Talk:20.109(F12): Journal Club I
Identifying your mutant candidates
Last time in lab you electroporated the reporter cells with a library of kinasing mutants. The transformed cells were selected based on their antibiotic resistances (Ampicillin, Chloramphenicol and Kanamycin) and today you will screen the transformants for a colony or two that have the phenotype you are looking for. Recall that you are hoping to find a variant of Cph8 that makes the S-gal media darker when the mutant cells are grown in the dark. To find a cell with such a behavior, you might think that plating the cells on S-gal would be most productive approach. Unfortunately when the photography strain is plated on top of S-gal (as opposed to embedded in the agar) the color cells grown in the light and in the dark are both dark black. A different indicator media is needed, one that will vary in the range of enzyme activities that are relevant for our screen.
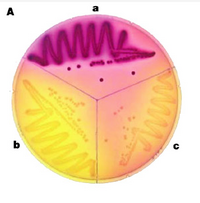
We will use MacConkey agar as the indicator of b-gal activity for our reporter strain that's been electroporated with the Cph8 library. MacConkey agar was developed more than 100 years ago [reference = MacCONKEY, A., "Lactose-fermenting bacteria in faeces" J. Hyg., 8; 333-379 (1905)]. MacConkey's media contains Neutral Red, a pH sensitive dye. When b-gal catalyzes the hydrolysis of lactose into glucose and galactose, it changes the pH by providing simple sugars for the cell to metabolize. Since H+ is a byproduct of metabolism and Neutral Red is a pH indicator, the cells growing on MacLac change color. The colonies are red when the b-gal activity is high and less red or yellow when the activity is lower.
Before we move to Journal club today, you and your partner should choose two candidates from your electroporation plates. To choose your candidates, try to find reasonably well isolated colonies to compare so you will have a clonal population to study. Then look for colonies of about the same size but different colors. The size of the colonies matters since very small colonies may not have metabolized as much lactose based simply on their reduced number of cells rather than the activity of the signaling pathway in that cell.
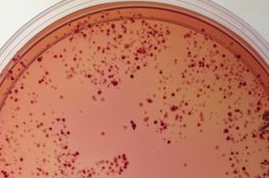
Once you have chosen your favorite 2 candidates, circle the colonies with a sharpie on the back of the petri dish, and then give the petri dish to the teaching faculty so the colony can be restreaked for additional colonies and then grown at 37° in the light and dark in LB+Amp25+Cam34+Kan10 liquid media in advance of the next lab.