Subsoontorn Lab:Research
Current Research

|
Banana (Musa spp.) is one of the most valuable agricultural crops in the world particularly in the tropical and subtropical countries. Nearly three fourth of banana plantation in Thailand grow a banana variety called “Kluai Numwa” which is highly susceptible for Panama disease caused by Fusarium oxysporum f.sp. cubense (Foc). The use of antagonistic microorganisms offers a more effective and sustainable way of controlling Foc comparing to chemical fungicides. While a number of antagonic microorganisms against Foc have been explored worldwide, biocontrol of Foc in Thailand almost exclusively focuses on the use of Trichoderma spp. In this project, we propose the development of alternative Foc biocontrol agents from rhizobacteria Pseudomonas fluorescens. Here, we plan to isolate, select, modify and prototype Foc biocontrol products from P. fluorescens in Thailand. Expected outcomes of this project includes a) discovery and characterisation of new P. fluorescens strains capable of suppressing Foc, b) foundational technology for non-transgenic genome edit P. fluorescens to have better antifungal properties, c) prototypes of P. fluorescens based bioproducts ready to field test in banana plantation in Thailand. This project is under collaboration with Dr. Wandee Wattanachaiyingcharoen (NU) and Dr. Panatda Jannoey (NU) |
|
Thailand is the forth largest ornamental fish exporter in the world and recently made siamese fighting fish national aquatic animals. Zooplanktons such as fairy shrimp and water flea are important for ornamental fish industry as a food source for nurturing fish larvae. Zooplanktons with optimal body size and nutritional contents help accelerate the growth of fish larvae and increase survival rate. Here, we propose to develop technologies for redesigning anatomy and biochemical content of fairy shrimp for siamese fight fish. Our approaches rely on a) encapsulation of high value chemical (such as nutrients, hormones or antibiotics) or cells (such as probiotic microbe) into fairy shrimp nauplii, b) selective breeding of fairy shrimp for desirable phenotype (such as optimal size or high pigment content) under microfluidic chips. The technology platforms developed in this project will help create next generation livefoods that can reduce cost and improve product quality for Thai ornamental fish industry. This project is under collaboration with Dr. Nukul Saengphan (Live feed research center) and Asso.Prof. Panwong Kuntanawat (SUT, Thailand)
|
|
Our ability to study and utilise microbiota is limited by the lack of tools for precisely perturbing and modulating microbial subpopulations of interest within a heterogenous population. Recently, CRISPR/Cas technology was used for creating antimicrobials with a programmable spectrum of activities.This strategy exploits the fact that CRISPR/Cas system can be designed to break a specific DNA sequence. In a prokaryotic cell without efficient DNA repair, such genomic cleavage often results in cell death. By delivering the designed CRISPR/Cas system to a microbial population one could selectively knockdown a subpopulation whose genomic DNA is targeted. Previous works demonstrated sequence-specific elimination of Escherichia coli and Staphylococcus aureus in mixed populations. Here, we are applying this strategy for targeted elimination of Vibrio harveyi, a pathogenic bacteria in black tiger shrimp and Pacific white shrimp. This project is under collaboration with Dr. Wanilada Rungrassamee (BIOTEC) and Prof. Jim Haseloff (University of Cambridge, UK)
|
Past Research

|
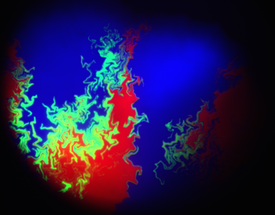
DNA transfer via conjugation plays a major role in the dissemination of antibiotic resistance among medically significant bacterial species. In this study, we developed a technique for visualising spatial distribution of conjugating bacterial population on a solid surface. Populations of donor, recipient and transconjugant cells can be distinguished using three different fluorescent reporters. We show that the fractal dimension of the interface between donor and recipient populations determines population-level conjugation efficiency. Additionally, competition for nutrients available at colony borders results in stochastic loss of cell diversity and increases variability of observed conjugation frequencies across different colonies. Our ability to monitor the dynamics of conjugation especially in the context of growth and competition within bacterial population would lead to better understanding of how antibiotic resistance spreads and how we might prevent it. This project was conducted at the laboratory of Prof. Jim Haseloff (University of Cambridge, UK)
|
.
|
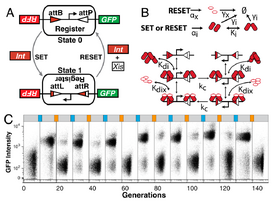
Synthetic biology promises to replace ad hoc small scale DNA engineering with formalized processes applied to realize much larger scale changes in genotypes and more radical changes in phenotypes. Our work here describes design principles and demonstrates applications of a recombinase device family to provide examples for how to compose reliable synthetic gene systems. This work includes: a) computational feasibility studies of synthetic cycle counter an example of complex synthetic gene systems requiring hundreds of genetic parts, b) experimental proof-of-concepts, failure analyses and design principles for a Recombinase Addressable Data (RAD) device built from bacteriophage integrases and excisionases. Our RAD devices are capable of storing state over one hundred cell generations and can be switched repeatedly, c) generalisation of rewritable RAD principles to three other integrase-excisionase pairs, and implement single-use two input logics, buffer gates and cascades, d) Implementation of autonomous recombinase switches driven by growth-phase dependent promoters. Taken together,the work developed here comprises an initial framework for composing complex yet reliable genetic systems using recombinase enzymes. This project was conducted at the laboratory of Prof. Drew Endy (Stanford University, USA)
|
|
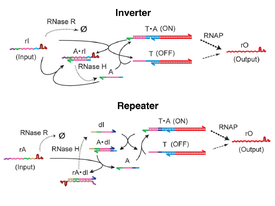
An overarching goal of synthetic and systems biology is to engineer and understand complex biochemical systems by rationally designing and analyzing their basic component interactions. Toward gradually increasing the complexity of systematically engineered systems, programmable synthetic circuits operating in cell-free in vitro environments offer a valuable testing ground for principles for the design, characterization, and analysis of complex biochemical systems. Here we illustrate this approach using in vitro transcriptional circuits (“genelets”) while developing an activatable transcriptional switch motif and configuring it as a bistable autoregulatory circuit, using just four synthetic DNA strands and three essential enzymes, bacteriophage T7 RNA polymerase, Escherichia coli ribonuclease H, and ribonuclease R. This simplicity encouraged us to characterize and model the system within a Bayesian inference framework. Our work demonstrates that programmable in vitro biochemical circuits can serve as a testing ground for evaluating methods for the design and analysis of more complex biochemical systems such as living cells. This project was conducted at the laboratory of Prof. Erik Winfree (California Institute of Technology, USA)
|