CH391L/S12/Selectablegeneticmarkers: Difference between revisions
No edit summary |
No edit summary |
||
| Line 14: | Line 14: | ||
===Antibiotic=== | ===Antibiotic=== | ||
In synthetic biology research, the primary forms of selectable markers are antibiotic resistant genes. Because a large portion of research takes place in vivo in E. coli, antibiotic selectable markers can be employed whenever transfecting DNA in order to distinguish wild-type cells from successfully transfected ones. | In synthetic biology research, the primary forms of selectable markers are antibiotic resistant genes. Because a large portion of research takes place in vivo in E. coli, antibiotic selectable markers can be employed whenever transfecting DNA in order to distinguish wild-type cells from successfully transfected ones. If ligating more than one gene of interest into a plasmid for transfection into E. coli, it is often beneficial to employ multiple antibiotic markers to ensure that both genes are present in resultant colonies. | ||
Common types of antibiotics used include ampicillin, tetracycline, chloramphenicol, and the many -mycins, including kanamycin. | Common types of antibiotics used include ampicillin, tetracycline, chloramphenicol, and the many -mycins, including kanamycin. A large range of antibiotic resistances are used as genetic markers. Because of this, each antibiotic resistance is often referred to by a three letter acronym, such as Amp, Tet, Chl, Cam and Kan. Plates containing these antibiotics can be made en mass, and used to grow appropriate cultures of transformed E. coli. | ||
Antibiotic markers are the most popular form of selectable genetic markers. As such, the field is quite large and constantly expanding in order to meet research needs. For instance, <i>Poggi et al.</i> recognized the mutation of antibiotic resistance towards gentamicin, kanamycin, streptomycin, and spectinomycin in leptospiral pathogens. The group was able to develop a cassette that included two antibiotic markers, along with a new gentamicin marker. Using multiple antibiotic markers greatly reduces the chance of background colonies that have spontaneously developed antibiotic resistance. | |||
===Herbicidal=== | ===Herbicidal=== | ||
| Line 72: | Line 74: | ||
#Parsons2011 [https://e-reports-ext.llnl.gov/pdf/476269.pdf D. Parsons, M. Tolmasky, P. Chain, B. W. Segelke] | #Parsons2011 [https://e-reports-ext.llnl.gov/pdf/476269.pdf D. Parsons, M. Tolmasky, P. Chain, B. W. Segelke] | ||
//Report by the Lawrence Livermore National Laboratory on a new system for selectable markers | //Report by the Lawrence Livermore National Laboratory on a new system for selectable markers | ||
#Poggi2010 pmid=20511419 | |||
//Multiple antibiotic resistances as selectable genetic markers in Leptospira. | |||
</biblio> | </biblio> | ||
Revision as of 05:06, 20 February 2012
Selectable Markers Overview
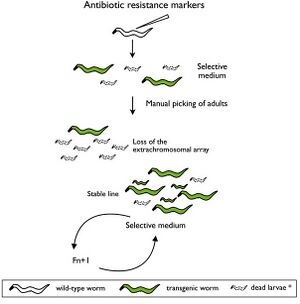
Selectable genetic markers are exogenous genes that are introduced into a cell, conferring a previously absent resistance. These markers are primarily used to "mark" the successful transformation of DNA into a plasmid. Oftentimes, selectable markers are accompanied by other exogenous genes that is the primary gene of interest; the marker simply serves to distinguish between successful transformations, and unaltered cells.
It is not atypical to witness transformation efficiencies as low as .05%, making it difficult to pick correct cellular colonies without additional techniques. This is where the selectable genetic markers prove their usefulness. For instance, selectable genetic markers can be used to confer ampicillin resistance to E. coli. These newly resistant E. coli can then be grown on culture plates with ampicillin, allowing only E.coli with successfully transformed DNA to proliferate.
In addition to selectable genetic markers are screenable genetic markers. Screenable genetic markers function in a similar manner in that they are exogenous genes that are transformed into a cell; however, they do not confer any new sort of resistance to the cell. Instead, they cause the cell to respond differently to environmental conditions in such a way as to distinguish transformed cells from untransformed cells. This can be useful when determining the transformation efficiency of a cell, or when carefully monitoring the activity of proteins.
Types of Selectable Markers
Antibiotic
In synthetic biology research, the primary forms of selectable markers are antibiotic resistant genes. Because a large portion of research takes place in vivo in E. coli, antibiotic selectable markers can be employed whenever transfecting DNA in order to distinguish wild-type cells from successfully transfected ones. If ligating more than one gene of interest into a plasmid for transfection into E. coli, it is often beneficial to employ multiple antibiotic markers to ensure that both genes are present in resultant colonies.
Common types of antibiotics used include ampicillin, tetracycline, chloramphenicol, and the many -mycins, including kanamycin. A large range of antibiotic resistances are used as genetic markers. Because of this, each antibiotic resistance is often referred to by a three letter acronym, such as Amp, Tet, Chl, Cam and Kan. Plates containing these antibiotics can be made en mass, and used to grow appropriate cultures of transformed E. coli.
Antibiotic markers are the most popular form of selectable genetic markers. As such, the field is quite large and constantly expanding in order to meet research needs. For instance, Poggi et al. recognized the mutation of antibiotic resistance towards gentamicin, kanamycin, streptomycin, and spectinomycin in leptospiral pathogens. The group was able to develop a cassette that included two antibiotic markers, along with a new gentamicin marker. Using multiple antibiotic markers greatly reduces the chance of background colonies that have spontaneously developed antibiotic resistance.
Herbicidal
Morphological
Other
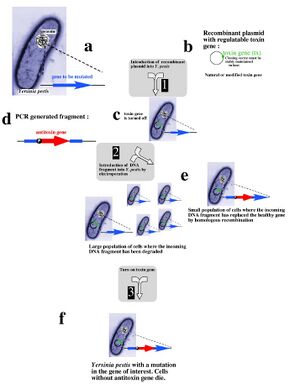
Recent research into selectable genetic markers has looked into pathways that avoid employing antibiotic and herbicidal resistance. This is due to rising concern over "wild" strains of bacteria or plants developing antibiotic or herbicidal resistance and proliferating rapidly in nature. Even in a laboratory environment, avoiding the resistance approach towards selectable markers can prove beneficial.
A novel approach towards selectable markers was developed in Lawrence Livermore National Laboratory, which employes a toxin/antitoxin combination of genes as a marker. The process, summarized in the figure to the left, effectively avoids the need to grow antibiotic resistant bacterial cultures on an antibiotic plate. An inducible zeta-toxin group of proteins is first introduced into an E. coli strain. A DNA strand of interest containing an zeta-antitoxin group is then transformed into the E. coli, and the entire culture is grown. The zeta-toxin group is then induced, killing off all E. coli that does not contain the antitoxin group. Besides for triggering the zeta-toxin group, no outside influence is required to select for the desired cells[2].
Types of Screening
Blue/White Screening
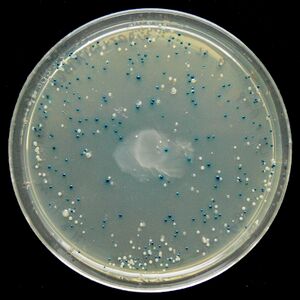
Blue/White Screening is commonly used in E. coli transformations. In this screening, cells are grown on agar plates in the presence of X-gal and IPTG to test for the presence of β-galactosidase enzyme. In the M15 strain of E. coli, part of the lacZ gene is deleted, removing the cell's ability to produce β-galactosidase. However, when transfected with a plasmid containing a lacZα domain, such as pUC19, the gene becomes operable and the cell produces β-galactosidase. It is possible to create a successful transformation in which β-galactosidase is not produced by inserting DNA into the lacZα domain. This is particularly useful to check for successful ligations. Successful ligations will not produce β-galactosidase, while unsuccessful ligations will.
X-gal, while normally colorless (i.e. white), will readily hydrolyze in the presence of β-galactosidase into a compound with a sharp blue color. Therefore, colonies with successfully transformed cells with the desired DNA will grow white, while background colonies will grow blue.
Green Fluorescent Protein Screening
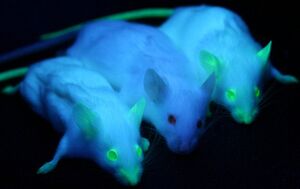
Green Fluorescent Protein, or GFP, was first isolated from the crystal jellyfish Aequorea victoria in the 1960s. In 1994, GFP was successfully cloned[4], allowing researchers to use the protein as a screenable marker for the first time. Virtually harmless in live cells, GFP has the unique pheotype of glowing bright green under ultraviolet light. GFP functions entirely of its own accord, and requires no exogenous material besides ionizing radiation in order to fluoresce. This allows GFP to be used as a marker accompanying transfected DNA, and has been used extensively in academia.
In 2011, GFP was used to create an in vivo mammary model to investigate tumorigenesis in mice. Tumor cells were introduced into the mice, accompanied with GFP as a screenable marker. As the mice tumors proliferated, so did GFP. This allowed for easy differentiate between tumors and stroma cells, greatly aiding cancer researchers[3].
Artificial Selection
Selectable markers have a remarkably relevant role in industrial applications. Because of their ability to distinguish cells from one another, selectable markers are an essential tool for artificial selection. While artificial selection of organisms is possible without the use of selectable markers, the process is significantly shorter with their use.
Artificial selection is a special instance in which selectable markers are often the desired gene to be introduced into a cell. For instance, rice has been transfected with a plethora of resistances using selectable markers. Glycopeptide binding protein, dihydrofolate reductase, and hygromycin phosphotransferase have all been introduced into rice, conferring resistance to bleomycin and pheomycin, methotrexate, and hygromycin B respectively. This allows farmers to use herbicides select for only rice with these markers, while eliminating the majority of invasive species[5].
Issues
Genetically Modified Organisms
Roundup Ready crops
References
- Giordano-Santini R and Dupuy D. Selectable genetic markers for nematode transgenesis. Cell Mol Life Sci. 2011 Jun;68(11):1917-27. DOI:10.1007/s00018-011-0670-1 |
Review article about selectable genetic markers as used in nematodes. Relatively new field for nematodes, possible due to the completion of the Caenorhabditis elegans genome.
-
D. Parsons, M. Tolmasky, P. Chain, B. W. Segelke
Report by the Lawrence Livermore National Laboratory on a new system for selectable markers
- Moen I, Jevne C, Wang J, Kalland KH, Chekenya M, Akslen LA, Sleire L, Enger PØ, Reed RK, Øyan AM, and Stuhr LE. Gene expression in tumor cells and stroma in dsRed 4T1 tumors in eGFP-expressing mice with and without enhanced oxygenation. BMC Cancer. 2012 Jan 17;12:21. DOI:10.1186/1471-2407-12-21 |
Observes tumor growth in mice by introducing GFP into the mice.
- Chalfie M, Tu Y, Euskirchen G, Ward WW, and Prasher DC. Green fluorescent protein as a marker for gene expression. Science. 1994 Feb 11;263(5148):802-5. DOI:10.1126/science.8303295 |
The first instance of using GFP as a marker.
- Caplan A, Dekeyser R, and Van Montagu M. Selectable markers for rice transformation. Methods Enzymol. 1992;216:426-41. DOI:10.1016/0076-6879(92)16039-m |
Various genetic markers used for artificial selection of rice crops.
- Poggi D, Oliveira de Giuseppe P, and Picardeau M. Antibiotic resistance markers for genetic manipulations of Leptospira spp. Appl Environ Microbiol. 2010 Jul;76(14):4882-5. DOI:10.1128/AEM.00775-10 |
Multiple antibiotic resistances as selectable genetic markers in Leptospira.