Skatebro:research prop
From OpenWetWare
Jump to navigationJump to search
Project Overview
There is much evidence to support that, under standard cellular conditions, Sus1 plays a key role in regulating SAGA activated genes by directing/confining them to the nuclear periphery for their preferential processing and export. We propose a 3-part study to look at changes in SAGA-dependent gene transcription, motility, and expression in wild-type and sus1Δ yeast strains under different cellular conditions, such as high temperature combined with galactose induction.
- Part 1: use dynamic 3D tracking in live yeast cells to visualize the motility of SAGA-dependent genes HXK1, GAL1, and INO1
- Part 2: use RNA-fluorescence in situ hybridization (FISH) to look at HXK1, GAL1, INO1 transcription levels
- Part 3: use DNA microarray to look at changes in genome-wide transcription levels in sus1Δ yeast.
Background Information
- SAGA complex dominates transcriptional activation at a minumum of 10% of the meausurable yeast genome, and these genes tend to be stress induced.
- Genes GAL1, HXK1, and INO1 have been previously studied and confirmed to be SAGA-dependent.
- Sus1 interacts with both the SAGA complex and the Sac3-Thp1-Cdc31 complex, which binds to Nuclear Pore Complexes (NPC) at the nuclear periphery.
- Under galactose induction, while Sus1 was found to be dispensable for GAL1 activation, Sus1 was found to be critical for gene motility and confinement of activated GAL loci to the nuclear periphery.
- Sus1 is hypothesized to be likely involved in transcription coupled mRNA export in a manner similar to the following schematic:
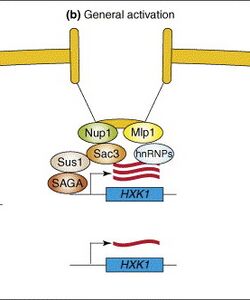
- Our Module 3 research suggested that, on galactose media and under increasing heat stress, sus1Δ yeast strains can grow more robustly than wild-type yeast strains.
- Intrigued by this interesting and unpredicted finding, we propose a research project aimed at studying Sus1's role in gene transcription, motility, and expression under different cellular conditions, for example high temperature combined with galactose induction.
Research Problem and Goals
- Specific problem: to determine if the role of Sus1 in gene regulation is altered by different cellular conditions.
- Does Sus1 somehow inhibit either the transcription, motility, or expression of SAGA-dependent genes at high temperatures?
- General goal: Expand previous research into the relation between the spatial organization of the genome and gene expression by investigating Sus1's role in the mechanism of peripheral gene recruitment in greater detail.
Project Details and Methods
- We will be working with FY2068 and the sus1Δ yeast strain we made in Module 3. Differences between these strains will give us insight into the role of Sus1 in overall gene regulation (extending our previous goal of analyzing Sus1 in SAGA).
- Methods:
- 3D live tracking of GAL1, INO1, and HXK1 in FY2068 and sus1Δ yeast at 30 C and 37 C.
- RNA-fluorescence in situ hybridization (FISH) to measure transcription levels of GAL1, INO1, and HXK1 in FY2068 and sus1Δ yeast at 30 C and 37 C.
- DNA microarray of FY2068 and sus1Δ yeast at different temperatures in the presence of different transcriptional inducers.
- Spot test to monitor viability of yeast under different conditions.
Predicted Outcomes
If all goes well:
- In wild-type yeast, GAL1 has been shown to be confined to the nuclear periphery upon intranuclear activation, while in sus1Δ yeast, GAL1 does not undergo this confinement (Cabal et al., 2006). These studies have also found that GAL1 transcription levels are unaffected by the presence or absence of Sus1. Our recent experiments suggested that the confinement of GAL genes to the nuclear periphery at higher temperatures is harmful to the yeast. We hope to confirm these findings and thus show that freer gene motility at higher temperatures allows certain genes to be better expressed.
- In wild-type yeast, INO1 has been shown to be recruited to the nuclear periphery for transcriptional activation by nuclear membrane associated Scs2 protein (Brown et al., 2007). Sus1's involvement in INO1 regulation has never been explicitly studied before, however, our recent experiments identified INO1 as having a statistically significant 8-fold increase in transcription level in sus1Δ yeast. Studying the movement of INO1 in the nucleus in both wild-type and sus1Δ yeast will allow us to further characterize the importance of Sus1 in peripheral gene recruitment. Again we hypothesize that freer gene motility at higher temperatures allows certain genes to be better expressed.
- We expect to repeat our findings of unaffected transcription levels for GAL1 and affected transcription levels for INO1 in sus1Δ yeast under normal cellular conditions. Furthermore, we expect to confirm that both genes have affected gene motility and thus gene expression in sus1Δ yeast. Also, we expect to find that high temperature affects one/all of the following in wild-type and sus1Δ yeast: gene transcription, gene motility, and/or gene expression --> and that thus sus1Δ and wild-type yeast will grow differently at high temperatures in the presence of different transcriptional inducers. Our hypothesis is that freer gene motility is crucial for gene expression and cell viability at higher temperatures.
- We found HXK1 to be down regulated 4-fold in sus1Δ yeast. HXK1 is also known to be SAGA regulated and is worthwhile to look into in addition to INO1 and GAL1.
Resources
References
- Huasinga K., Pugh B., A genome-wide housekeeping role for TFIID and a highly regulated stress-related role for SAGA in Saccharomyces cerevisiae. Molecular Cell. 2004;13(4):573-85
- Cabal G., Genovesio, A., Rodriguez-Navarro S., Zimmer C., Gadal O., Lesne A., Buc H., Feuerbach-Fournier F., Olivo-Martin J., Hurt E., Nehrbass U. SAGA interacting factors confine sub-diffusion of transcribed genes to the nuclear envelope. Nature. 2006;441(8):770-773
- Zanton S., Pugh F., Changes in genomewide occupancy of core transcriptional regulators during heat stress. PNAS:101(48);16843-16848
- Brown CR, Silver PA (2007) Transcription regulation at the nuclear pore complex. Current Opinion in Genetics & Development 17 (2): 100-106.