User:Alexa Coon/Notebook/Biology 210 at AU
PCR 16S Sequence Analysis 26 February 2015
Purpose
This 16S Sequence analysis was done with bacterial colonies from our Hay Infusions in order to determine the types of bacteria present in our cultures. This procedure will replicate the 16S rRNA gene and read the order to determine the bacteria based on their RNA sequence.
Material and Methods • Transfer bacteria to 100 microliters of water • Incubate for 10 minutes at 100 degrees Celcius • Centrifuge for 5 minute at 13400 rpm • Prepare the PCR tube by adding 20 microliters of primer and water mix to the bead • Transfer 5 microliter of the supernatant to the PCR reaction • Leave in the PCR machine • Put the PCR’d product in an agarose gel to sequence • Send sequences to be analyzed where the RNA strand was compared to other strands in order to match it to the DNA in an already-known bacteria
Data Observations
The highest match for MB 35 was Chryseobacterium which is a gram-negative, non-motile, rod shaped, yellow bacteria found primarily in soil and water but also in raw milk in Israel and some other food products. It was previously categorized as a flavobacterium. MB35 NNNNNNNNNNNNNGCTNNGCAGCCGAGCGGTAGAGATTCTTCGGAATCTTGAGAGCGGCGTACGGGTGCGGAACACGTGT GCAACCTGCCTTTATCAGGGGGATAGCCTTTCGAAAGGAAGATTAATACCCCATAATATATTAAGTGGCATCACTTGATA TTGAAAACTCCGGTGGATAAAGATGGGCACGCGCAAGATTAGATAGTTGGTAGGGTAACGGCCTACCAAGTCAGTGATCT TTAGGGGGCCTGAGAGGGTGATCCCCCACACTGGTACTGAGACACGGACCAGACTCCTACGGGAGGCAGCAGTGAGGAAT ATTGGACAATGGGTGAGAGCCTGATCCAGCCATCCCGCGTGAAGGATGACGGCCCTATGGGTTGTAAACTTCTTTTGTAT AGGGATAAACCTTTCCACGTGTGGAAAGCTGAAGGTACTATACGAATAAGCACCGGCTAACTCCGTGCCAGCAGCCGCGG TAATACGGAGGGTGCAAGCGTTATCCGGATTTATTGGGTTTAAAGGGTCCGTAGGCGGATCTGTAAGTCAGTGGTGAAAT CTCACAGCTTAACTGTGAAACTGCCATTGATACTGCAGGTCTTGAGTAAGGTAGAAGTAGCTGGAATAAGTANTGTAGCG GTGAAATGCATANATATTACNTANNNNACCAATTGCGAAGGCAGGTTACTATGTCTTAACTGACNCTGATGGACNAAAGC GTGGGGAGCGAACAGGATTNNATACCCTGGTAGTCCACNCCGTAAACGATGCTAACTCGTTTTTGGGNCTTCGGATTCNN ANACTAANCGAAAGTGATAAGTTAGCCNACCTGGGAGTACNGTTNNNNAGAATGAAACTNNNANNANNNACGGGGGCCNN NNNNACCGGNNNNTATNTGGTTTNAGTGCNATGATNCGCGAGNANCNNTACCACGCNTNAANTGGGGAAGTGAGGNNGGT TNGNNNGNNNNCGTNNNTNCNNCAANTTTCAANNNNNTCCNGGGTGNNGGNGGGNNGTNNNNNANNGNTANGTNTAGGTN CNNNGACNAGCNCNNCCCCCNNNNNCGGGNNNGGGCGGNTNNNTTNCNGGANNNCNNNNNGNGNNNCCCTTGNNNNN
The MB36 matched most highly with the Variovorax bacteria. This bacteria is a gram-negative bacteria that helps with breaking down organisms and helps plants grow. It makes yellow colonies and has a rod-shape.
MB36
NNNNNNNNNNNNNNNCGNGNNNNNNNNNNTNNTTGCANNNNNNNNCTAACTACTTCTGGCAGAACCCGCTCCCATGGTGT
GACGGGCGGTGTGTACAAGACCCGGGAACGTATTCACCGTGACATTCTGATCCACGATTACTAGCGATTCCGACTTCACG
CAGTCGAGTTGCAGACTGCGATCCGGACTACGACTGGTTTTATGGGATTAGCTCCCCCTCGCGGGTTGGCAACCCTTTGT
ACCAGCCATTGTATGACGTGTGTAGCCCCACCTATAAGGGCCATGAGGACTTGACGTCATCCCCACCTTCCTCCGGTTTG
TCACCGGCAGTCTCATTAGAGTGCCCAACTGAATGTAGCAACTAATGACAAGGGTTGCGCTCGTTGCGGGACTTAACCCA
ACATCTCACGACACGAGCTGACGACAGCCATGCAGCACCTGTGTTACGGTTCTCTTTCGAGCACTAAGCCATCTCTGGCG
AATTCCGTACATGTCAAAGGTGGGTAAGGTTTTTCGCGTTGCATCGAATTAAACCACATCATCCACCGCTTGTGCGGGTC
CCCGTCAATTCCTTTGAGTTTCAACCTTGCGGCCGTACTCCCCAGGCGGTCAACTTCACGCGTTAGCTTCGTTACTGAGT
CAGTGAAGACCCAACAACCAGTTGACATCGTTTAGGGCGTGGACTACCAGGGTATCTAATCCTGTTTGCTCCCCACGCTT
TCGTGCATGAGCGTCAGTACAGGTCCAGGGGATTGCCTTCGCCATCGGTGTTCCTCCGCATATCTACGCATTTCACTGCT
ACACGCGGAATTCCATCCCCCTCTACCGTACTCTAGCTATGCAGTCACAGATGCAGTTCCCAGGTTGAGCCCGGGGATTT
CACAACTGTCTTACATAACCGCCTGCGCACGCTTTACGCCCAGTAATTCCGANTAACGCTTGCACCCTACGTATTACCGC
GGCTGCTGGCACGTAGTTNGCCNNGCTTATTCTTACNNTACCGTCATTAGCCCTCTTTATNNAAAANGCCGTTTCGTTCC
GTACAAANCAGTTTANANCCGAAGGCCTTCATCCNGCNNNGNATGNNGNTCNGCTTNNNCCNTNNCAAAATNCCNACTGC
NNCNCCGNNNNANTCNGGGNNNNNNNN
Conclusions and Future Directions
There were bacteria living in the soil of our transect that we were able identify by analyzing the RNA that we isolated via PCR. This helped us to figure out what their roles in the ecosystem were and why they might have been present in our transect. In the future we could do this with more of the bacterial colonies we were looking at so that we could get a fuller picture of what is in the transect and how all the microbacteria interact to keep the systems working.
4.7.15 Include gel image if possible. Also how did your gram stain results compare with the information you found on the sequenced bacteria. SK
Lab 6 Zebrafish Experiment 19/26 February 2015
Purpose
The purpose of this lab was to determine how the addition of estrogen at various levels affected the development of zebrafish embryos. This was done by adding zebrafish to water with various levels of estrogen and watching how they developed. Our hypothesis was that higher estrogen doses in solution slow down the speed of the development of embryos.
Materials and Methods
• Get 20 zebrafish embryos for each of the petri dishes • Have one petri dish of embryos with 20ml of Deerpark water and 0 g/L of estrogen, a second with water and 12.5 g/L estrogen and a final with 25 g/L of estrogen • Check every few days to remove dead ones and make observations • 2nd day in our fish had died so we transferred them into 2 ml containers individually and added embryos to counterbalance the ones we had lost • Observations of developments of body and eyes made again on day 4 and dead fish removed and documented • Feed on day 7 and observe • Final observations day 13 with documentation and the disposal of all the embryos
Data and Observations
Our lab found that zebrafish embryos that were exposed to heightened levels of estrogen had difficulty developing. This ranged anywhere from meaning full deformation (see picture below) or simply that the reaction times of the living hatchlings were significantly slower than those of their counterparts who had not received the treatment. Some were not hatched or died prior to hatching, many had slow responses, and others had large eyes or shortened tails. Along the way we had many zebreafish die, but the overwhelming majority of them were the treated fish. I would suppose that the fish that died without the treatment had issues with bacteria or died of some kind of natural cause.
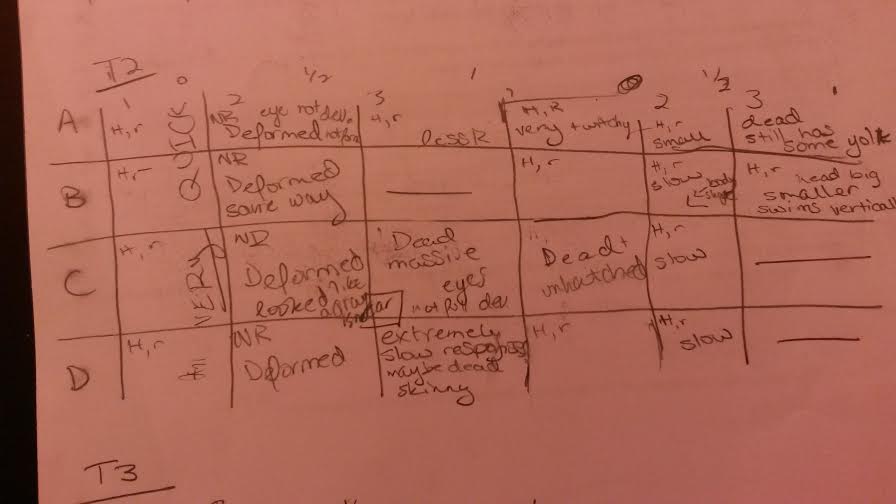
Day 4 Observations

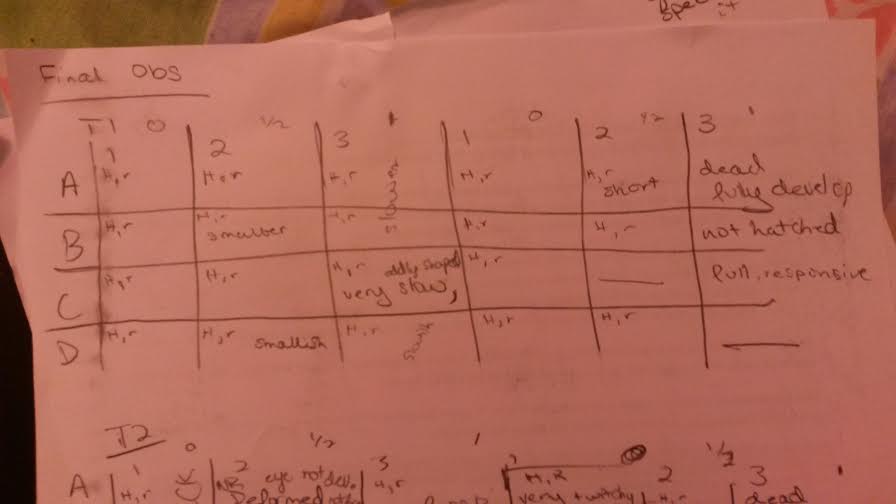
Final Day Observations
These graphics are set up so that H means hatched and R means responsive and any other observations were written in the squares. The first column of each was the control, the second got the half dose, the third got the full dose and this pattern repeats for the next three rows. Those with slashes through them were already empty chambers.
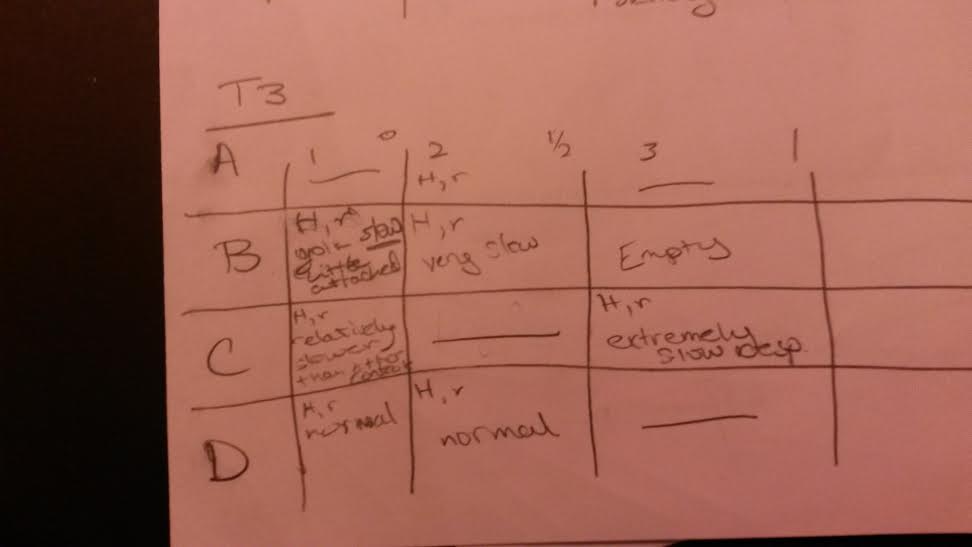
Dead Embryo Final Day
This image is a fish from the half dose (2nd column) that exhibited deformation consistent with the rest of its row.
Conclusions and Future Direction
This experiment supported our hypothesis that embryos would show slowed growth from heightened levels of estrogen in their growing environment. Since many of the embryos had issues with stunted responses and deformations we know now that the estrogen present in the water did slow the way that the fish developed. In the future, if we were to redo the experiment it would be better to have some more specific ways to measure the development of the embryos. If we could get more accurate measurements on size of eyes or tails we would have more quantitative proof that the estrogen had an effect on the fish. It could also be interesting to do a follow up experiment to this with testosterone in the same manner to compare the two.
4.7.15 Good entry. Some of the data difficult to read. SK
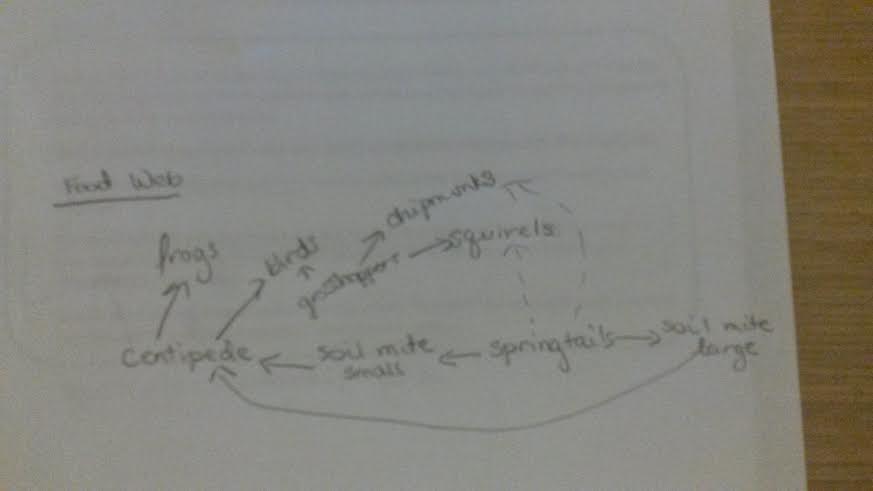
2.24.15 Good entry. Need to include a food web describing relationships between organisms in transect. SK
Lab 5 Invertebrates 12 February 2015
Purpose
This lab was a final look at the transect our lab group received during the first week. We have already looked at the micro level particles that live there, so we are now branching out to the macro level invertebrates that inhabit the space. We hypothesized that since this transect is part of a larger ecosystem it had to have macro level organisms that contributed to the way the transect functioned. We were looking for invertebrates so that we could identify which types were in our transect, how big they were, and look at some of the attributes that we could see in a dissecting scope.
Materials and Methods
• Our group first looked at the nemotodes, planaria, and annelida (earthworms) to see how they moved • We then looked at various arthropods to make observations about their bodies • We then broke down our Berlese funnel o We threw away the contents of the funnel o We poured half of the liquid into one petri dish and the other half into the other o Our group looked at these and other petri dishes from other groups to find invertebrates from our transect We tried to figure out what the organism was, the length, the number we found in the sample, and then we made basic observations about it • We then hypothesized about what vertebrates could live in our transect
Data and Observations
In the earlier parts of this lab we found that nematodes squirm in”s” shapes, small and slow while the Annelidas squirm in an accordion –like, flexible motion. The planaria seemed like they had no body cavity and slinked or curled to get around. When looking at our Berlese Funnel we found 4 types of invertebrates. There was on centipede that was about 0.8 m in length. It was white, long, had lots of legs, and two antennae at both the anterior and posterior ends. There were about 6 small soil mites at about 0.2 mm in our transect that we described as ant-like, with small wings, antennae, is white, and has black eyes. There was one large soil mite that looked round with a spherical midbody, and legs spread out in a sun pattern. That was 0.7 mm long. The springtail came out at 1 mm, the largest in the transect, with lots of legs, antennae like a grasshopper, and with what looked like a hard outer shell.
I suppose that there are probably squirrels, chipmunks, and an assortment of birds who like berries and worms that inhabit this transect in the summer and spring. There are also probably mice that wind up in the area or frogs that come up from the small man-made stream nearby.
Conclusions and Future Directions
The results of the experiment confirm the hypothesis that there were in fact macrolevel invertebrates that help the ecosystem to function. We were able to provide four different types of invertebrates by looking at the water through a dissection microscope, each with unique attributes and unique functions in the ecosystem. This shows that even man-made gardens have dynamic ecosystems. We were also able to gain a background in the breadth of invertebrates that exist. If we were to do this again I would like to pick a different transect since our transect was man-made. I would have liked to see a natural occurring habitat. I would also have preferred to do this in later spring or summer since there would be more life since it would be warmer.
2.19.15
Good entry. Data would be a little clearer if used a data table. You can make this in excel and then transfer it to OWW format using http://excel2wiki.net then just copy and paste table back into OWW. Images would also add some information.
SK
Lab 4 Plants and Fungi 5 February 2015
Purpose
This lab was an investigation of our transect from a macro level, with an eye to the plants and the fungi that are present. Not only were we learning about which organisms were present, we were also looking at the vascularization, the specialization of structures, and the ways in which these organisms reproduced so that we could determine to which phyla each belonged. If any of the samples were fungi, we followed similar patterns to figure out which phyla these belonged to and learn about their basic functions. Finally we set up for next week’s investigation of invertebrates.
Materials and Method
• We took two Ziploc baggies out to our transect and collected some underbrush in one bag and we collected 5 different leaves in the other bag, since leaves seemed to represent the plants we saw in our patch • We didn’t collect any seeds or pine cones or flowers because they weren’t present • We took them back and observed for vascularization, which seemed to be present in each of the plants o We also examined part of a lily and pictures of vascularization to more fully understand the types of vascularization • We observed the leaves closely for any specialization of the cells and observed the pattern in which the leaves grew • Next we looked at the leaves and figured that since there were no flowers on any of the plants and they were all only leaved plants that they probably underwent alternation of generations o We didn’t see any flowers or spores out there, but it seemed based on the leaves that they were all dicots • We observed some fungi under dissecting microscopes • We used the other bag of samples from our transect • We added this to 25 mL of ethanol/water to a 50 mL conical tube with a screening material in the bottom of the funnel on top of the screen • We put this on the rung stand and put it under a light which was all covered with foil and left it for a week
Data and Observations
Our lab found only leaves from our transect. One was on a low ivy-like bush, one was from a big push toward bender arena, another from a short bush in the center of the “garden”, a high tree toward the road, and a far tree toward Leonard. There wasn’t much variation among them. Each was vascularized (so were not bryophytes), similarly sized, between 3 and 1.5 inches by 0.5 to 1 inch, and each seemed to reproduce through alternation of generations. Only one of them did not have cuticle, the leave from the high tree toward the road, which was covered with a soft, prickly layer. They all seemed to be dicots and none of them had flowers.
Conclusion and Future Direction
We found that there are in fact a number of different plants in our transect, but because our transect is man-made and kempt, a lot of what we found there was incredibly similar. We found that the majority of plants that we came across have similar attributes and reproduce similarly. I think that if we had done these with a larger transect or done it later in the year, we would have had more luck with finding differentiation between the species that we were observing. We might have found a flower or spores that would have led to variation in our results.
This is my attempt to figure this site out...hope it worked.
Lab 1 Biological Life at AU
Purpose
This experiment involved observing a tract of land at American University, a niche, as a jumping point for discussing ecosystems. Each group was expected to take down various biotic, or living, and abiotic, or nonliving, things they found in their swatch of land. This helped to assess the biodiversity present across the campus and which populations were found in which locations.
Materials and Methods
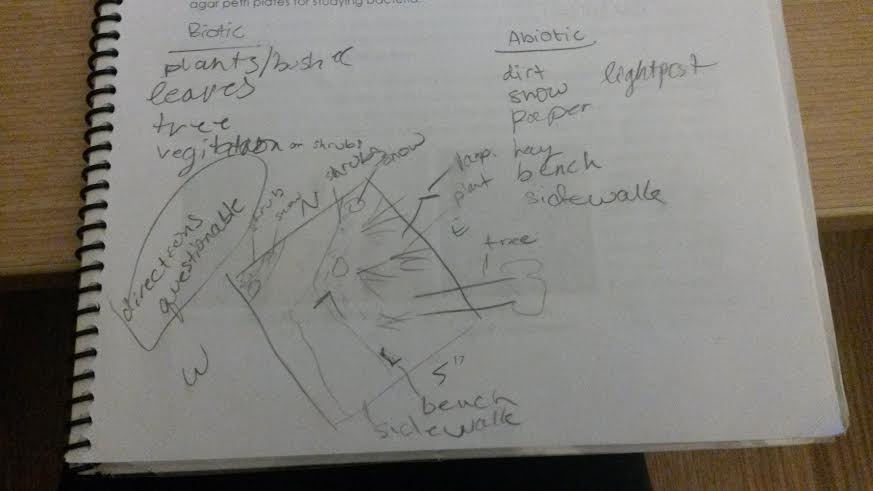
The group in which I worked was assigned a transect near the gym. At the site of the transect we found each of the Popsicle sticks that marked the edges and drew from an overhead perspective what we saw. Each item we observed was labeled. After this we took note of what biotic and abiotic material was present in the space we had mapped. Using a 5mL conical tube we collected 10 to 12 grams of soil as a sample of our transect. This we took back to the lab and added to 500 mLs of deerpark water in a plastic jar as well as 0.1 grams of dried milk. This was all mixed so that the organisms present in the sample could grow so that they could be observed at a later date. The prepared jar was left uncovered at the end of class.
Data and Observations
The area that was assigned to my group was part of a man-made garden which had elements of human impact. So some of abiotic features we found there include a bench, a lightpost, paper, and a sidewalk, as well as more natural things like hay, snow, and dirt. The biotic features included shrubs, trees, leaves, vines, and bushes.
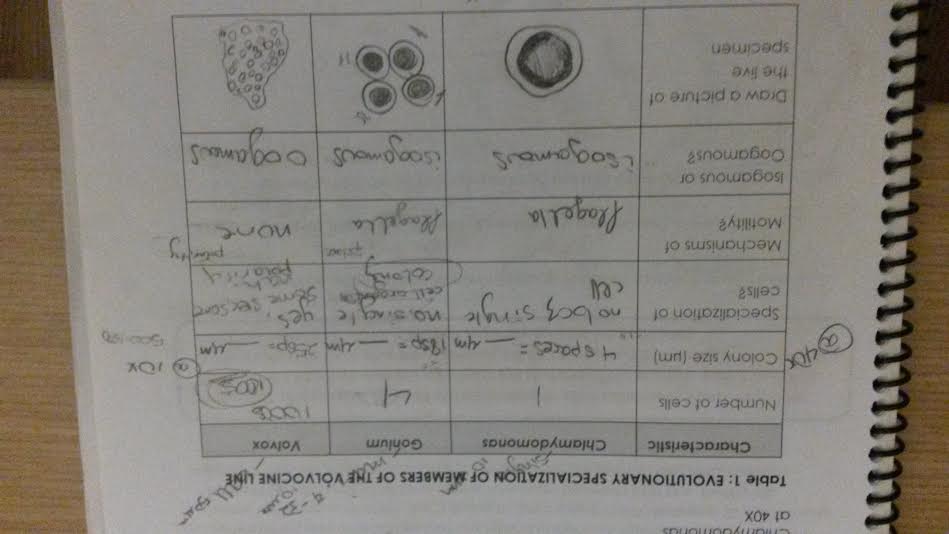
Conclusions and Future Directions
The Hay Infusion that was created at the end of the lab period will wait until the following week to be tested for organisms, but since there are already a good number of living things present in the transect, like trees and bushes, it is safe to say that there will be many more living organisms to discover under a microscopic lens. In the future if this experiment were to be repeated, it would be interesting to have multiple infusions with various amount of milk in them. This would be likely to change how much growth there is in the infusion and is likely to result in much higher rates of bacterial presence.
Lab 2 Identifying Algae and Protist 22 January 2015
Purpose
This lab was focused on identifying various unicellular eukarya so that we could use our knowledge of the eukarya to determine what organisms were present in the Hay Infusion we created during the last experiment. Given that there were many biotic features present in our Hay Infusion, we hypothesized that there would be a lot of unicellular eukarya growing in our jar.
Materials and Methods
•Wet mount of known organisms o Focus on the organism and describe it, measure it (at 40x the objective spaces are 2.5 micrometers) •Use dichotomous key to identify the organism observed •Repeat with samples of Hay Infusion o Two samples from different parts of the jar: 1. Settled film at the top, 2. Bottom near the sediment •Keep track of the organisms in notebook, measuring size and using the dichotomous key for identification •Prep for next lab: labeled four 10 mL tubes of sterile broth with 10^-2, 10^-4, 10^-6 and 10^-8 •Four nutrient agar plates and four nutrient agar plates with tet, labeled each 10^-3, 10^-5, 10^-7, 10^-9 •100 microliters of the Hay Infusion is added to the broth with the 10^-2 label, 100 microliters of that is added to the 10^-4, repeat for 10^-6 and 10^-8 •Plated by adding 100 microliters of the sample 10^-2 to the 10^-3 agar plate •Left to incubate for 1 week
Data and Observations
The Hay Infusion culture that our group looked at had turned greenish brown and the leaves and large debris had settled to the bottom. It smelled like rotting and trash. When we took the first sample it was off the top where the film had settled. Here we found chlamydomonas around 8 to 10 micrometers long, colpidium that seemed to be around 50 micrometers, and paramecium bursaria that were near 70 micrometers long. The sample we took from the bottom yielded an unidentifiable paramecium that we though could be either eudorina or pandoma paramecium, a couple peranema around 30 micrometers, and a chlamydomonas. A final sample that we looked at since we had time came from the middle where a lot of the leaves had settled. In this sample we found more paramecium bursaria around 80 micrometers.
Conclusions and Future Directions Our hypothesis that there would be a number of microorganism in our Hay Infusion was correct, we were able to find a lot of eukarya living in the samples that we pulled and investigated. If we had done a more thorough survey of the infusion, we would have probably found more types living in the samples. Varying other factors like time, leaving the infusion for 2 or 3 weeks, or adding more milk could have produced more eukarya to observe in culture.
2.10.15 Each new entry should be entered at the top, moving older entries down. You need to include tables of results and images. You can make tables in excel and then convert them to OWW format using this website: http://excel2wiki.net. Also the text needs to be in past tense. You should be writing about what you did not just writing down the instructions from the lab manual. SK
Lab 3 Studying Bacteria
January 29, 2015
Purpose
This experiment was conducted in order to identify and categorize the microorganisms that are present in the Hay Infusions that were collected. After doing so, these microorganisms were tested for antibacterial resistance on agar plates. The final goal of this lab was to compile this information so that it could be compared against a key to determine the names of the bacteria we had uncovered.
Materials and Methods
• Observe the number of colonies on each of the plates, both agar and agar plus tet o Convert these numbers to colonies/mL • Prepare wet mounts by sterilizing a loop and scraping up part of the growth, add water • Observe it at 40x for motility and cell shape • Gram stain piece of growth by sterilizing a loop and taking some of the growth, adding water • Pass through the flame until water evaporates • Cover with crystal violet for one minute • Rinse off with water • Cover in Gram’s iodine mordant for one minute • Rinse • Flood with 95% alcohol for 10 to 20 seconds • Rinse until it runs clear • Stain with safranin for 20 to 30 seconds • Rinse • Allow to airdry, do not use coverslip • Observe at 40x to 100x (with oil) for gram + or – • Set up for PCR: transfer 1 colony to 100 micriometers of water • Incubate 10 minutes at 100 degree C o Tubes must float • Centrifuge 5 mins at 13,400rpm • Add 20 microliters of primer to PCR tube o Mix to dissolve PCR bead • Take 5 microliters of superfnatant to the 16S PCR reaction, put in PCR machine
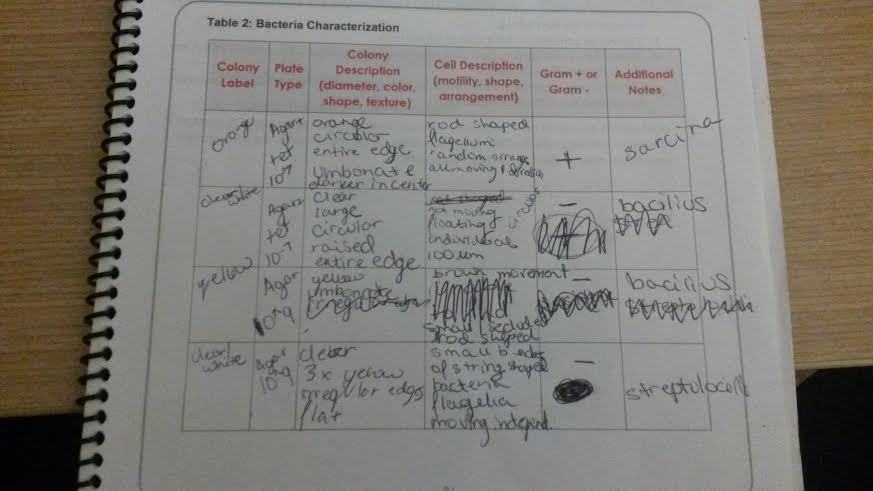
Data and Observations
In the initial colony observation, our group found that the plates with tet, the antibiotic, had significantly fewer colonies present , with the most diluted plate not having any at all and the least diluted plate with tet only having about 108 total. On the other hand the plates without tet had many more. The most diluted plate without tet had 8 colonies, while the least diluted had over 750 colonies that we could count. When we chose 4 colonies to test for motility, shape, arrangement, and gram positivity or negativity, we found that the majority of our samples were motile, with only one group floating instead of moving and the majority of the groups gram negative. The gram positive group was also motile and rod shaped, so we determined that was sarcina. The non-motile group was clear, circular, and gram negative so we determined it to be bacilius. The next followed brown movement, was gram negative, and was yellow, we determined this to be bacilius as well. The last was clear, small string shaped, and gram negative which we deemed to be streptolocilli.
Conclusions and Future Directions
This experiment was able to prove that there are in fact many bacteria present in our cultures that survived the agar and tet plates and many that prospered on just the agar plates. We were able to identify 4 instances of bacteria total on 2 different plates. In finding them we were also able to determine their resilience to the antibiotics with which they interacted. For example on the tet and agar plates 10^-7, the orange bacteria were the only resilent while on the 10^-5 the orange survived as well as some clear. However in the nontet plates the 10^-7 plates had orange and clear and white that survived. In the future, it could be cool to vary the amount of time that the antibiotics were allowed to fight so that we could determine if there is one specific type of bacteria that will beat all others out for each dilution.