Silver: Assembly
Summary
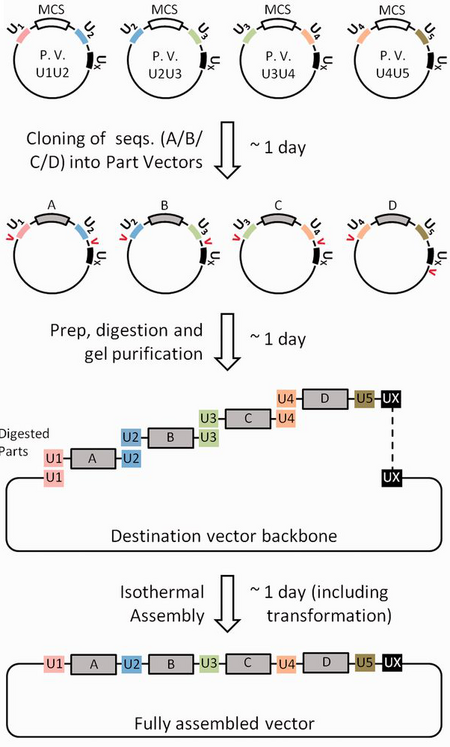
Unique nucleotide sequence (UNS)-guided DNA assembly is a variation on Gibson/isothermal assembly that is more tolerant to repeated sequence elements, e.g. those common in synthetic metabolic pathways and genetic circuits. In this approach, 40 bp UNSes are designed computationally to be unstructured and biologically inert, then attached to the termini of each DNA part to facilitate accurate recombination-based assembly. Most commonly, we perform UNS-guided assembly using a combination of standardized "Part" and "Destination" vectors containing previously designed UNSes. The sequences to be assembled are first cloned into the part vectors, attaching them to UNSes. The UNS-flanked parts are then digested out and inserted into the destination vector backbone via isothermal assembly. Assembly accuracy is high (typically > 80%), and by including multiple versions of each part in the assembly, complex libraries may be constructed.
Previous Implementations of UNS-Guided Assembly
The details required for implementation of UNS-guided assembly are described in Torella et al. 2014, "Rapid construction of insulated genetic circuits via synthetic sequence-guided isothermal assembly", as are examples of its use for the construction and optimization of bacterial metabolic pathways. We have also used this approach to build and genomically integrate mammalian genetic circuits in Lienert et al. 2014, "Two- and three-input TALE-based AND logic computation in embryonic stem cells." A new manuscript detailing the protocol we use, as well as troubleshooting steps, is forthcoming.
Implementing UNS-Guided Assembly
Basic UNS-Guided Assembly Protocol
List of Part and Destination Vectors
Obtaining Vectors for UNS-Guided Assembly
A list of available Part and Destination Vectors can be found at our Available Vectors page.
Part and Destination vectors can be obtained from the Silver Lab. E-mail jtorella@fas.harvard.edu.
Software for Designing new UNSes