SPRI bead mix
Curators
Other formats
Abstract
This protocol describes the preparation of stocks and buffers for inexpensive, convenient, and scalable DNA and RNA purification from aqueous solutions by solid-phase reversible immobilization (SPRI) on carboxylated paramagnetic beads. It also describes how to validate the effectiveness of the mixes before use.
The bead mixes described in this protocol are drop-in substitutes for AMPure XP and RNAClean XP beads (Beckman Coulter), but at about 1/100 of the cost (~$0.55/mL vs. $15–$70/mL at current Canadian prices).
Materials
Beads
- Sera-Mag™ Magnetic SpeedBeads™, carboxylated, 1 μm, 3 EDAC/PA5 (GE Healthcare Life Sciences #65152105050250) – warning: the bead suspension contains 0.05% sodium azide
Chemicals (molecular biology grade)
- Common
- Sodium chloride (NaCl)
- Poly(ethylene glycol), avg. mol. wt. 8000 (PEG 8000)
- Polysorbate 20 (Tween 20)
- Hydrochloric acid (HCl) concentrate
- Nuclease-free water
- For DNA mix only
- Tris(hydroxymethyl)aminomethane (Tris base)
- Disodium ethylenediaminetetraacetate dihydrate (EDTA)
- For RNA mix only
- Trisodium citrate dihydrate
Consumables
- 50 mL conical tubes
- 1.5 mL microcentrifuge tubes
- Disposable weighing vessels
- Disposable Pasteur pipettes
- Parafilm
- 0.22 μm syringe filters
- 10 mL disposable syringes
- 25 mL, 10 mL, 5 mL serological pipettes
- 1000 μL, 200 μL micropipette tips
Equipment
- Milligram-range balance
- Funnels
- Spatulas
- Heating plate
- Rotary mixer
- Microcentrifuge
- 25 mL graduated cylinder
- 50 mL volumetric flasks and stoppers
- 1000 μL, 200 μL adjustable-volume micropipettes
- Squirt bottle
- Magnetic separation block for 1.5 mL microcentrifuge tubes
Stock solutions
- Common solutions
- 1 N HCl
- 5 M NaCl
- 10% (v/v) Tween 20
- 50% (w/v) PEG 8000
- DNA solutions
- 1 M Tris base
- 0.1 M EDTA
- RNA solution
- 1 M trisodium citrate
Preparing stock solutions
Making 1 N HCl
Prepare at least 10 mL 1 N HCl in a glass bottle from available concentrated stock.
Making 50 mL of 1 M Tris base, 0.1 M disodium EDTA, 1 M trisodium citrate and 5 M NaCl stocks
In 50 mL volumetric flasks, prepare a separate stock solution for each of the following components with the specified weights of solids.
- Common solution
| 5 M NaCl | 14.610 g |
- DNA solutions
| 1 M Tris base | 6.057 g |
| 0.1 M Na2-EDTA•2H2O | 1.861 g |
- RNA solution
| 1 M Na3-citrate•2H2O | 14.705 g |
Some gentle heating may be necessary. Ensure the solution comes back to room temperature before completing the volume to the mark on the flask. Store in 50 mL conical tubes. ![]() Optional: filter the stocks with the syringes and filters to remove undissolved solids. It is strongly recommended to filter the solutions used for making RNA mix for sterilization.
Optional: filter the stocks with the syringes and filters to remove undissolved solids. It is strongly recommended to filter the solutions used for making RNA mix for sterilization.
Making 50 mL of 10% (v/v) Tween 20 stock
- Place a labeled 50 mL conical tube on the balance and tare it.
- With a new disposable Pasteur pipette, aspirate 1-5 mL of Tween 20.
- Slowly dispense the Tween 20 into the 50 mL conical tube to reach 5.475 g.
- Remove the tube from the balance and add 45.0 mL of water with a 25 mL serological pipette.
- Cap the tube and mix on a rotary mixer for one hour to dissolve the viscous liquid.
Making 25 mL of 50% (w/v) PEG 8000 stock
- Place the 25 mL graduated cylinder on the balance and tare it.
- Weigh 12.5 g of PEG 8000 powder directly into the cylinder. It is recommended to use a fresh pair of gloves to reduce static charges that make the powder fly off the spatula.
- Add no more than 14 mL of nuclease-free water with a serological pipette on top of the PEG powder in the cylinder. The water level will reach over the 25 mL mark as the cylinder already contains about 20 mL of dry powder. Be sure not to fill the cylinder completely, as some air is required to make mixing possible. If the cylinder is too small, use a 50 mL one.
- Seal the cylinder with a double layer of Parafilm.
- Shake vigorously to suspend the powder in the water until there are no more lumps of dry solid sticking to the cylinder wall. It will be very viscous and clumpy.
- Let the suspension stand at room temperature for at least an hour to allow the solids to dissolve and the air bubbles to rise.
- Remove the Parafilm and complete the volume with nuclease-free water to the 25 mL mark.
- Seal the cylinder again and mix well by inverting. The solution is very viscous and homogenizing it can take a while.
- Transfer the solution to a 50 mL conical tube for storage. There will be some loss inside the cylinder but you need only 20 mL for one batch of bead mix.
This recipe can be scaled up with larger cylinders to make mixing easier, for example making 50 mL of solution in a 100 mL cylinder.
Buffer recipes
Nucleic acid elution and storage buffers
- DNA buffer
- TE+Tween (10 mM Tris base, 1 mM EDTA, 0.05% Tween 20, pH 8.0 @ 25 °C)
- RNA buffer
- Citrate+Tween (1 mM trisodium citrate, 0.05% Tween 20, pH 6.4 @ 25 °C)
Ingredients for 50 mL
- DNA buffer
| Nuclease-free water | 48.564 mL |
| Tris base, 1 M | 0.500 mL |
| Disodium EDTA, 0.1 M | 0.500 mL |
| Tween 20, 10% (v/v) | 0.250 mL |
| HCl, 1 N | 0.186 mL |
- RNA buffer
| Nuclease-free water | 49.679 mL |
| Trisodium citrate, 1 M | 0.050 mL |
| Tween 20, 10% (v/v) | 0.250 mL |
| HCl, 1 N | 0.021 mL |
These solutions are used for preparing the beads before adding them to the mix. They are also useful for DNA and RNA elution and storage. It is possible to make concentrates of these solutions for convenience. Keep them refrigerated.
Nucleic acid binding bead mixes
- DNA mix
- 10 mM Tris base, 1 mM EDTA, 2.5 M NaCl, 20% PEG 8000, 0.05% Tween 20, pH 8.0 @ 25 °C
- RNA mix
- 1 mM trisodium citrate, 2.5 M NaCl, 20% PEG 8000, 0.05% Tween 20, pH 6.4 @ 25 °C
![]() Read the mixing instructions below before starting to combine the ingredients.
Read the mixing instructions below before starting to combine the ingredients.
Ingredients for 50 mL
- DNA binding bead mix
| NaCl, 5 M | 25.000 mL |
| Nuclease-free water | 3.582 mL |
| Tris base, 1 M | 0.500 mL |
| Disodium EDTA, 0.1 M | 0.500 mL |
| HCl, 1 N | 0.168 mL |
| PEG 8000, 50% (w/v) | 20.000 mL |
| Tween 20, 10% (v/v) | 0.250 mL |
| Sera-Mag bead suspension | 1.000 mL |
- RNA binding bead mix
| NaCl, 5 M | 25.000 mL |
| Nuclease-free water | 4.672 mL |
| Trisodium citrate, 1 M | 0.050 mL |
| HCl, 1 N | 0.028 mL |
| PEG 8000, 50% (w/v) | 20.000 mL |
| Tween 20, 10% (v/v) | 0.250 mL |
| Sera-Mag bead suspension | 1.000 mL |
Mixing instructions
- Mix the Sera-Mag beads very well to resuspend.
- Quickly transfer 1 mL to a 1.5 mL microcentrifuge tube (the beads settle quickly).
- Place the tube on a magnet stand until the supernatant is clear, about 30 s.
- Remove and discard the supernatant.
- Add 1 mL of previously prepared "DNA buffer" or "RNA buffer", depending on the kind of bead binding mix you are preparing, to the bead pellet and close the tube.
- Remove the tube from the magnet and resuspend the beads by vortexing for at least 15 seconds. Spin down the liquid with a microcentrifuge.
- Put the tube back on the magnet until the beads clear.
- Remove and discard the supernatant.
- Repeat steps 5 to 8 twice, for a total of 3 washes with the appropriate buffer, leaving the supernatant in the tube after the last wash.
- In a new 50 mL conical tube, combine the nuclease-free water, NaCl and HCl. For DNA, also add the Tris and EDTA. For RNA, add only the trisodium citrate instead. Cap and mix well.
- Remove the buffer supernatant from the bead tube still on the magnet.
- Add 1 mL of incomplete binding buffer (prepared at step 10) to the bead tube on the magnet.
- Remove the bead tube from the magnet and resuspend by vortexing for 15 s. Briefly spin down the liquid without pelleting the beads.
- Add the washed beads to the incomplete binding buffer. Cap and vortex for 30 s.
- With a 25 mL serological pipette, add 20 mL of 50% PEG stock. Dispense slowly and allow the viscous liquid to slide down the inside walls of the pipette to ensure an accurate volume is added.
- Add the Tween 20.
- Cap the tube and mix by inversion gently but thoroughly, until the color appears homogeneous.
The bead binding mix is ready to be used or validated. Store at 4 °C.
Important considerations and recommendations
Chelating agents
EDTA and citrate may interfere with some enzymatic reactions by sequestering divalent cations such as Mg2+ and Mn2+. These ions may damage nucleic acids or activate contaminating nucleases. On the other hand, sequestering these ions may interfere with downstream reactions that require them; if so, you can compensate by adding ions equimolar to the EDTA or citrate.
pH
The pH titrations for the buffers and bead mixes were calculated with the Python package ionize 0.8.0. They may be inaccurate for the bead mixes due to the very high ionic strengths of those solutions. Colour-change pH indicators will also be inaccurate for the same reason. A properly calibrated pH meter may be able to measure these solutions correctly. Keep in mind that the bead mix will be diluted during use when added to the sample to be purified, which will change the ionic strength and thus the pH.
Tween 20
Adding Tween 20 to the solutions described in the protocol is optional but provides multiple benefits. It reduces adhesion of nucleic acids to plastics, which is increased during SPRI due to the high ionic strength. This improves sample recovery. Tween 20 also reduces surface tension, which can pull beads off the pellet during supernatant removal. This effect becomes very useful if the pellet is very small. If Tween 20 is not compatible with your downstream processes or if foaming is a problem, replace its volume with nuclease-free water when mixing the solutions.
Validation
It is recommended to validate the bead mixes before use to ensure their effectiveness. They can be compared to AMPure XP or RNAClean XP, or to a previous batch of homemade mix. Validation can be done with DNA or RNA that is representative of a typical usage scenario, a DNA ladder (note that NEB ladders may contain modifications that make their SPRI behaviour unrepresentative of normal DNA), fragmented DNA across a range of sizes, or an RNA standard.
Example with DNA smear
Fragment λ phage genomic dsDNA (e.g. Thermo Scientific #SD0011) with the NEBNext® dsDNA Fragmentase® kit (NEB #M0348S) for 20 minutes, according to the manufacturer's instructions, and purify the reaction product with 2 volumes of previously validated beads, then elute in TE+Tween. This produces a flat smear (50 to 2000 bp) that is easy to see on an agarose gel or an Agilent Bioanalyzer DNA 1000 chip.
Re-purify the DNA with the reference DNA bead mix and the new DNA bead mix side-by-side using 2 volumes of beads. After elution, test the DNA on agarose gel or Bioanalyzer to compare the repurified samples with each other and with the original fragmented stock. If the yields and size distributions from the two bead mixes are equivalent, the new mix is ready for use.
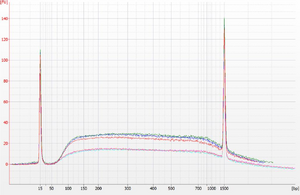
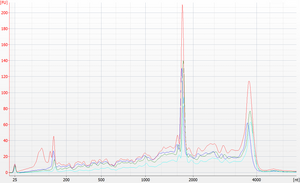
Example with RNA standard
To test the RNA bead mix, dilute FirstChoice Human Brain Reference RNA (Life Tech #AM6050) 100-fold to 10 ng/μL in Citrate+Tween. Purify the dilution using the RNA bead mix and elute in Citrate+Tween. Load the original input and the purified output on an Agilent Bioanalyzer RNA 6000 Pico chip.