SBB11Ntbk-Keith Licardo
~~!~~
To My Project Page
To Keith's Personal Page
To Other Notebooks/Protocol
Keith Licardo 14:02, 15 February 2011 (EST)
sbb1103 part PCR
Expand PCR with oligos ss28f and ss28r on MG1655 genome.
The oligos were already diluted at 10 uM and so I used them provided oligos directly.
Followed the Expand PCR protocols as listed in Spring 140L website. Used Expand Buffer.
The part was around 700 bp long so I decided to use 2K55 for the thermocycler algorithm.
Keith Licardo 14:01, 17 February 2011 (EST)
sbb1103 PCR Analytical Gel
Set-up into 0.5 mL tube: 5 uL of loading dye and only 2 uL of PCR product. Someone else did the gel upstairs.
Lane 11 was mine. The fragment in that lane matched the 700-kb fragment on the DNA ladder lane. This indicates that I was able to take out the part from the MG1655 genome.
The rest of the PCR product was then put through zymo clean up using the protocol from the BE140L site. I left the DNA fragments in the regular zymo filter for around 15 minutes as I had to go upstairs to do the preparative gel (gel purification) for Bth8112. The DNA might be damaged if it dried up in the zymo filter. I followed the rest of the protocol and washed the fragments twice with the zymo wash buffer and dried it up with the centrifuge for 90 secs.

Keith Licardo 14:16, 17 February 2011 (EST)
cut-and-paste from one vector to another for Bth8112
Digesting out Bth8112
Cutting Bth8112 out from vector pBca9145.
Followed the protocol for "EcoRI/BamHI Digest of PCR Products." Used 0.5 uL of the pBca9145-Bth8112. Added everything to the original tube for pBca9145-Bth8112. Then added the following: 3 uL of DDW, 1 uL of NEB buffer, 0.5 uL of EcoRI, 0.5 uL of BamHI. Spun with the mini spin after adding each. Pipetted all of the 10 uL from the original into a PCR tube but had to pipet out the rest at 0.5 uL three times. The PCR tube was then mini-spun and incubated for an hour.
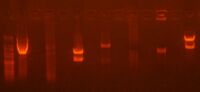
The following is gel picture for the gel purification step.
Lane 5 was mine. Although the lane shows two blurry bands, the bands became more vivid with more time running the gel. Professor Anderson cut mine for me. The smaller fragment was put into a tube with 600 uL of ADB buffer and left in the 55 deg Celsius bath for about 15 minutes.
Keith Licardo 16:26, 18 February 2011 (EST)
cut-and-paste from one vector to another for part Bth8112
Zymo Gel Purification of Bth8112
Followed the protocol for the Zymo gel purification of Bth8112 from yesterday. (Note: Sung-won mentioned that leaving the fragments in the gel might have been a better stopping point than leaving the fragments in the gel and ADB buffer.) The professor cut the gel for me, and no isopropanol was added as my fragment was larger than 300 bp. After the ADB buffer and liquified gel was filtered out, the fragments was then washed twice with 200 uL AW wash. The fragments was then dryed out for 90 seconds in the centrifuge. Finally, the fragments were eluted out with 8.5 uL of DDW.
Keith Licardo 13:38, 22 February 2011 (EST)
The class as a whole had been using the AW wash buffer instead of A4 Wash Buffer for the zymo clean up. Because of this, Prof. Anderson redid all our PCR and zymo cleanups.
There were no cut-and-paste material for EcoRI/BamHI digest PCR products (for part Bth8112) and so only one of my parts (sbb1103) could be digested and ligated.
I did the EcoRI/BamHI digestion for sbb1103. Added the following to a PCR tube: 8 uL of the part, 1 uL NEB buffer 2, 0.5 of EcoRI, and 0.5 of BamHI. I minispun the tube and incubated for an hour in the thermocycler. After the incubation, the DNA fragments were gel purified. The one who designated to do the gel prep added the loading dye with the fragments and mixed the solution before adding to the wells. The gel ran until the fragments were 3/4 across the gel. Afterwards, I cut the fragment out of the gel (700 bp) and melted the gel away by adding 600 uL of ADB buffer and incubating for 15 minutes.
Keith Licardo 13:56, 24 February 2011 (EST)
sbb1103
I did the Zymo gel purification for sbb1103 which was in a solution with 600 uL of ADB buffer. The liquid content was first taken out by spinning the solution in a Zymo filter twice to completely remove the ADB buffer. Then it was washed twice with 200 uL of A4 wash buffer and dried for 90 seconds. The fragments were then eluted out with 8 uL of DDW.
Bth8112
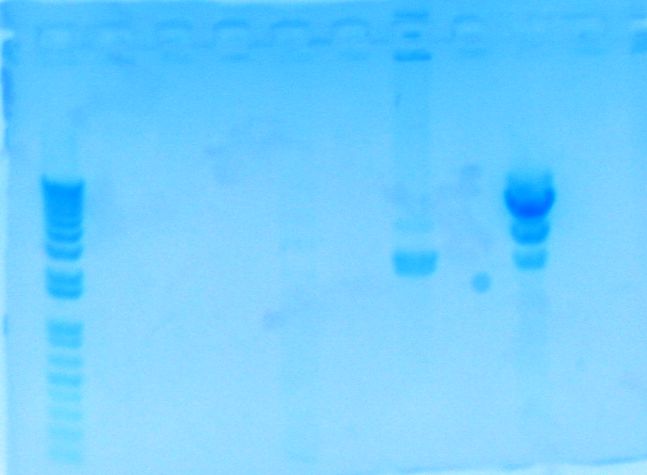
Digested the PCR product labelled "8112" with EcoRI/BamHI with the following added into a PCR tube: 6 uL of what I assumed was the PCR product of part Bth8112, 2 uL of DDW, 1 uL NEB buffer 2, 0.5 uL of EcoRI, and 0.5 uL of BamHI. I then incubated the PCR tube at 37 degrees Celsius for an hour in the thermocycler. After an hour, the tube was sent upstairs for the gel prep. The following is the image of the gel. My fragments in lane 1 were not present.
Keith Licardo 12:53, 1 March 2011 (EST)
sbb1103
Last week, the digestion fragments were obtained from gel prep last week and cleaned up with Zymo gel purification. The eluted products were then prepared.
The ligation and transformation of sbb1103 was done today. The ligation reaction consists of the following: 6.5 uL of DDW, 1 uL of T4 Ligase Buffer, 1 uL of 1601KC, 1 uL insert digest, and 0.5 uL of T4 DNA ligase. The reaction mixture was mini-spun and left on the benchtop for 30 minutes. The cell aliquot (yellow tube) was thawed on ice. 50 uL of DDW and 30 uL of KCM were added. 70 uL of the cell cocktail was then added to the ligation reaction mixture, stirred, and left to sit for 10 minutes. After 10 minutes, the ligation mixture with the cell cocktail was heat-shocked for 1.5 minutes at 42 degrees Celsius and put back on ice for 1 minute. 100 uL of LB was added and left to incubate at 37 degrees Celsius for an hour. After the incubation in the shaker at 37 degrees Celsius, the cells were plated on the Kanamycin resistant plates. The magic wand was submerged in the ethanol and put over the burner flame. The plate was then incubated over night. The GSIs picked the colonies and grew them for the miniprep.
Keith Licardo 13:39, 8 March 2011 (EST)
sbb1103 miniprep
Four colonies were picked and grown on culture tubes over night. All the colonies on the plate were white.
The 2 mL from the saturated culture tubes was centrifuged in 2 mL tube. None of the pellet was red. The supernantn was dumped. 250 uL of P1 buffer was added to the 2mL tube to resuspend the pellets with. 250 uL of P2 buffer was then added to the tube and the mixture was inverted a few times. The mixture became clear. Finally, 350 uL of N3 buffer was added and the tubes were then inverte da few times again. The tubes were centrifuged at 14000 rpm for 5 minutes.
The mixture was then poured into the blue column. The columns were centrifuged for 15 seconds. This was repeated again to remove all the liquid. The column was then washed with 500 uL of PB buffer (15 seconds) and 750 uL PE buffer(15 seconds), the dried for 90 seconds. The samples were then eluted with 50 uL of water (30 seconds).
Keith Licardo 14:24, 8 March 2011 (EST)
sbb1103 analytical mapping
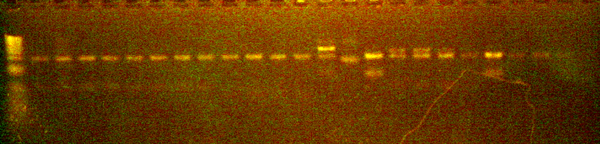
The following were added to PCR tubes for each of the miniprep product: 4 uL DDW, 4 uL miniprep product, 1 uL NEB buffer, 0.5 uL EcoRI, and 0.5 uL BamHI. The tubes were incubated at 37 degrees Celsius for 30 minutes and run on an analytical gel.
Keith Licardo 11:34, 31 March 2011 (EDT)
sbb1103 sequencing
The analytical gel verified the insertion of the part. Plasmids from colonies 1 and 2 were sent out for sequencing. The sequencing data was then analyzed by 3 groups. The chromatogram and sequence analysis was sent to JCA.
Keith Licardo 10:50, 7 April 2011 (EDT)
Team 3: "Joey's Angels"
Keith, Vini, Joey, Shane, Amy
We received our cells (promoters with GFP upstream) in a 96-well block on Tuesday. The cells were then plated for Kan resistance. We used 12 plates overall. Each plate was divided three-ways to accomodate all 35 cells with different plasmids. After speaking with the GSIs, we decided not to do one of our negative controls. This control entailed measurement of green fluorescence on untransformed MG1610. The plates were grown overnight.
The colonies were picked the next day. Two colonies were picked for each type of transformed MG1610. Because 70 colonies would need to be grown, we decided to grow them in 24-well blocks. Three blocks were used.
5 ml of LB with Kan was used to grow the cells. The LB was first transferred on a 50-ml centrifuge tube to prevent contamination of the stock. 5-ml pipette tips were then used to transfer LB from this tube and into the blocks. The pipette tips were flame-sterilized on all axis to prevent contamination. The toothpicks used to transfer the colonies were similarly sterilized. The toothpicks were left on the blocks first and removed one at a time after the entire block was filled with colonies. The block was then covered with an adhesive sticker which allows for Oxygen to diffuse through. The blocks were left to incubate at 37 deg Celsius and shaken at 224 rpm in the incubation room in the fourth floor. The plates were covered with parafilm and left in the Anderson lab refrigerator.
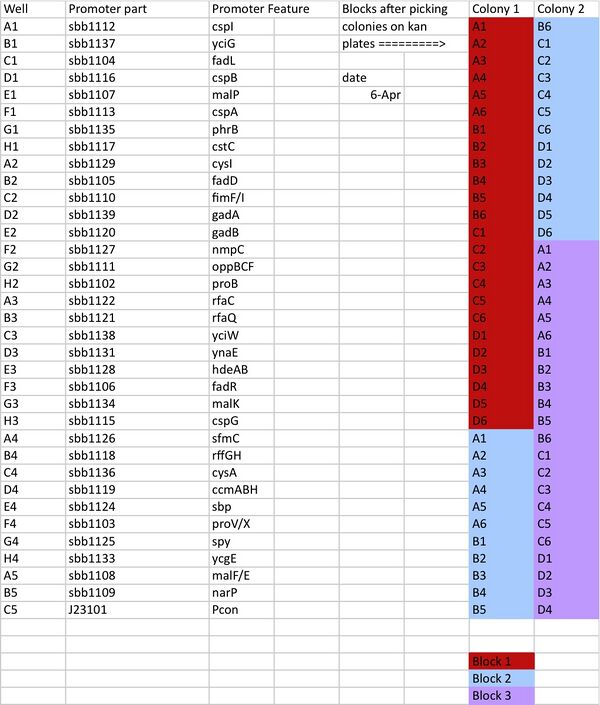
The following spreadsheet points from which transformed cell line each well in the 24-well block came.
Keith Licardo 18:29, 7 April 2011 (EDT)
Team 3: "Joey's Angels"
Keith, Vini, Joey, Shane, Amy
The 24-well blocks were shaken in the incubation room in the 4th floor. However, the blocks were designed to be placed in the shaker in the Andersen lab. Because of this, the blocks detached despite the tape used and one block placed near the edge of the shaker fell. This was block 3 which contained half of the second set of colonies. A new set of colonies was picked today to replace block 3.
Keith Licardo 13:22, 19 April 2011 (EDT)
Team 3: "Joey's Angels"
Keith, Vini, Joey, Shane, Amy
Shane transferred 1 mL from each of the 24-well blocks into a 96-well block. This was grown overnight without any shaking.
Acidic Condition
The acidic solutions were made by Shane and Joey. 10 uL of the confluent Kan LB was added to 90 uL of acidic Kan LB into the 96-well Tecan plate. This was run using the overnight program for the Tecan. OD and GFP was measured.
Heat Shock Condition
100 uL from the confluent Kan LB was added to 900 uL in a 96-well block. The block was the one added to the heat bath (47 deg C) 10 minutes prior to the Tecan measurement. The data points were for 0, 10, 30, 90, and 180 minutes. OD and GFP was measured.
Cold Shock Condition
100 uL from the confluent Kan LB was added to 900 uL in the 96-well plates. The block was the one added to the cold bath (15 deg C) 10 minutes prior to the Tecan measurement. he data points were for 0, 10, 30, 90, and 180 minutes. OD and GFP was measured.
Keith Licardo 14:38, 21 April 2011 (EDT)
The rest of my notebook entries are in the Team 3 "Joey's Angel" notebook.