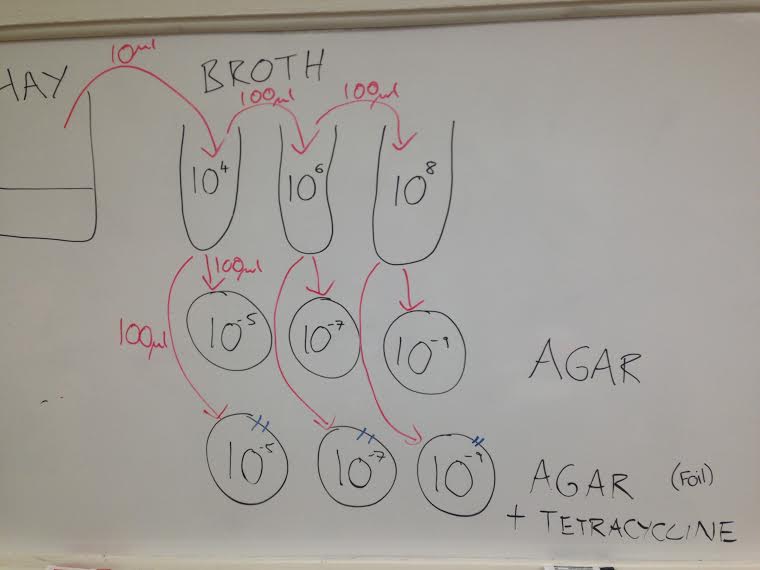
Materials and Methods In this observational experiment, the hay infusion culture was brought to the work station. Using transfer pipettes, two different wet mounts were made with samples from different parts of the culture (top and bottom). Each sample was observed under a microscope to identify different living organisms. A paper and pen was used to draw picture of the observed organisms (Images 1-6). After sample collection was complete, the hay infusion culture was returned to the back table. Part II of the experiment dealt with preparing and plating serial dilutions. To do this, three tubes of 10 mls sterile broth were labeled 10^-4, 10^-6 and 10^-8. Three nutrient agar plates and three nutrient agar plates with tetracycline were obtained and labeled accordingly. One plate from each of the two groups was labeled 10^-5, 10^-7 and 10^-9 with initials, date and group number. Using a micropippetor set at 100 ul and a box of tops, 100 ul of broth from tube 10^-4 was added to tube 10^-6 and mixed. That step was repeated two more times to make the 10^-6 and 10^-68dilutions (Image 5). To plate the serial dilutions on the agar nutrient plates, 100 uL was pipetted from the 10^-4 plate. A glass spreader soaking in alcohol and run through a bunsen burner was used to spread the dilution evenly across the agar nutrient plate. The same procedure was repeated for the rest of the plates, including the +tet plates. Agar nutrient plates with tetracycline were all covered in aluminum foil and both sets of plates were set in the back of the room to incubate for one week.
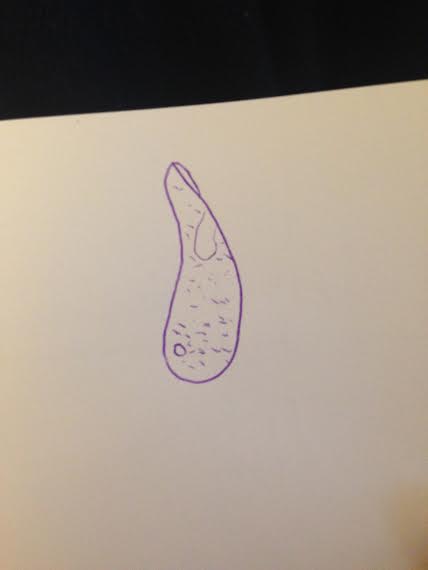
Data and Observations The hay infusion appeared brown in color and had a very putrid odor. The odor was particularly strong with the culture was moved, allowing sealed in smells to escape. There was no life, such as mold or green shoots, apparent on the surface of the liquid. Two samples were taken from the culture- one from the surface of the liquid and one from the bottom of the culture near a leaf. After observation and analysis, the sample from the top resulted in three different organisms- Chlamydomonas, Colpidium and Paramecium Bursaria (Images 1-3). The Chlamydomonas is an algae that are by far the smallest, measuring at 5 micrometers and move by way of flagellum. The Paramecium Bursaria is a protozoan that swim in a corkscrew fashion, which helps determine its identity and use cilia for movement. The Colpidium is a protozoa measuring about 50 micrometers and moves by way of cilia. All three organisms photosynthesize. The sample from the bottom of the culture near a leaf presented three life forms as well- Chlamydomonas, Peranema and Paramecium Bursaria. The one organism that differs is the Peranema (Image 4), which is a photosynthesizing protozoa that moves by way of flagellum and measured to be 40 micrometers.
Image 3: Paramecium Bursaria, 100 um
Conclusion
Two samples were taken from varying parts of the hay infusion culture. The samples showed different organisms living in each. This can be explained by many factors, one of which being more sunlight reaches the top of the culture, versus the bottom of the culture, which could be a more ideal living environment for certain organisms than others. The sample from the bottom of the culture and near the leaf had appearance of Peranema, which was not present in the sample from the top of the culture and not near vegetation. Peranema show to thrive better when surrounded by decaying material, such as the leaf, than in free floating water. Paramecium Bursaria meets all the needs of life- it is comprised of cells, it reproduces (asexually), evolves, maintains homeostasis, undergoes metabolism through photosynthesis and grows.
If the Hay Infusion culture "grew" for another two months, various changes would be predictable. The majority of the water would most likely evaporate, which would ultimately kill the organisms in the mini-habitat. Prior to evaporation, new organisms would grow and develop based on the changes happening to the vegetation and chemical make up of the culture. The selective pressures that would affect the community of samples would be the amount of water, sunlight and useable vegetation to maintain life.
The experiment and the identified life forms supported the hypothesis that if a hay infusion culture was created and then observed after one week, multiple different life forms (algae and protists) would appear and be viewable under a microscope for analysis and observation.
Work Cited
Bentley, M., Walters-Conte, K., Zeller, N. (2015). Identifying Algae and Protists. General Biology II, 1(1), 18-24.
ASK
1.27.15 Good first lab book entry. The transect diagram is a little simple. List biotic and abiotic features. SK
Biological Transect: Observing a Niche at American University January 26, 2015
Purpose The purpose of this experiment was to observe a 20x20 transect at AU for the Biology-210 lab. This experiment and the corresponding set of observations will act as the basis for future experiments, using the samples collected during this initial inspection of the transect. After taking a sample of abiotic and biotic components from the transect, the plot of land was discussed in class. This lab was purely observational and did not yield any findings. A Hay Infusion was created with the collected sample to aid in the following week's lab.
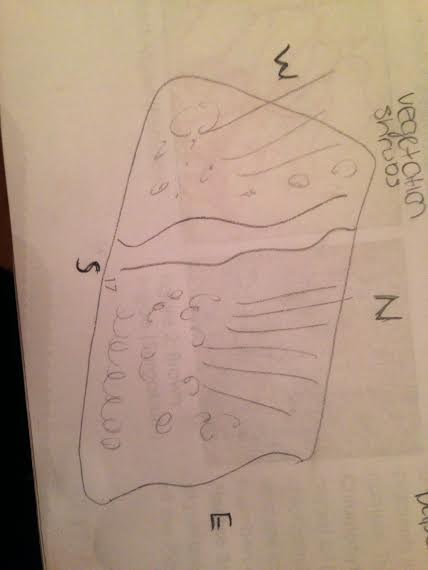
Materials and Methods In this observational experiment, each group was given a flashlight and a sterile 50 ml conical tube for sample collection representative of the transect. After being dropped off at the appropriate transect, the 20x20 plot of land was examined and drawn for reference. The transect is located in the gardens between Hughes Hall and Bender Arena and a concrete pathway running from north to south. The allotted plastic tube was filled with dirt, soil and vegetation from the transect. The transect was drawn, with directions labeled (Image 1). Upon return to the lab, a Hay Infusion Culture was made from the collected sample. 10 grams of soil and vegetation from the conical tube sample was placed in a plastic jar containing 500 ml of deer park water. 0.1 g of dried milk was added into the jar and the jar was mixed. After being labeled, the jar was set down without a lid for a week.
Data and Observations This transect is located on an uneven plot of land that contained many different forms of vegetation (see Image 1). The vegetation found was largely evenly distributed across the transect. The biotic organisms specified are: trees, leaves, bushes, vegetation and shrubs. The abiotic components specified are: snow, dirt, hay, wood chips and paper. During the making of the Hay Infusion Culture, dried milk was added as a source of food for the vegetation in the culture.
Conclusion The jar was set in the back of the lab without a lid for examination of protists and to "inoculate agar petri plates for studying bacteria" (Bentley et al 2015) in the following week.
Work Cited
Bentley, M., Walters-Conte, K., Zeller, N. (2015). Biological Life at AU. General Biology II, 1(1), 13-17.
ASK