Talk:20.109(F13): Mod 1 Day 5 Examine candidate clones & tissue culture
T/R lab
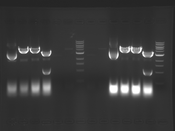
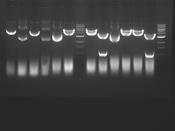
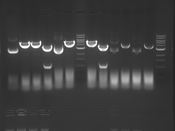
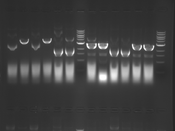
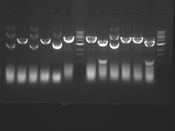
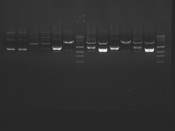
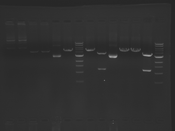
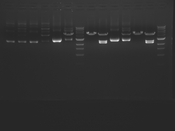
Digest Results
Note: most of these gels were run using loading dye without RNase (except for team Platinum). In our misguided attempt to keep your methods section simple, we made the call to stick with the NEB loading dye instead of our home-brew loading dye -- sorry, all! Most of the gels seem straightforward to interpret: just ignore the very thick small bands of RNA, and note that your other DNA bands may be shifted down slightly and are also curved. For example, in several images the uncut plasmid band is shifted to below the 3kbp MW marker. However, in a couple of cases we may re-run digests/gels, and will keep you posted about it. For the teams with unknown enzymes and/or samples (Green, Blue, and Pink), please modify this page and fill in these details. Feel free to provide feedback about how "interpretable" you think your gel is directly to us.
W/F lab
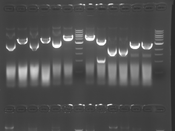
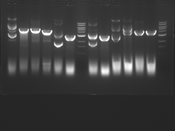
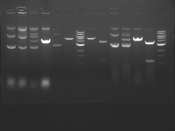
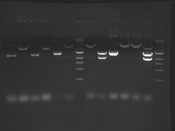
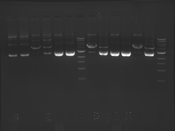
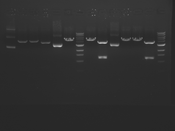
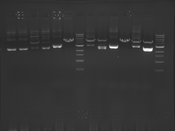
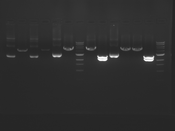
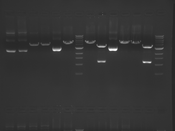
Digest Results
Note: Blue and White teams please edit the details for your enzymes :)
Ligation Results
Please put your colony count data in the correct row.
Note: "hypothetical data" just shows made-up numbers similar to what you might expect.
| Group Colour | pCX-EGFP (#) | bkb + ins, no lig (#) | bkb + lig, no ins (#) | bkb + ins, lig 1 (#) | bkb + ins, lig 2 (#) |
|---|---|---|---|---|---|
| Hypothetical Data | 1000 | 2 | 20 | 100 | 100 |
| Pink (W/F) | 896 | 3 | 1 | 337 | |
| Red (W/F) | 1096 | 0 | 6 | 67 | 100 |
| Blue (W/F) | 1140 | 1 | 4 | 231 | 270 |
| Purple (W/F) | 860 | 5 | 4 | 158 | 133 |
| White (W/F) | 1380 | 21 | 25 | 152 | 98 |
| Yellow(W/F) | 950 | 1 | 2 | 102 | 127 |
| Green(W/F) | 944 | 20 | 29 | 90 | 132 |