Sbb14-Tae Won Chung
Project Outline
The purpose of the project is to transform thermostable aminoacyl-tRNA synthetase genes from thermophilic bacteria into E. coli. This involves following steps:
- PCR Cloning of desired genes from bacterial genome.
- Removal of Internal restriction site in case there is internal restriction site.
This is particularly done by splicing with overlap extension (SOE)
- Digesting the PCR product and ligating on to a vector.
- Transforming the plasmid into host cell
- Mini-preping sample colonies of the cell to analyze whether transformation was successful or not.
Organism
prolyl-tRNA synthetase
locus tag = TM_0514 length = 1734 bp
>gb|AE000512.1|:540171-541904 Thermotoga maritima MSB8, complete genome TCACCTGACAGCCTCCTTGGGATCGTACTCCCGCTTCATCTTTTCCAGTGTCTCCAGAAGCGCACCGAAA CCGTTTTTGATGTTCACCTTGACGAGTTCTTTCGAATAGCGTTTCTTCAGTTCAACAACACCTTCCTTGA GGGATCTTCCCACGTTTATTCTTATGGGAAAGCCTATGAGATCGGCGTCTTTGAATTTGAAACCAGGCGA GACTTCTCTGTCATCCAGAACAACCTCTTCTCCTTTTTCTGAGAGAACCTGGTAAATCTTCTCACCCACC TGCTTCTGTTCAGCGTCGTTCATGTTCAGAATGTCCACCACAACGGTGTAAGGGGCGATCGAAAGGGGCC AGATCATACCGTTCTCATCGTGGAAATGTTCCACAACAGCCGCCATCGTTCTGGAAACTCCCCAGCCATA ACAGCCCATGATGAAAGGCTTCATCTCACCGTTTTCATCCATGAAGTAGGCCTTCATGGCTTCGGAGTAT TTTGTGCCGAGTTTGAAGATGTGACCAAGCTCTATCCCTTTTGTCGCCTTGAGAGGCTCACCACAGACGG GACAGGGATCACCCTCCACCACCGTTCTCAGATCGTACCACTCGTCCACCTTGAAATCTCTGGGGTGGTT TGCGTTTACGTAGTGCGTATCCTCTTCCATACCCCCAACGACGAAGTTTTTCAGTCCTCTGACGCTGTGA TCGGCCACTTTCTTCACATCCACACCGATGGGTCCAATGAATCCGACGGGAACTCCGAAGTCCTTCAGAA TTTCTTCTGGGGTTGCCATTCTCAGCGACTGATCTTTCAAATGTGCTTTGAGCTTCGCTTCGTTGAGCTC CAGGTCTCCCCTTATAAGTACCATTACGTATCCTTCTCTTCCTTTGTAAAGAAGGGACTTCACAATCTTC GACGGAGGAACACCGAGAAACTCTGAAACTTCCTCGATCGTCTTCACCCCGGGTGTGGGAACCTTCTTGA GGGGTTTCTCTTCTTCCTGTTCCTGAGTGTATTCACCTTTGTATTCGGCTTTTTCGTCGCTTGCCTGGTA GCCGCACTTTTCACAGAAGAGTACGTTCGTCTCTCCTATTTTCGCAGGAACGACGAACTCGTGGGAAGCG TTCCCTCCGATGGCACCCGTTTCTGCCTCTATGATCATGTACCGCACACCAAGCCTTTCCATGATTCGGG AGTACGCTTTTTTGAACTGTTCGTACGTCTCGTCCAGAGATTCCCAGCTTGCGTGAAAGCTGTAAGCGTC CTTCATGATGAATTCTCTCGCCCTGAGAAGGCCGAAGCGTGGTCTGATTTCGTCTCTGTACTTGTTCGCT ATCTGATACAGAGTGAGAGGAAGCTGTTTGTATGAACGAAGTTCGTTCTTCACAAGGTCCGTGACGATCT CCTCGTGCGTGGGACCGAGTGTGAAGTCTCTTTCGTGCCTGTCTTTGAGTTTCATCATTTCGGGACCGTA ATCGTCCCACCTTCCTGACTGTTTCCAGAGTTCCGCAGGTTGAAGGATGGGCATCAGAATTTCCTGTGCC CCTATCCTGTTCATCTCTTCTCGAACTATGTTCTCTATTTTGAGAAGCACTCTTCTTCCTAATGGGAGGT AGGTGTATATACCGGCGGCTACTTTTCTTATGAAGCCTCCTCGAAGAAGATACTCGTGGCTTACTGTCTC AACATCGGAAGGGGTTTCTTTGAGAGTAGGAGCATAGAGGTCTTTCATCCGCAA
glutamyl tRNA-Gln amidotransferase, subunit A
locus tag = TM_1272 length = 1428 bp
>gb|AE000512.1|:1297318-1298745 Thermotoga maritima MSB8, complete genome GTGATCGATTTGGACTTCAGAAAACTCACAATAGAAGAGTGTCTGAAACTTTCCGAAGAAGAAAGGGAGA AACTCCCACAACTTTCCCTGGAAACCATCAAAAGACTCGATCCACACGTGAAGGCGTTCATCTCCGTAAG AGAGAATGTTTCCGTTGAGAAGAAGGGAAAATTCTGGGGAATTCCAGTTGCGATCAAGGACAACATTCTC ACACTGGGCATGAGAACGACCTGTGCCTCCAGGATACTCGAAAACTACGAATCCGTTTTCGATGCCACAG TGGTGAAGAAGATGAAGGAAGCAGGCTTTGTCGTGGTTGGGAAGGCAAACCTCGATGAGTTTGCGATGGG CTCGAGCACCGAAAGATCCGCATTCTTTCCCACAAGAAACCCCTGGGACCTGGAAAGGGTTCCCGGCGGA AGCAGTGGGGGTTCTGCCGCGGCCGTTTCTGCAGGAATGGTGGTGGCAGCACTCGGAAGCGACACGGGAG GCTCTGTGAGACAGCCCGCTTCCCTGTGCGGTGTTGTTGGGTACAAGCCCACCTACGGTCTCGTCTCAAG GTACGGTCTTGTAGCGTTTGCAAGTTCTCTGGACCAGATCGGACCCATCACAAAGACCGTAAGAGACGCT GCAATCCTCATGGAGATCATCTCGGGCCGGGACGAAAACGATGCCACGACGGTGAACAGGAAGGTGGACT TCCTCTCGGAGATAGAAGAGGGAGTCTCCGGCATGAAGTTTGCCGTTCCTGAAGAGATCTACGAACACGA TATAGAAGAAGGAGTTTCGGAAAGATTCGAAGAGGCCCTGAAACTTCTCGAGAGACTCGGTGCAAAGGTG GAGAGAGTGAAGATACCCCACATCAAGTACTCTGTTGCCACCTACTACGTCATCGCTCCAGCCGAGGCGA GTTCAAACCTTGCCAGATTCGACGGTGTGAAGTACGGCCTCAGGATAAAAGAGAAAGGCCTCAGGGAGAT GTACATGAAGACAAGAAACGTGGGCTTCGGTGAGGAAGTGAGAAGAAGGATCATGATCGGAACCTTCACC CTCAGCGCTGCCTACTACGAAGCCTATTTCAACAAGGCCATGAAAGTGAGGAGAAAGATCTCCGATGAGC TCAACGAGGTGCTCTCGCAGTACGACGCGATCCTCACTCCCACCTCCCCGGTGACGGCCTTCAAGATCGG AGAGATAAAAGATCCTCTCACCTACTACCTCATGGACATCTTCACGATACCCGCAAACCTCGCGGGACTT CCCGCGATCAGTGTGCCGTTCGGGTTCTCAAACAATCTCCCCGTTGGTGTTCAGGTGATAGGCAGAAGGT TCGCGGACGGGAAAGTCTTCAGAATAGCGAGGGCCATAGAGAAGAACTCCCCATACAACGAAAACGGCAT GTTCCCGCTCCCGGAGGTGAAAGCATGA
glutamyl tRNA-Gln amidotransferase, subunit B
locus tag = TM_1273 length = 1449 bp
>gb|AE000512.1|:1298742-1300190 Thermotoga maritima MSB8, complete genome ATGAGATACAGACCCGTCATAGGTCTTGAGATACACGTTCAGCTTTCAACGAAGACAAAAGCGTTCTGCT CCTGCCCGGCGGATGTCTTCGAGCTTCCTCCGAACACGGCGATCTGTCCCGTCTGCACCGGCCAGCCGGG GGCCCTTCCCGTTCCCAACGAAGAGATGATCAGGTTCGCCGTGAAGACCGCCCTTGCGTTGAACTGCAAG ATTCACAAGTATTCCAGATTCGACAGAAAGAACTACTTCTATCCTGACCTTCCCAAGGGATACCAGATCA GCCAGTACTTCTACCCGATCGCAACCGAGGGTTTCCTGGAGATAGACGGAGACGAAGGAAGGAAAAAAGT CAGAATCAGAAGGCTCCACCTCGAGGAAGACGCTGGGAAACTCGTCCACGAAGGGGACTCCATCACCCGT GCGAGCTACTCTCTGGTCGATATGAACAGATGCGGTGTTCCTCTCATCGAGATCGTCACAGAGCCCGATA TTTCTTCTCCGAGAGAAGCTCGCGTTTTCATGGAGAAGCTGAGATCCATCGTGAGGTACCTCGGCGTGAG CACGGGAGACATGGAAAAGGGAGCGCTCAGATGCGACGCGAACATCTCCGTTGTTGACACAGAAACGGGA AGACAGAGCAACAGGGTGGAAGTGAAGAACATGAACTCTTTCAGGTTCGTCGAAAGGGCTTTGGAGTACG AGTTCGAGCGAATTGTGAAGGCGATGGAGAGAGGAGAAGACGTGGAAAGAGAGACGAGAGGCTGGGACAT GGCCACGAAGATCACCGTTTCCATGAGAGGAAAAGAAGAAGAGAGCGACTACAGGTACTTCCCGGAGCCG GACATACCGCCCGTTGTTCTCTCCGACGAGTATCTCGAAGAGGTGAAAAAGGAACTCCCAGAGCTTCCAG ACGAGAAGGCAGAGCGATTCATGAGAGAGTACGGCCTTCCGGAGTACGACGCGAAGGTGCTCACCTCGAG TAAAGAACTCGCTGAGTTCTTCGAAGAGTGTGTGAAGGTGGTGAACAGACCGAAGGATCTCAGCAACTGG ATCATGACGGAAGTTCTGAGAGAACTCAACGAAAGAAACATCGAAATCACAGAGTCAAAACTGACCCCTC AGCACTTCGCTGATCTCTTCAAATTGATGGACGAAGGAAAAATCTCCATAAAGATCGCAAAGGAGATATT CCCGGAAGTCTTCGAAACAGGAAAGATGCCCTCCCAGATCGTGGAGGAAAAGGGTTTGACTCAGATCAAC GACGAAAAACTCATCGAGGAACTGGTGAAAAAGGCGATGGAACAGAATCCAAAGGCCGTTCAGGATTACA AATCAGGCAAAAAGAAAGCGGCAGGGTTTTTCGTGGGATACGTGATGAGAGAGACAAAAGGAAAAGCGAA TCCGGAACTCACGAACAGGATCATTCAAAAACTCCTCGAGGGGGAGTGA
glutamyl tRNA-Gln amidotransferase, subunit C
locus tag = TM_1273 length = 291 bp
>gb|AE000512.1|:264194-264484 Thermotoga maritima MSB8, complete genome ATGATAAAGGTTACAAAGGACCTAGTGCTTCATCTTGAAAATCTGGCAAGGTTAGAACTCTCCGAAGACC AGAGAGAAAGTCTGATGAAAGATTTTCAGGAGATACTCGATTACGTGGAGCTCCTCAACGAAGTCGACGT GGAGGGTGTGGAGCCAATGTACACACCCGTTGAGGACAGCGCCAAACTCAGAAAAGGAGATCCGAGGTTC TTTGAAATGCGGGACCTCATAAAGAAGAACTTCCCGGAAGAAAAAGACGGTCACATAAAAGTCCCCGGAA TCCACAGATGA
- Note that prolyl-tRNA aa synthetase is backward.
- Subunit A and Subunit B of the Gln amidotransferase are neighbors. There is a 4bp overlap between two genes. How this overlap is transcribed is unknown. I tried submitting the sequences to BLAST to find the actual sequence of the whole enzyme, but I did not retrieve any useful information.
Construction Files
Prolyl-tRNA aa synthetase
PCR ProRS-F/EcoRI-SOE-R on pcrpdt (485 bp, SOE_A) PCR EcoRI-SOE-F/ProRS-R on pcrpdt (1296bp, SOE_B) PCR ProRS-F/ProRS-R on A+B (1755 bp, newpcrpdt) Digest newpcrpdt (NcoI/EcoRI, 1739+10+6, L, pcrdig) Digest pBAD-myc-his-A (NcoI/EcoRI, 4053+41, L, vectdig) Ligate pcrdig and vectdig (pBAD-ProRS) ----------------------------------------- >ProRS-Fwd Forward primer for PCR of ProRS ccagtCCATGGcacggatgaaagacctctatgc >ProRS-Rv Reverse primer for PCR of ProRS ccagtGAATTCtcacctgacagcctccttggg >EcoRI-SOE-F Forward primer for SOEing PCR of ProRS CAGGGCGAGAGAGTTCATCATGAAGG >EcoRI-SOE-R Reverse primer for SOEing PCR of ProRS CCTTCATGATGAACTCTCTCGCCCTG
Glutamyl-tRNA Gln amidotransferase
PCR Gln AB RS-F/EcoRI-SOE-R on pcrpdt (205 bp, A) PCR EcoRI-SOE-F/Gln AB RS-R on pcrpdt (2718bp, B) PCR Gln AB RS-F/Gln AB RS-R on A+B (2894 bp, newpcrpdt) Digest newpcrpdt (NcoI/EcoRI, 2878+10+6, L, pcrdig) Digest pBAD-myc-his-A (NcoI/EcoRI, 4053+41, L, vectdig) Ligate pcrdig and vectdig (pBAD-Gln AB) ----------------------------------------- >Gln AB RS-Fwd Forward primer for PCR of Gln AB RS ccagtCCATGGcaatcgatttggacttcagaaaac >Gln AB RS-Rv Reverse primer for PCR of Gln AB RS ccagtGAATTCtcactccccctcgaggag >EcoRI-SOE-F Forward primer for SOEing PCR of Gln AB RS gaaaattctgggGAATcCcagttgcgatc >EcoRI-SOE-R Reverse primer for SOEing PCR of Gln AB RS gatcgcaactgGgATTCcccagaattttc
Glutamyl-tRNA Gln amidotransferase
PCR GlnC-F/GlnC-R on Thermotoga Maritima genome (312 bp, pcrpdt) Digest pcrpdt (NcoI/EcoRI, 300+10+6, L, pcrdig) Digest pBAD-myc-his-A (NcoI/EcoRI, 4053+41, L, vectdig) Ligate pcrdig and vectdig (pBAD-Gln C) ----------------------------------------- >GlnC-F Forward primer for PCR of Gln C RS ccagtCCATGGcaataaaggttacaaaggacctagtgc >GlnC-R Reverse primer for PCR of Gln C RS ccagtGAATTCtcatctgtggattccgggg
May 1st and Result
This was the last lab period for the semester.
I did mini-prep purification procedure to extract the plasmid DNA from the cells. The DNA was then submitted to DNA sequencing.
- Miniprep purification of DNA
MINIPREP (2mL) Procedure for Plasmid DNA Purification 1. Pellet 2 mL saturated culture by spinning full speed, 30 seconds in a 2mL Microcentrifuge tube. 2. Dump supernatant 3. Add 250uL of P1 buffer into each tube. Resuspend the cells thoroughly 4. Add 250uL of P2 buffer (a base that denatures everything and causes cells to lyse). Gently mix up and down. Solution should become clearer. 5. Add 350uL of N3 buffer (an acid of pH ~5 that causes cell junk - including protein and chromosomal DNA - to precipitate, and leaves plasmids and other small molecules in solution). Slowly invert a few times, then shake. 6. Spin in centrifuge at top speed for 5 minutes. 7. Label blue columns with an alcohol-resistant lab pen. 8. Pour liquid into columns, and place the columns into the centrifuge. Spin at full speed for 15 seconds. 9. Dump liquid out of the collectors under the columns (the DNA should be stuck to the white resin) 10. Wash each column with 500 uL of PB buffer. 11. Spin in centrifuge at full speed for 15 seconds, then flick out the liquid again. 12. Wash with 750uL of PE buffer (washes the salts off the resins). 13. Spin in centrifuge at full speed for 15 seconds and flick out liquid again. 14. Spin in centrifuge at full speed for 90 sec to dry off all water and ethanol. 15. Label new Microcentrifuge tubes and put columns in them. 16. Elute them by adding 50uL of water down the middle of the column (don't let it stick to the sides). 17. Spin in centrifuge at top speed for 30 seconds. 18. Take out columns and cap the tubes.
In this way, I prepared DNA extractions of prolyl-tRNA synthetase and glutamyl tRNA-Gln amidotransferase subunit C, which I then submitted to sequencing.
On May 2nd, the sequencing results came back.
- Pro RS Sample Sequencing Result
GGGGCTAAATAAGGCAGAAGTCCACATTGATTATTTGCACGGCGTCACACTTTGCTATGCCATAGCATTTTTATCCATAAGATTAGCGGATCCTACCTGACGCTTTTTATCGCAACTCTCTACTGTTTCTCCATACCCGTTTTTTGGGCTAACAGGAGGAATTAACCATGGCACGGATGAAAGACCTCTATGCTCCTACTCTCAAAGAAACC CCTTCCGATGTTGAGACAGTAAGCCACGAGTATCTTCTTCGAGGAGGCTTCATAAGAAAAGTAGCCGCCGGTATATACACCTACCTCCCATTAGGAAGAAGAGTGCTTCTCAAAATAGAGAACATAGTTCGAGAAGAGATGAACAGGATAGGGGCACAGGAAATTCTGATGCCCATCCTTCAACCTGCGGAACTCTGGAAACAGTCAGGAAG GTGGGACGATTACGGTCCCGAAATGATGAAACTCAAAGACAGGCACGAAAGAGACTTCACACTCGGTCCCACGCACGAGGAGATCGTCACGGACCTTGTGAAGAACGAACTTCGTTCATACAAACAGCTTCCTCTCACTCTGTATCAGATAGCGAACAAGTACAGAGACGAAATCAGACCACGCTTCGGCCTTCTCAGGGCGAGAGAGTTCA TCATGAAAGACGCTTACAGCTTTCACGCAAGCTGGGAATCTCTGGACGAGACGTACGAACAGTTCAAAAAAGCGTACTCCCGAATCATGGAAAGCTCTTGGTGTGCGGTACATGATCATAGAGCAGACGGTGCGGCATCGGAGGGACGCTCCACGAGTCGTCGTCTGCAAATAGACAGAAGACGACGTACTCTTCTGTGAAAGCGGTCCTCC AGC AGCGAAAGCGATACACGGGATACTCTGACGGAGTGTTGACGAAGAAACCTCCAGTTCACCGCGGAGTGGATCGTTCAGGTCCGTCCCCGATGAGCTCTCCAAAAGAAGAGTTTGGAGCG
- Gln ADT C Sample Sequencing Result
GGGCTTATACTGCGAAGTCCACATTGATTATTTGCACGGCGTCACACTTTGCTATGCCATAGCATTTTTATCCATAAGATTAGCGGATCCTACCTGACGCTTTTTATCGCAACTCTCTACTGTTTCTCCATACCCGTTTTTTGGGCTAACAGGAGGAATTAACCATGGCAATAAAGGTTACAAAGGACCTAGTGCTTCATCTTGAAAATCTG GCAAGGTTAGAACCCTCCGAAGACCAGAGAGAAAGTCTGATGAAAGATTTTCAGGAGATACTCGATTACGTGGAGCTCCTCAACGAAGTCGACGTGGAGGGTGTGGAGCCAATGTACACACCCGTTGAGGACAGCGCCAAACTCAGAAAAGGAGATCCGAGGTTCTTTGAAATGCGGGACCTCATAAAGAAGAACTTCCCGGAAGAAAAAGA CGGTCACATAAAAGTCCCCGGAATCCACAGATGAGAATTCGAAGCTTGGGCCCGAACAAAAACTCATCTCAGAAGAGGATCTGAATAGCGCCGTCGACCATCATCATCATCATCATTGAGTTTAAACGGTCTCCAGCTTGGCTGTTTTGGCGGATGAGAGAAGATTTTCAGCCTGATACAGATTAAATCAGAACGCAGAAGCGGTCTGATAA AACAGAATTTGCCTGGCGGCAGTAGCGCGGTGGTCCCACCTGACCCCATGCCGAACTCAGAAGTGAAACGCCGTAGCGCCGATGGTAGTGTGGGGTCTCCCCATGCGAGAGTAGGGAACTGCCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCCTAAGTAGGAC AAATCCGCCGGGAGCGGATTTGAACGTTGCGAAGCAACGGCCCGGGAGGGTGGCGGGTATGACCCCCGTCATAGACTGCTTGGGCATCAATTTAGGCGTGATGGCGTTGTTGTCTGATGTCATTTGTGGGTTTCTACAGATCTATAGTTATA
Both results indicate that cloning and transformation of Pro RS and Gln ADT C were successful. The sequences are almost identical to its construction files.
April 29th and 30th
The ligated plasmids from previous week were transformed into the host cell using heat-shock transformation.
- Transformation by heat-shock procedure
1. Thaw a 200 uL aliquot of cells on ice 2. Add 50 uL of water to the cells (if greater volume is desired) 3. Add 30 uL of KCM to the cells 4. Put your ligation mixture on ice, let cool a minute or two (for Miniprep product, dilute by 10, then use 1uL of dilution) 5. Add 70 uL of the cell cocktail to the ligation, stir to mix 6. Let sit on ice for 10 min 7. Heat shock for 90 seconds at 42 (longer incubation may work better) 8. Put back on ice for 1 min 9. Add 100uL of 2YT, let shake in the 37 degree incubator for 1 hour 10. Plate 70+ uL on selective antibiotics, let incubate at 37 degrees overnight
The lab did not have enough plates for the entire class, so I divided a single plate into two parts, each labeled "Pro" for ProRS, and "GlnC" for GlnADT C transformation.
The next day, I observed that there were many colonies on both sides of the plate, indicating that transformation of either original Pbad plasmid or our ligated plasmid had occurred.
I picked out 2 colonies for each part (2 for ProRS, and 2 for GlnADT C), and incubated in 4ml LB media overnight, for the mini-prep purification. The rest of the plate was stored in the refrigerator.
April 24th
The vectors (from Karim) were ligated today, following the procedure:
- Ligation procedure:
1. Set up the following reaction: 6.5uL ddH2O 1uL T4 DNA Ligase Buffer 1uL Vector digest 1uL Insert digest 0.5uL T4 DNA Ligase 2. Pound upside down on the bench to mix 3. Give it a quick spin to send it back to the bottom of the tube 4. Incubate on the benchtop for 30min 5. Put on ice and proceed to the transformation
The two ligated products were then labeled after the inserts, ProRS and GlnADT C, and were stored in ther freezer.
April 22nd
No fruitful experiment could be done, as all the gel trays were taken by other groups. I tried digesting my Pbad vector, except I needed the gel purifying step for this, and without the trays, nothing could be done. I decided to use Karim's digested Pbad vector for ligation step.
April 17th
I gel-purified the digestion products from 4/10. I first ran a agarose gel, then cut out the band which were in their approximate location. For a higher resolution, I rant the gel for longer period of time.
April 15th
Purified DNAs from 04/10 were digested with NcoI and EcoRI. (as with our vector)
- Digestion procedure(30ul reaction mixture):
1. Set up the following reaction: 24uL of eluted PCR product 3uL of NEB Buffer 2 1.5uL NcoI 1.5uL EcoRI 2. Incubate at 37 degrees on the thermocycler for 1hr 3. Run an agarose gel, and melt with 600uL ADB buffer at 55 degrees. ****NOTE: If you are running short of time, this is an acceptable stopping point 4. If the DNA is shorter than 300bp, add 250uL of isopropanol and mix prior to loading it on the column
As step 3 says, since I did not have enough time to purify the DNAs, I stopped at mixing the products with ADB, and then stored the products in the freezer
April 10th
I examined my last attempt to SOE the pieces.
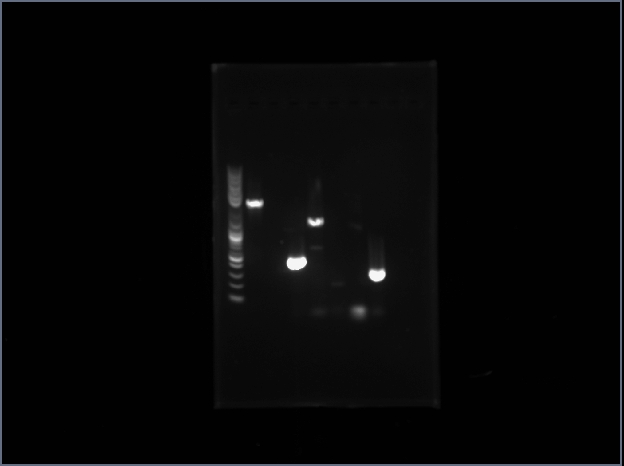
ProRS seemed to have been made judging by the size of the band.
GlnAB still failed. At this point I decided to stop working on GlnAB, and there could be some possible errors associated with this.
The first lane is the ProRS, and the next two lanes are their subparts. We see that the two parts have nicely annealed onto a whole product.
The 5th lane is where GlnAB should have been. Yet I do not see any visible trace of the product.
April 8th
April 3rd
I retrieved the PCR products from the freezer. I run an agarose gel to perform gel-purification step. I saved the GlnADT C for digestion step later. For ProRS SOE_A and ProRS SOE_B, the bands were cut and dissolved in a same vial in 650 ml ADB buffer. Similarly,Gln AB SOE_A and Gln AB SOE_B bands were cut and dissolved in a same vial in 650 ml ADB buffer.
These products (except GlnC) were then Zymo-cleaned, each dissolved in 12ul H2O.
So far, I have 12ul solution of ProRS SOE A+B, GlnAB SOE A+B, and 30ul solution of GlnC.
I set up the SOEing reaction, but with DMSO.
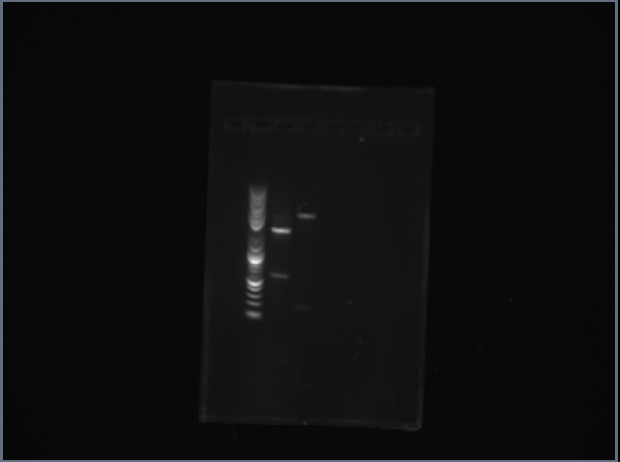
The 1st lane is ProRS subparts, and the 2nd lane is Gln AB subparts.
April 1st, Restarting
I restarted the experiment from the scratch again. This time, I replaced all the PCR step with DMSO version for better accuracy.
Following reaction mixture was made, and then put into a cycler.
20.7uL ddH2O 3.3ul DMSO 3.3uL 10x Expand Buffer (labeled 2) 3.3uL dNTPs (2mM in each) 1uL Oligo 1, 10uM 1uL Oligo 2, 10uM 0.5 uL Template DNA (Thermotoga maritima genomic DNA) 0.5uL Expand Polymerase (labeled 1)
Instead of 2K55 and 4K55 cycles, I chose 2K45 and 4K45 this time.
March 13-20th
The SOEing from 03/11 was a messy product. SOE would not anneal into a single product. GlnAB subunits especially demonstrated a large product that seemed to be larger than 4k bp. Over this period, I tried to fix the problem in one way or another, say carrying the SOE reaction in DMSO etc. After some analysis, I concluded that at some point, oligos were switched and the subparts (SOE) of the SOEing were also probably wrong.
March 11th
I gel-purified the PCR products from 03/06, and proceeded to SOEing reaction for ProRS and GlnADT AB.
- Splicing by Overlap Extention procedure:
1. Set up PCR reactions according to your construction file as normal 33uL reactions as described in Cloning by PCR 2. For each PCR, load 6uL of PCR product premixed with 4uL of loading buffer in a single well of a 1% agarose gel 3. Cut out the bands, put them into a single 1.5mL microcentrifuge tube 4. Add 650uL of ADB Buffer 5. Proceed with the Zymo Gel Purification procedure 6. Elute the DNA in 50uL of water 7. Set up your second round of PCR as a normal 33uL reaction using the eluted mixture of fragments as template
March 6th
First set of PCR reactions were set up. Following reaction mixture was made, and then put into a cycler.
24uL ddH2O 3.3uL 10x Expand Buffer (labeled 2) 3.3uL dNTPs (2mM in each) 1uL Oligo 1, 10uM 1uL Oligo 2, 10uM 0.5 uL Template DNA (Thermotoga maritima genomic DNA) 0.5uL Expand Polymerase (labeled 1)
So 3 sets of reactions were carried.
1) Prolyl-tRNA synthetase (ProRS)
2) Gln amidotransferase subunit AB (GlnAB)
3) Gln amidotransferase subunit C (GlnC)
March 4th
Genomic DNA and Oligos arrived. The oligos came in following amounts.
1. Pro RS Fwd: 32.4 nmol
2. Pro RS Rv: 31.0 nmol
3. Pro SOE Fwd: 25.1 nmol
4. Pro SOE Rv: 31.5 nmol
5. Gln subunit A+B Fwd: 36.0 nmol
6. Gln subunit A+B Rv: 29.1 nmol
7. Gln subunit A+B SOE Fwd 34.8 nmol
8. Gln subunit A+B SOE RV: 37.0 nmol
9. Gln subunit C Fwd: 37.0 nmol
10. Gln subunit C Rv: 29.1 nmol
These oligoes were diluted to 10 uM solution by first diluting the oligos into 100uM mixture (for example, for Pro RS Fwd, I diluted it in 324ul water), and then diluted 10-fold again.